Team:Lethbridge/Notebook/Lab Work/September
From 2010.igem.org
(→September 2, 2010) |
Liszabruder (Talk | contribs) (→AV) |
||
| (45 intermediate revisions not shown) | |||
| Line 279: | Line 279: | ||
</table><br> | </table><br> | ||
| - | Added 48(µL) to each reaction. | + | Added 48(µL) to each reaction. <br> |
| + | [[image:Lethbridge_100903AssemblyADS.jpg|200px]] | ||
| - | |||
---- | ---- | ||
<b>Objective:</b> Characterize xylE in LB Media vs. M9 Minimal Media.<br> | <b>Objective:</b> Characterize xylE in LB Media vs. M9 Minimal Media.<br> | ||
| Line 295: | Line 295: | ||
---- | ---- | ||
| + | ===<font color="white">JV=== | ||
| + | |||
| + | <b>Objective:</b> PCR the pSB1C3 plasmid from part J04450 miniprep.<br> | ||
| + | <table border ="3"> | ||
| + | <tr><td><b>Component</b><td><b>1X(µL)</b> | ||
| + | <tr><td>Milli-Q H<sub>2</sub>O<td>12.8 | ||
| + | <tr><td>10x Pfu Buffer with MgSO<sub>4</sub><td>2 | ||
| + | <tr><td>dNTPs<td>1 | ||
| + | <tr><td>SB-prep-26 Primer<td>1 | ||
| + | <tr><td>SB-prep-3P Primer<td>1 | ||
| + | <tr><td>Template DNA<td>2 | ||
| + | <tr><td>Pfu polymerase<td>0.2 | ||
| + | </table><br> | ||
| + | |||
| + | Used iGEM cycle PSB1C3 in thermocycler. | ||
| + | Ran PCR product on 1% TAE agarose gel. | ||
| + | |||
| + | ---- | ||
| + | ===<font color="white">KG=== | ||
| + | |||
| + | <b>Objective:</b> PCR of xylE (mRBS-xylE K118021) with fusion standard so that an arginine tail can be attached.<br> | ||
| + | |||
| + | <table border ="3"> | ||
| + | <tr><td><b>Component</b><td><b>1X(µL)</b><td><b>Master Mix(x6)(µL)</b> | ||
| + | <tr><td>Milli-Q H<sub>2</sub>O<td>12.8<td>76.8 | ||
| + | <tr><td>10x Pfu Buffer with MgSO<sub>4</sub><td>2<td>12 | ||
| + | <tr><td>dNTPs<td>1<td>6 | ||
| + | <tr><td>xylE Fusion Prefix Forward Primer<td>1<td>6 | ||
| + | <tr><td>xylE Fusion Suffix Reverse Primer<td>1<td>6 | ||
| + | <tr><td>Template DNA<td>2<td>12 | ||
| + | <tr><td>Pfu polymerase<td>0.2<td>1.2 | ||
| + | </table><br> | ||
| + | |||
| + | PCR- Conditions: | ||
| + | 1. 95<sup>o</sup>C for 180 sec | ||
| + | 2. 95<sup>o</sup>C for 30 sec | ||
| + | 3. 56.6 +/- 5<sup>o</sup>C for 30 sec | ||
| + | 4. 72<sup>o</sup>C for 150 sec | ||
| + | 5. 72<sup>o</sup>C for 600 sec | ||
| + | (35 cycles) | ||
| + | |||
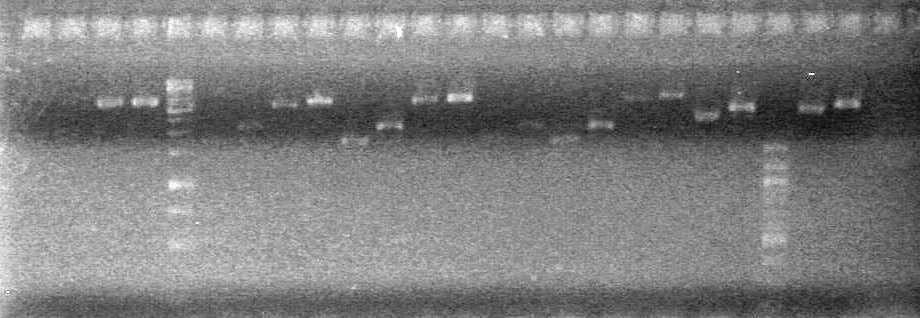
| + | Analyzed PCR on 2% agarose gel which ran at 100 V for 90 minutes.<br> | ||
| + | [[image:100915PCR AV KG JV.jpg|200px]] | ||
| + | |||
===<font color="white">AS=== | ===<font color="white">AS=== | ||
<b>Objective:</b> Assemble xylE-dT using 3AB assembly and 3AB assembly.<br> | <b>Objective:</b> Assemble xylE-dT using 3AB assembly and 3AB assembly.<br> | ||
| Line 385: | Line 429: | ||
<b>Method:</b> Run 1% TAE agarose gels at 100 V for 90 minutes. <br> | <b>Method:</b> Run 1% TAE agarose gels at 100 V for 90 minutes. <br> | ||
| - | + | [[image:100914ADSAssemblyPrep.jpg|100px]] | |
==<font color="white">September 16, 2010== | ==<font color="white">September 16, 2010== | ||
| Line 393: | Line 437: | ||
<b>Results:</b> Gel of miniprepped RFP against previously prepared RFP showed a band of the right size.<br> | <b>Results:</b> Gel of miniprepped RFP against previously prepared RFP showed a band of the right size.<br> | ||
| - | + | [[image:100916 JV RFP BW.jpg|50px]] | |
==<font color="white">September 17, 2010== | ==<font color="white">September 17, 2010== | ||
| Line 417: | Line 461: | ||
<tr><td>Pfu DNA Polymerase</td><td>0.2</td></tr> | <tr><td>Pfu DNA Polymerase</td><td>0.2</td></tr> | ||
</table> | </table> | ||
| + | |||
| + | PCR conditions: | ||
| + | <table><table border ="3"> | ||
| + | <tr><td><b>Step</b></td><td><b>Temperature (<sup>o</sup>C)</b></td><td><b>Time (mins)</b></td><td><b>Number of cycles</b></td></tr> | ||
| + | <tr><td>Initial Denaturation</td><td>95</td><td>2</td><td>1</td></tr> | ||
| + | <tr><td>Denaturation</td><td>95</td><td>0.5</td><td>25</td></tr> | ||
| + | <tr><td>Annealing</td><td>48.2, 52.4, 56.0 (gradient)</td><td>0.5</td><td>25</td></tr> | ||
| + | <tr><td>Extension</td><td>72</td><td>2</td><td>25</td></tr> | ||
| + | <tr><td>Final Extension</td><td>72</td><td>10</td><td>1</td></tr> | ||
| + | </table> | ||
| + | |||
| + | [[image:100915PCR AV KG JV.jpg|200px]] | ||
| + | |||
| + | Results: Amplification showing in gel, but fragment amplified was ~1000bp. RFP and dT should be ~800bp. Primers were checked and discovered the YFP/CFP primers were not compatible with the RFP. Concluded the amplification resulted from sloppy annealing of N-term fus prefix to the bio prefix resulting in the full construct being amplified (~1000bp) | ||
| + | ---- | ||
| + | |||
| + | ===<font color="white">KG=== | ||
| + | |||
| + | <b>Objective:</b> PCR xylE to attach fusion standard to the N-term and standard suffix to C-term.<br> | ||
| + | |||
| + | <table border ="3"> | ||
| + | <tr><td><b>Component</b><td><b>1X(µL)</b><td><b>Master Mix(x6)(µL)</b> | ||
| + | <tr><td>Milli-Q H<sub>2</sub>O<td>12.8<td>76.8 | ||
| + | <tr><td>10x Pfu Buffer with MgSO<sub>4</sub><td>2<td>12 | ||
| + | <tr><td>dNTPs<td>1<td>6 | ||
| + | <tr><td>Fusion Forward Primer<td>1<td>6 | ||
| + | <tr><td>Standard Reverse Primer<td>1<td>6 | ||
| + | <tr><td>Template DNA<td>2<td>12 | ||
| + | <tr><td>Pfu polymerase<td>0.2<td>1.2 | ||
| + | </table><br> | ||
| + | |||
| + | Fusion standard prefix has melting temperature of 66.5<sup>o</sup>C. Standard suffix has melting temperature of 70.5<sup>o</sup>C. | ||
| + | Annealing temperature of 61.5<sup>o</sup>C with 5 <sup>o</sup>C gradient. | ||
| + | PCR- Conditions: | ||
| + | 1. 95<sup>o</sup>C for 180 sec | ||
| + | 2. 95<sup>o</sup>C for 30 sec | ||
| + | 3. 61.5 +/- 5<sup>o</sup>C for 30 sec | ||
| + | 4. 72<sup>o</sup>C for 150 sec | ||
| + | 5. 72<sup>o</sup>C for 600 sec | ||
| + | (35 cycles) | ||
| + | |||
| + | ---- | ||
| + | <b>Objective:</b> PCR xylE to attach fusion standard to the C-term and standard suffix to N-term.<br> | ||
| + | |||
| + | <table border ="3"> | ||
| + | <tr><td><b>Component</b><td><b>1X(µL)</b><td><b>Master Mix(x6)(µL)</b> | ||
| + | <tr><td>Milli-Q H<sub>2</sub>O<td>12.8<td>76.8 | ||
| + | <tr><td>10x Pfu Buffer with MgSO<sub>4</sub><td>2<td>12 | ||
| + | <tr><td>dNTPs<td>1<td>6 | ||
| + | <tr><td>Standard Forward Primer<td>1<td>6 | ||
| + | <tr><td>Fusion Reverse Primer<td>1<td>6 | ||
| + | <tr><td>Template DNA<td>2<td>12 | ||
| + | <tr><td>Pfu polymerase<td>0.2<td>1.2 | ||
| + | </table><br> | ||
| + | |||
| + | Standard prefix has melting temperature of 68.7<sup>o</sup>C. Standard suffix has melting temperature of 61.6<sup>o</sup>C. | ||
| + | Annealing temperature of 56.6<sup>o</sup>C with 5 <sup>o</sup>C gradient. | ||
| + | |||
| + | PCR- Conditions: | ||
| + | 1. 95<sup>o</sup>C for 180 sec | ||
| + | 2. 95<sup>o</sup>C for 30 sec | ||
| + | 3. 56.6 +/- 5<sup>o</sup>C for 30 sec | ||
| + | 4. 72<sup>o</sup>C for 150 sec | ||
| + | 5. 72<sup>o</sup>C for 600 sec | ||
| + | (35 cycles) | ||
==<font color="white">September 18, 2010== | ==<font color="white">September 18, 2010== | ||
| Line 437: | Line 546: | ||
* Plated cells in presence of IPTG, Fe or IPTG and Fe on AMP(+) plates. Incubated at 37 <sup>o</sup>C. | * Plated cells in presence of IPTG, Fe or IPTG and Fe on AMP(+) plates. Incubated at 37 <sup>o</sup>C. | ||
** Preparation of these samples was achieved by taking a 60(µL) cell sample and inoculating 5 mL LB media. These solutions were incubated at 37 <sup>o</sup>C for 30 min. To 1 mL of culture, the appropriate amount of Fe2+ [0.938(µL)], Fe3+[1.88(µL)] or IPTG [1(µL)] was added. | ** Preparation of these samples was achieved by taking a 60(µL) cell sample and inoculating 5 mL LB media. These solutions were incubated at 37 <sup>o</sup>C for 30 min. To 1 mL of culture, the appropriate amount of Fe2+ [0.938(µL)], Fe3+[1.88(µL)] or IPTG [1(µL)] was added. | ||
| + | ---- | ||
| + | ===<font color="white">TF=== | ||
| + | |||
| + | <b>Objective:</b> Obtain preparations of B0015 (dT) and J04450 (RFP). | ||
| + | |||
| + | <b>Method:</b> Used [[Team:Lethbridge/Notebook/Protocols|Plasmid DNA Purification by Alkaline Lysis (Large Scale AKA Maxiprep]] protocol. | ||
| + | |||
| + | <b>Results:</b> Chloroform, DNA and isopropanol were mixed and therefore no products could be recovered. | ||
| + | ---- | ||
| + | |||
| + | |||
==<font color="white">September 20, 2010== | ==<font color="white">September 20, 2010== | ||
| Line 454: | Line 574: | ||
<tr><td>8</td><td>RFP2</td><td>10</td><td>2</td></tr> | <tr><td>8</td><td>RFP2</td><td>10</td><td>2</td></tr> | ||
<tr><td>9</td><td>RFP3</td><td>10</td><td>2</td></tr></table><br> | <tr><td>9</td><td>RFP3</td><td>10</td><td>2</td></tr></table><br> | ||
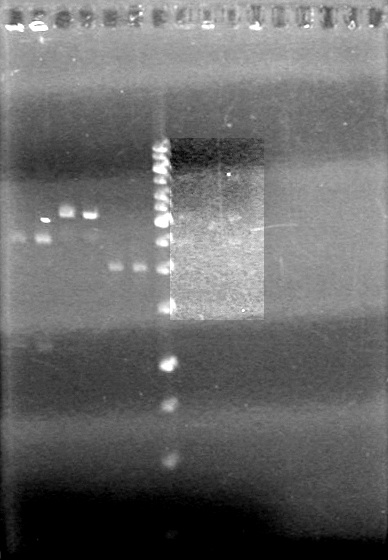
| - | <b>Results:</b | + | <b>Results:</b> <br> |
| + | [[image:100920.jpg|100px]] | ||
*mms6 was not amplified | *mms6 was not amplified | ||
*RFP was amplified | *RFP was amplified | ||
| Line 461: | Line 582: | ||
***We believe that the suffix segment of the fusion primer annealed rather than the FP segment. This would cause the promoter (R0040) of J04450 to be amplified, adding ~200bp, giving a final size of >1000bp. | ***We believe that the suffix segment of the fusion primer annealed rather than the FP segment. This would cause the promoter (R0040) of J04450 to be amplified, adding ~200bp, giving a final size of >1000bp. | ||
---------------------------------------------------------------------------------------------------------------------- | ---------------------------------------------------------------------------------------------------------------------- | ||
| - | <b>Objective:</b> Confirm assembly (3 antibiotic) of K118021 and B0015 from Sept. | + | <b>Objective:</b> Confirm assembly (3 antibiotic) of K118021 and B0015 from Sept. 14, 2010 via PCR.<br> |
<b>Method:</b> Set up 50uL reaction mixture with 2uL of ligated DNA. Used VF2 and VR primers.<br> | <b>Method:</b> Set up 50uL reaction mixture with 2uL of ligated DNA. Used VF2 and VR primers.<br> | ||
Used PFU setting on Thermocycler<br> | Used PFU setting on Thermocycler<br> | ||
| - | <b>Results:</b | + | <b>Results:</b> <br> |
| + | [[image:100915Assembly.jpg|100px]] | ||
<b>Conclusion:</b> Assembly did not work as intended. | <b>Conclusion:</b> Assembly did not work as intended. | ||
| + | ---- | ||
| + | |||
| + | ===<font color="white">HB=== | ||
| + | |||
| + | <b>Objective:</b> PCR mms6 to attach fusion standard to the N-term and standard suffix to C-term.<br> | ||
| + | |||
| + | <table border ="3"> | ||
| + | <tr><td><b>Component</b><td><b>1X(µL)</b><td><b>Master Mix(x6)(µL)</b> | ||
| + | <tr><td>Milli-Q H<sub>2</sub>O<td>12.8<td>76.8 | ||
| + | <tr><td>10x Pfu Buffer with MgSO<sub>4</sub><td>2<td>12 | ||
| + | <tr><td>dNTPs<td>1<td>6 | ||
| + | <tr><td>Fusion Forward Primer<td>1<td>6 | ||
| + | <tr><td>Standard Reverse Primer<td>1<td>6 | ||
| + | <tr><td>Template DNA<td>2<td>12 | ||
| + | <tr><td>Pfu polymerase<td>0.2<td>1.2 | ||
| + | </table><br> | ||
| + | |||
| + | Prefix fusion sense primer has melting temperature of 56.1<sup>o</sup>C. Standard suffix primer has melting temperature of 61.3<sup>o</sup>C. | ||
| + | |||
| + | PCR- Conditions: | ||
| + | 1. 95<sup>o</sup>C for 180 sec | ||
| + | 2. 95<sup>o</sup>C for 30 sec | ||
| + | 3. 51.1<sup>o</sup>C for 30 sec | ||
| + | 4. 72<sup>o</sup>C for 150 sec | ||
| + | 5. 72<sup>o</sup>C for 600 sec | ||
| + | (35 cycles) | ||
| + | |||
| + | Program HBPREFU: | ||
| + | 1. 49.1<sup>o</sup>C 2. 50.2<sup>o</sup>C 3. 51.4<sup>o</sup>C 4. 52.6<sup>o</sup>C 5. 53.5<sup>o</sup>C | ||
==<font color="white">September 21, 2010== | ==<font color="white">September 21, 2010== | ||
| Line 485: | Line 636: | ||
<b>Follow-up:</b><br> | <b>Follow-up:</b><br> | ||
Screen white colonies by addition of catechol to solution containing white cells. | Screen white colonies by addition of catechol to solution containing white cells. | ||
| + | ---- | ||
| + | ===<font color="white">TF, MC=== | ||
| + | |||
| + | <b>Objective:</b> Obtain a preparation of B0015 (dT). | ||
| + | |||
| + | <b>Method:</b> Used [[Team:Lethbridge/Notebook/Protocols|Plasmid DNA Purification by Alkaline Lysis (Large Scale AKA Maxiprep]] protocol. | ||
| + | ---- | ||
| + | ===<font color="white">AV=== | ||
| + | <b>Objective:</b> PCR amplify the signal sequences synthesized by Mr. Gene (K331007, K331008, K331009, K331012).<br> | ||
| + | |||
| + | Composition of each PCR tube: | ||
| + | <table><table border ="3"> | ||
| + | <tr><td><b>Component</b></td><td><b>Volume(µL)</b></td></tr> | ||
| + | <tr><td>10x Pfu Buffer w/ MgSO<sub>4</sub></td><td>2</td></tr> | ||
| + | <tr><td>dNTP (10mM)</td><td>1</td></tr> | ||
| + | <tr><td>VF2</td><td>1</td></tr> | ||
| + | <tr><td>VR</td><td>1</td></tr> | ||
| + | <tr><td>MilliQ H<sub>2</sub>O</td><td>12.8</td></tr> | ||
| + | <tr><td>Template DNA</td><td>2</td></tr> | ||
| + | <tr><td>Pfu DNA Polymerase</td><td>0.2</td></tr> | ||
| + | </table><br> | ||
| + | Used iGEM thermocycler setting: PFU.<br> | ||
| + | |||
==<font color="white">September 21, 2010== | ==<font color="white">September 21, 2010== | ||
===<font color="white">ADS=== | ===<font color="white">ADS=== | ||
| Line 509: | Line 683: | ||
<tr><td>16</td><td>Empty</td><td></td><td></td></tr> | <tr><td>16</td><td>Empty</td><td></td><td></td></tr> | ||
<tr><td>17</td><td>Empty</td><td></td><td></td></tr></table><br> | <tr><td>17</td><td>Empty</td><td></td><td></td></tr></table><br> | ||
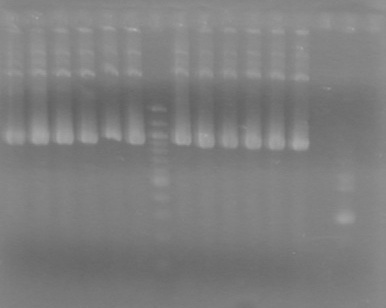
| - | <b>Results:</b | + | <b>Results:</b> <br> |
| + | [[image:100921.jpg|200px]] | ||
*Both KG PCR (Standard-Fusion and Fusion-Standard) amplified | *Both KG PCR (Standard-Fusion and Fusion-Standard) amplified | ||
*ADS PCR Amplified, but no insert present. | *ADS PCR Amplified, but no insert present. | ||
| Line 565: | Line 740: | ||
*4) Verify sequence | *4) Verify sequence | ||
*5) Submit to registry for sequencing | *5) Submit to registry for sequencing | ||
| + | ---- | ||
| + | ===<font color="white">TF, MC=== | ||
| + | |||
| + | <b>Objective:</b> Obtain a preparation of J04450 (RFP).<br> | ||
| + | |||
| + | <b>ethod:</b> Used [[Team:Lethbridge/Notebook/Protocols|Plasmid DNA Purification by Alkaline Lysis (Large Scale AKA Maxiprep]] protocol. | ||
==<font color="white">September 24, 2010== | ==<font color="white">September 24, 2010== | ||
| Line 639: | Line 820: | ||
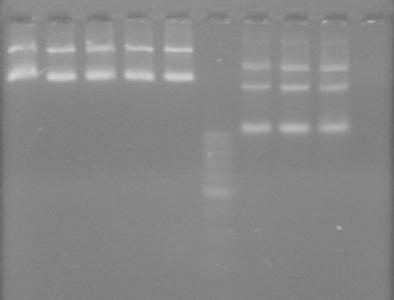
<b>Results:</b> Obtained <font color="red">RESULTS!!!!</font color> positive colonies.<br> | <b>Results:</b> Obtained <font color="red">RESULTS!!!!</font color> positive colonies.<br> | ||
Also analyzed minipreps on 1.5% TAE Agarose Gel<br> | Also analyzed minipreps on 1.5% TAE Agarose Gel<br> | ||
| - | + | [[image:100926Minipreps.jpg|200px]] | |
==<font color="white">September 29, 2010== | ==<font color="white">September 29, 2010== | ||
| Line 652: | Line 833: | ||
Obtained TNTC colonies<br> | Obtained TNTC colonies<br> | ||
Started overnight 5mL cultures for miniprep and insertion into pSB1C3 vector. | Started overnight 5mL cultures for miniprep and insertion into pSB1C3 vector. | ||
| + | |||
| + | ---- | ||
| + | ===<font color="white">AV=== | ||
| + | |||
| + | <b>Objective:</b> PCR the N-term fusion tag onto RFP (E1010) using sloppy annealing of the YFP/CFP fusion primers. <br><br> | ||
| + | |||
| + | Composition of each PCR tube: | ||
| + | <table><table border ="3"> | ||
| + | <tr><td><b>Component</b></td><td>Volume(µL)</td></tr> | ||
| + | <tr><td>10x Pfu Buffer w/ MgSO<sub>4</sub></td><td>2</td></tr> | ||
| + | <tr><td>dNTP (10mM)</td><td>1</td></tr> | ||
| + | <tr><td>N-term Fus Prefix</td><td>1</td></tr> | ||
| + | <tr><td>Biobrick Suffix</td><td>1</td></tr> | ||
| + | <tr><td>MilliQ H<sub>2</sub>O</td><td>12.8</td></tr> | ||
| + | <tr><td>Template DNA (J04450)</td><td>2</td></tr> | ||
| + | <tr><td>Pfu DNA Polymerase</td><td>0.2</td></tr> | ||
| + | </table><br> | ||
| + | |||
| + | PCR conditions: | ||
| + | <table><table border ="3"> | ||
| + | <tr><td><b>Step</b></td><td><b>Temperature (<sup>o</sup>C)</b></td><td><b>Time (mins)</b></td><td><b>Number of cycles</b></td></tr> | ||
| + | <tr><td>Initial Denaturation</td><td>95</td><td>2</td><td>1</td></tr> | ||
| + | <tr><td>Denaturation</td><td>95</td><td>0.5</td><td>25</td></tr> | ||
| + | <tr><td>Annealing</td><td>47.6, 50.5, 53.8 (gradient)</td><td>0.5</td><td>25</td></tr> | ||
| + | <tr><td>Extension</td><td>72</td><td>2</td><td>25</td></tr> | ||
| + | <tr><td>Final Extension</td><td>72</td><td>10</td><td>1</td></tr> | ||
| + | </table><br> | ||
| + | |||
| + | Results: Gel has not been run. Have decided to order RFP specific primers for next year.<br><br> | ||
| + | <br> | ||
 "
"