Team:Kyoto/Notebook
From 2010.igem.org
(→Digestion of the PCR products of S gene by EcoRI and SpeI and E0840 by EcoRI and XbaI to insert S gene to E0840) |
(→Transformation) |
||
| Line 38: | Line 38: | ||
|<partinfo>J06702</partinfo>||2-8-E||1||20||21||○ | |<partinfo>J06702</partinfo>||2-8-E||1||20||21||○ | ||
|- | |- | ||
| - | |<partinfo>pSB4K5</partinfo>||1-5-G||1||20||21||&# | + | |<partinfo>pSB4K5</partinfo>||1-5-G||1||20||21||× |
|- | |- | ||
| - | |<partinfo>B0015</partinfo>||1-23-L||1||20||21||LB (Kan+)||&# | + | |<partinfo>B0015</partinfo>||1-23-L||1||20||21||LB (Kan+)||× |
|} | |} | ||
A vector of "pSB4K5" is Kanamycin-resistance, however, we plated it to LB plate (Amp+). And We didn't pre-culture "B0015" despite its vector is Kanamycin-resistance. So, it was predicted that we will fail the transformation of "pSB4K5" and "B0015". | A vector of "pSB4K5" is Kanamycin-resistance, however, we plated it to LB plate (Amp+). And We didn't pre-culture "B0015" despite its vector is Kanamycin-resistance. So, it was predicted that we will fail the transformation of "pSB4K5" and "B0015". | ||
</div> | </div> | ||
| + | |||
===Wednesday, July 21 <span class="by">By: Wataru, Ken, Makoto, Takuya Y.</span>=== | ===Wednesday, July 21 <span class="by">By: Wataru, Ken, Makoto, Takuya Y.</span>=== | ||
<div class="culture lysis measure"> | <div class="culture lysis measure"> | ||
Revision as of 13:50, 30 September 2010
Index
Notebook
Tuesday, July 20 By: Wataru, Tomo, Yuki, Kazuya, Ken, Makoto
Solubilization of Antibiotics
| Ampicillin(Amp) | Kanamycin(Kan) | |
| Mix 1.0g Amp and 20ml MilliQ | Mix 0.5g Kan and 10ml MilliQ | Final concentration is 50mg/ml |
| Dispense 1.1ml of the solution into 1.5ml tubes | ||
| Store in the freezer (-20℃) | ||
Making plates for LB (Amp+) and LB (Kan+)
Transformation
| Name | Well | Sample (µl) | Competent Cells (µl) | Total (µl) | Plate | Incubation | Result |
|---|---|---|---|---|---|---|---|
| <partinfo>J23100</partinfo> | 1-18-C | 1 | 20 | 21 | LB (Amp+) | At 37℃, 7/20 20:50 - 7/21 17:00 | ○ |
| <partinfo>J23105</partinfo> | 1-18-M | 1 | 20 | 21 | ○ | ||
| <partinfo>J23116</partinfo> | 1-20-M | 1 | 20 | 21 | ○ | ||
| <partinfo>R0011</partinfo> | 1-6-G | 1 | 20 | 21 | ○ | ||
| <partinfo>E0840</partinfo> | 1-12-O | 1 | 20 | 21 | ○ | ||
| <partinfo>J06702</partinfo> | 2-8-E | 1 | 20 | 21 | ○ | ||
| <partinfo>pSB4K5</partinfo> | 1-5-G | 1 | 20 | 21 | × | ||
| <partinfo>B0015</partinfo> | 1-23-L | 1 | 20 | 21 | LB (Kan+) | × |
A vector of "pSB4K5" is Kanamycin-resistance, however, we plated it to LB plate (Amp+). And We didn't pre-culture "B0015" despite its vector is Kanamycin-resistance. So, it was predicted that we will fail the transformation of "pSB4K5" and "B0015".
Wednesday, July 21 By: Wataru, Ken, Makoto, Takuya Y.
Culture of plates in which colonies was observed at 37℃ from 07/21 20:50 to 07/22 17:00
Making a master plate of the above plates
Retry Transformation
| Name | Well | Sample (µl) | Competent Cells (µl) | Total (µl) | Plate | Incubation | Result |
|---|---|---|---|---|---|---|---|
| <partinfo>pSB4K5</partinfo> | 1-5-G | 1 | 20 | 21 | LB (Kan+) | At 37℃ 7/21 20:50 - 7/22 16:30 | ○ |
| <partinfo>B0015</partinfo> | 1-23-L | 1 | 20 | 21 | ○ |
PCR for S-R-Rz/Rz1 and S
| No. | Water | 25mM MgSO4 | 2mM dNTPs | 10xBuffer for KOD Plus ver.2 | TemplateDNA (5ng/µl) | Primer Forward (10µM) | Primer S-R-Rz/Rz1 Reverse (10µM) | Primer S Reverse (10µM) | KOD Plus ver.2 | Total |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 28µl | 3µl | 5µl | 5µl | 5µl | 1.5µl | 1.5µl | - | 1µl | 50µl |
| 2 | 28µl | 3µl | 5µl | 5µl | 5µl | 1.5µl | 1.5µl | - | 1µl | 50µl |
| 3 | 28µl | 3µl | 5µl | 5µl | 5µl | 1.5µl | - | 1.5µl | 1µl | 50µl |
| 4 | 28µl | 3µl | 5µl | 5µl | 5µl | 1.5µl | - | 1.5µl | 1µl | 50µl |
| 5 | 28µl | 3µl | 5µl | 5µl | 5µl | 1.5µl | 1.5µl | - | 1µl | 50µl |
| 6 | 28µl | 3µl | 5µl | 5µl | 5µl | 1.5µl | 1.5µl | - | 1µl | 50µl |
| 7 | 28µl | 3µl | 5µl | 5µl | 5µl | 1.5µl | - | 1.5µl | 1µl | 50µl |
| 8 | 28µl | 3µl | 5µl | 5µl | 5µl | 1.5µl | - | 1.5µl | 1µl | 50µl |
| 94℃ | 2min | |
| 98℃ | 10sec | 30 cycles |
| 55℃ | 30sec | |
| 68℃ | 4min | |
| 4℃ | forever |
Thursday, July 22 By: Wataru
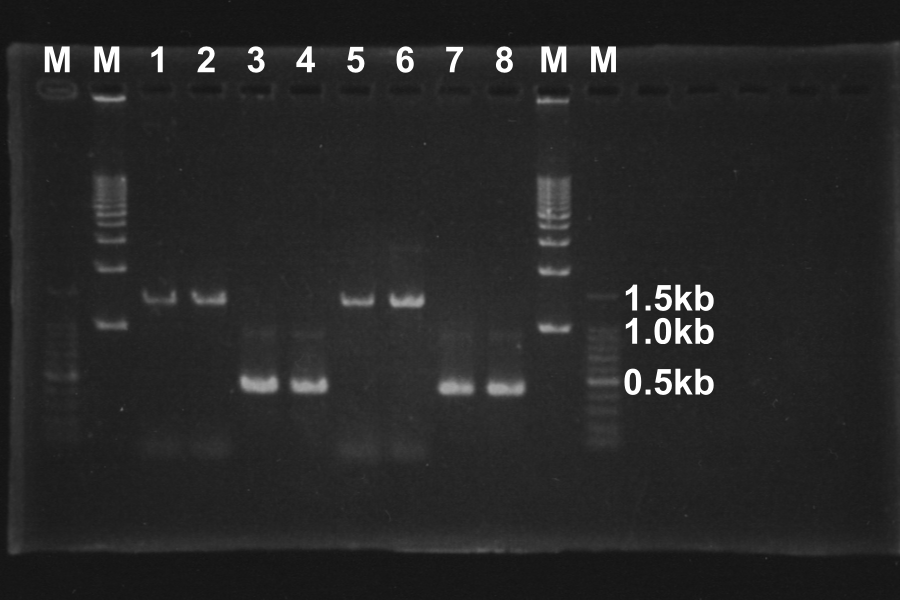
Electrophoresis of the PCR products for 40min
Length of S and S-R-Rz/Rz1 is 370bp and 1300bp, so PCR succeeded.
Miniprep
| Name | Concentration(ng/µl) |
|---|---|
| <partinfo>J23100</partinfo> | 18.5 |
| <partinfo>J23105</partinfo> | 12.5 |
| <partinfo>J23116</partinfo> | 14.6 |
| <partinfo>R0011</partinfo> | 8.6 |
| <partinfo>E0840</partinfo> | 12.1 |
| <partinfo>J06702</partinfo> | 14.7 |
The concentration of all samples was very week. Probably our shaking incubation was week.
Culture of plates and making master plates of <partinfo>pSB4K5</partinfo> and <partinfo>B0015</partinfo> from 07/22 17:00 to 07/23 10:00
Friday, July 23 By: Wataru, Tomo, Makoto
Miniprep
| Name | Concentration(ng/µl) |
|---|---|
| <partinfo>pSB4K5</partinfo> | 79.2 |
| <partinfo>B0015</partinfo> | - |
We lost <partinfo>B0015</partinfo> by our mistake. The concentration of <partinfo>pSB4K5</partinfo> is high, so this condition of shaking incubation is moderate.
Picking up number 1, 3, 5, and 7 of the products of PCR, and PCR-purification
| No. | Name | Concentration (ng/µl) | New Name |
|---|---|---|---|
| 1 | S-R-Rz/Rz1 | 18.6 | - |
| 3 | S | 77.6 | S1 |
| 5 | S-R-Rz/Rz1 | 33.6 | - |
| 7 | S | 65.4 | S2 |
The concentration of sample number 1 and 5, the PCR products of S-R-Rz/Rz1, is week, so we desided to retry PCR.
Retry of Standard PCR for S-R-Rz/Rz1
| No. | Water | 25mmol/l MgSO4 | 2mmol/l dNTPs | 10×Buffer for KOD plus ver.2 | Template DNA (5ng/µl) | Primer S-R-Rz/Rz1 Forward (10µmol/l) | Primer S-R-Rz/Rz1 Reverse (10µmol/l) | KOD plus ver.2 | Total |
|---|---|---|---|---|---|---|---|---|---|
| 1 | 28µl | 3 | 5 | 5 | 5 | 1.5 | 1.5 | 1 | 50 |
| 2 | 28 | 3 | 5 | 5 | 5 | 1.5 | 1.5 | 1 | 50 |
| 3 | 26.5 | 4.5 | 5 | 5 | 5 | 1.5 | 1.5 | 1 | 50 |
| 4 | 26.5 | 4.5 | 5 | 5 | 5 | 1.5 | 1.5 | 1 | 50 |
| 5 | 25 | 6 | 5 | 5 | 5 | 1.5 | 1.5 | 1 | 50 |
| 6 | 25 | 6 | 5 | 5 | 5 | 1.5 | 1.5 | 1 | 50 |
| 94℃ | 2min | |
| 98℃ | 10sec | 30 cycles |
| 55℃ | 30sec | |
| 68℃ | 4min | |
| 4℃ | forever |
Restriction Digestion of <partinfo>J06702</partinfo> by EcoRI, XbaI, SpeI, and PstI to check function of our Restriction Enzyme
| No. | 10xBuffer | BSA | Enzyme | MilliQ | Total | Incubation |
|---|---|---|---|---|---|---|
| 1 | 5µl | 1 | EcoRI 0.1 | 3.6 | 10 | At 37℃ 7/23 18:00 - 7/23 18:30 |
| 2 | 5 | 1 | XbaI 0.1 | 3.6 | 10 | |
| 3 | 5 | 1 | SpeI 0.1 | 3.6 | 10 | |
| 4 | 5 | 1 | PstI 0.1 | 3.6 | 10 | |
| 5 | 5 | 1 | - | 3.7 | 10 |
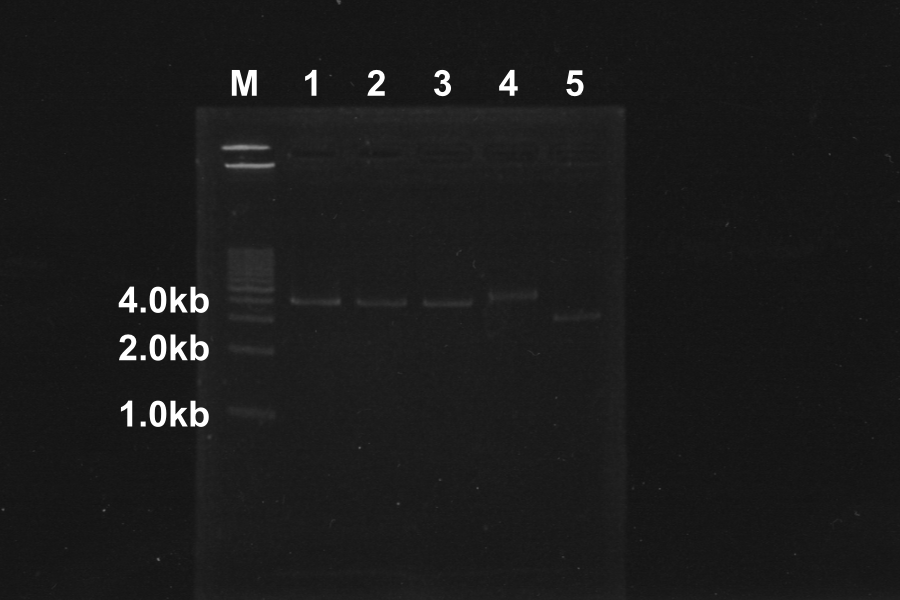
Electrophoresis of above sample for 35min
Comparison to sample 5(control, circular DNA), the bands of sample 1, 2, 3, 4 was shifted. The DNA of sample 1, 2, 3, 4 was linearized by Restriction enzymes. So, our restriction enzymes work correctly.
Digestion of the PCR products of S gene by EcoRI and SpeI and <partinfo>E0840</partinfo> by EcoRI and XbaI to insert S gene to <partinfo>E0840</partinfo>
| Name | Sample Volume (µl) | 10×Buffer | Enzyme 1 | Enzyme 2 | MilliQ | Total | Incubation |
|---|---|---|---|---|---|---|---|
| S1 | 11 | 5 | EcoRI 0.2 | SpeI 0.2 | 33.6 | 50 | At 37℃ for 2h |
| S2 | 11 | 5 | EcoRI 0.2 | SpeI 0.2 | 33.6 | 50 | |
| <partinfo>E0840</partinfo> | 45 | 5 | EcoRI 0.2 | XbaI 0.2 | 0 | 50 |
After PCR purification, evaporated them and diluted 3ul.
Ligation
| Name | Vector | Insert | Ligation High | Total |
|---|---|---|---|---|
| S-E08401 | <partinfo>E0840</partinfo> 0.5µl | S1 0.5 | 1 | 2 |
| S-E08402 | <partinfo>E0840</partinfo> 0.5 | S2 0.5 | 1 | 2 |
 "
"