Team:Washington/Gram Negative/Build
From 2010.igem.org
(→3OC6HSL as a proxy for a pathogenic signal) |
(→Engineering the Toxin Antitoxin HSL-inducible circuit) |
||
| (86 intermediate revisions not shown) | |||
| Line 26: | Line 26: | ||
<!---------------------------------------PAGE CONTENT GOES BELOW THIS----------------------------------------> | <!---------------------------------------PAGE CONTENT GOES BELOW THIS----------------------------------------> | ||
| - | |||
| - | = | + | =Recombineering the Promoter Region of the Type VI Secretion System= |
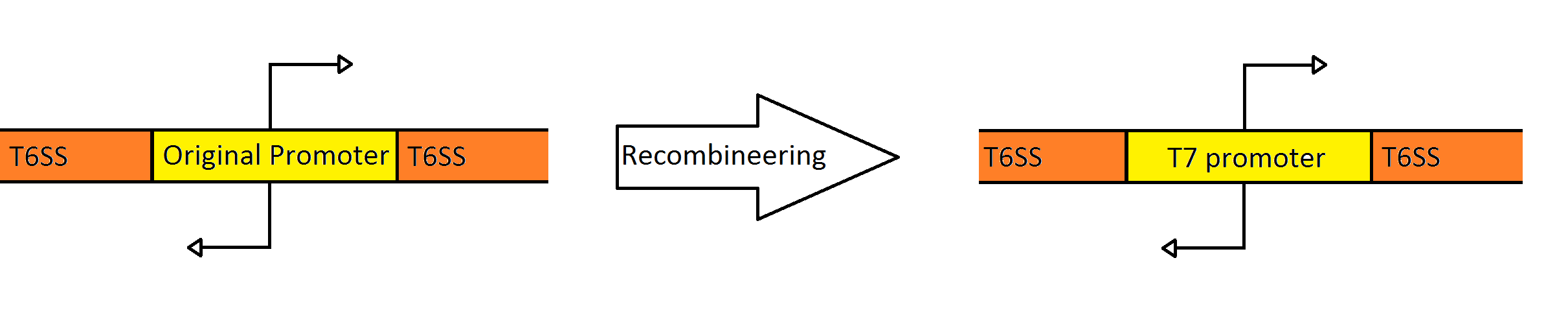
| - | + | [[Image:Washington_Recombineering_basic.png|750px|center|thumb|Schematic Showing the Engineering of the Fosmid to Contain a Divergent T7 Promoter Instead of the Native ''Pseudomonas'' Promoter]] | |
| - | + | Our goal was to replace the native ''P. aeruginosa'' regulation of the Type VI Secretion System with one that can be utilized by the ''E. coli'' transcriptional machinery. In order to replace the native promoters with more robust T7 promoters we used a technique called [https://2010.igem.org/Team:Washington/Protocols/Recombineering Recombineering] to flip out the region containing both promoters and replace it with a gene cassette as follows. We designed primers to create a cassette using PCR, containing the ''galK'' galactose metabolism gene flanked by 50 base pairs of homology just outside of the promoter region in the fosmid. The cassette was inserted, giving us a selection factor for our final insertion of our new promoters. A new promoter cassette was designed in total using oligos that encoded the two T7 promoters flanked by the same 50 base pairs of homology used previously, and inserted using recombineering into the fosmid that contained ''galK'' in the promoter region. | |
| - | + | ||
| + | =Engineering the Toxin Antitoxin HSL-inducible circuit= | ||
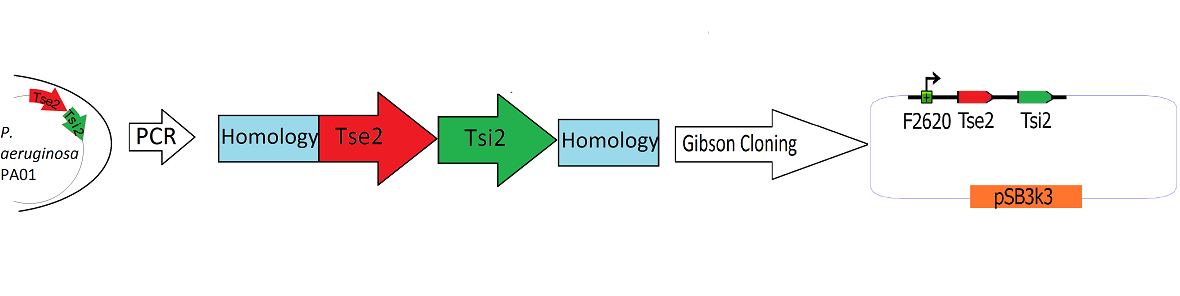
| + | [[Image:Washington_Building_Tse2_circuit.png|center|750px|thumb|Schematic Detailing the Transfer of Tse2 and Tsi2 from ''P. Aeruginosa'' into pSB3k3]] | ||
| + | The Tse2/Tsi2 locus was amplified from ''P. aeruginosa'' genomic DNA using PCR. Primers were used that amplified the region from the start codon of Tse2 to the stop codon of Tsi2. The Tse2 and Tsi2 locus was then placed downstream from F2620 via [[Team:Washington/Tools_Used/Next-Gen_Cloning|Gibson Cloning]]. Gibson cloning is a method that joins regions of homology found on the 5' and 3' ends of linearized molecules. The end result of this Gibson cloning reaction is a circularized plasmid containing the final F2620.Tse2.Tsi2 construct in pSB3K3. The circularized plasmid is then transformed into electrocompetent ''E. coli'' Dh5a and the resulting colonies were screened for the insert of expected length via double restriction digest followed by agarose gel electrophoresis. The plasmids that showed inserts of expected length were sequenced, and it was determined that multiple plasmids had correct F2620.Tse2.Tsi2 inserts. | ||
| + | |||
| + | |||
<!---------------------------------------PAGE CONTENT GOES ABOVE THIS----------------------------------------> | <!---------------------------------------PAGE CONTENT GOES ABOVE THIS----------------------------------------> | ||
<div style="text-align:center"> | <div style="text-align:center"> | ||
| - | '''← [[Team:Washington/ | + | '''← [[Team:Washington/Gram Negative/Design|Designing the Gram(-) Therapeutic]]''' |
| | ||
| | ||
| | ||
| - | '''[[Team:Washington/ | + | '''[[Team:Washington/Gram Negative/Test|Testing the Gram(-) Therapeutic]] →''' |
</div> | </div> | ||
{{Template:Team:Washington/Templates/Footer}} | {{Template:Team:Washington/Templates/Footer}} | ||
Latest revision as of 16:49, 26 October 2010
Recombineering the Promoter Region of the Type VI Secretion System
Our goal was to replace the native P. aeruginosa regulation of the Type VI Secretion System with one that can be utilized by the E. coli transcriptional machinery. In order to replace the native promoters with more robust T7 promoters we used a technique called Recombineering to flip out the region containing both promoters and replace it with a gene cassette as follows. We designed primers to create a cassette using PCR, containing the galK galactose metabolism gene flanked by 50 base pairs of homology just outside of the promoter region in the fosmid. The cassette was inserted, giving us a selection factor for our final insertion of our new promoters. A new promoter cassette was designed in total using oligos that encoded the two T7 promoters flanked by the same 50 base pairs of homology used previously, and inserted using recombineering into the fosmid that contained galK in the promoter region.
Engineering the Toxin Antitoxin HSL-inducible circuit
The Tse2/Tsi2 locus was amplified from P. aeruginosa genomic DNA using PCR. Primers were used that amplified the region from the start codon of Tse2 to the stop codon of Tsi2. The Tse2 and Tsi2 locus was then placed downstream from F2620 via Gibson Cloning. Gibson cloning is a method that joins regions of homology found on the 5' and 3' ends of linearized molecules. The end result of this Gibson cloning reaction is a circularized plasmid containing the final F2620.Tse2.Tsi2 construct in pSB3K3. The circularized plasmid is then transformed into electrocompetent E. coli Dh5a and the resulting colonies were screened for the insert of expected length via double restriction digest followed by agarose gel electrophoresis. The plasmids that showed inserts of expected length were sequenced, and it was determined that multiple plasmids had correct F2620.Tse2.Tsi2 inserts.
 "
"