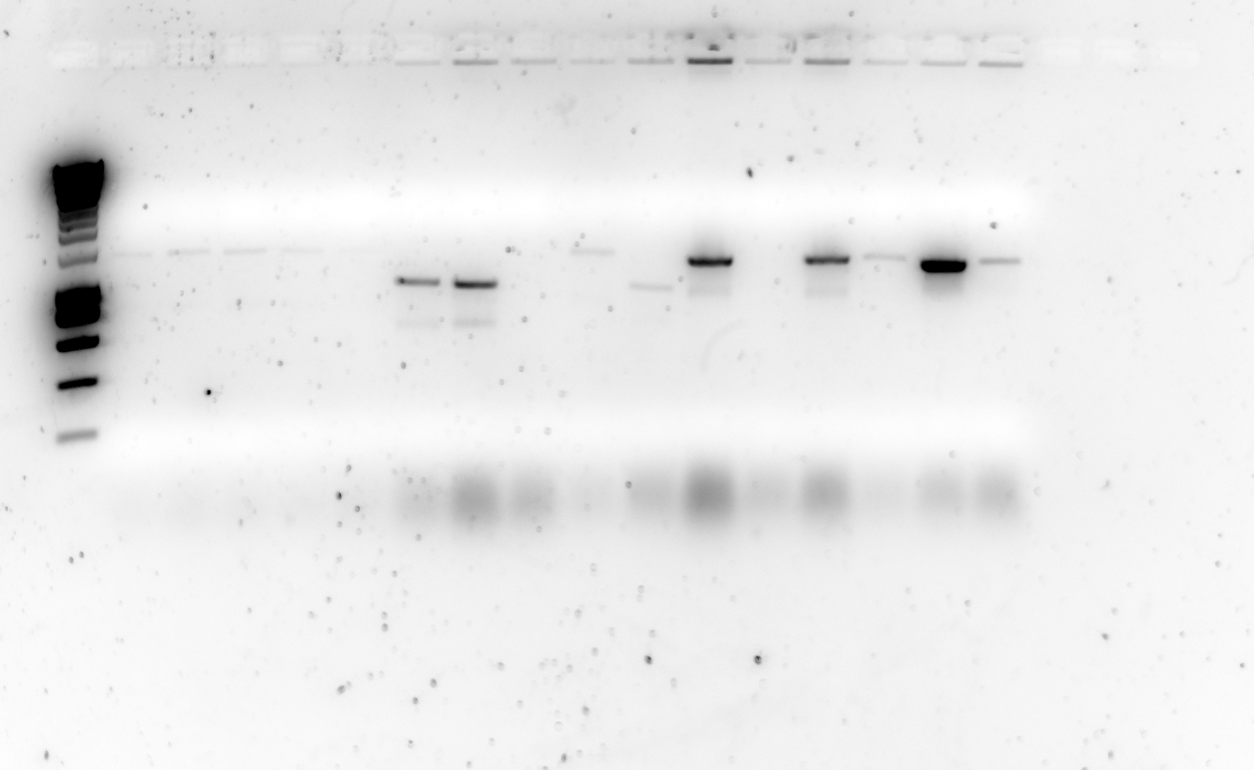Team:TU Delft/14 July 2010 content
From 2010.igem.org
Contents |
Lab work
Ordered DNA
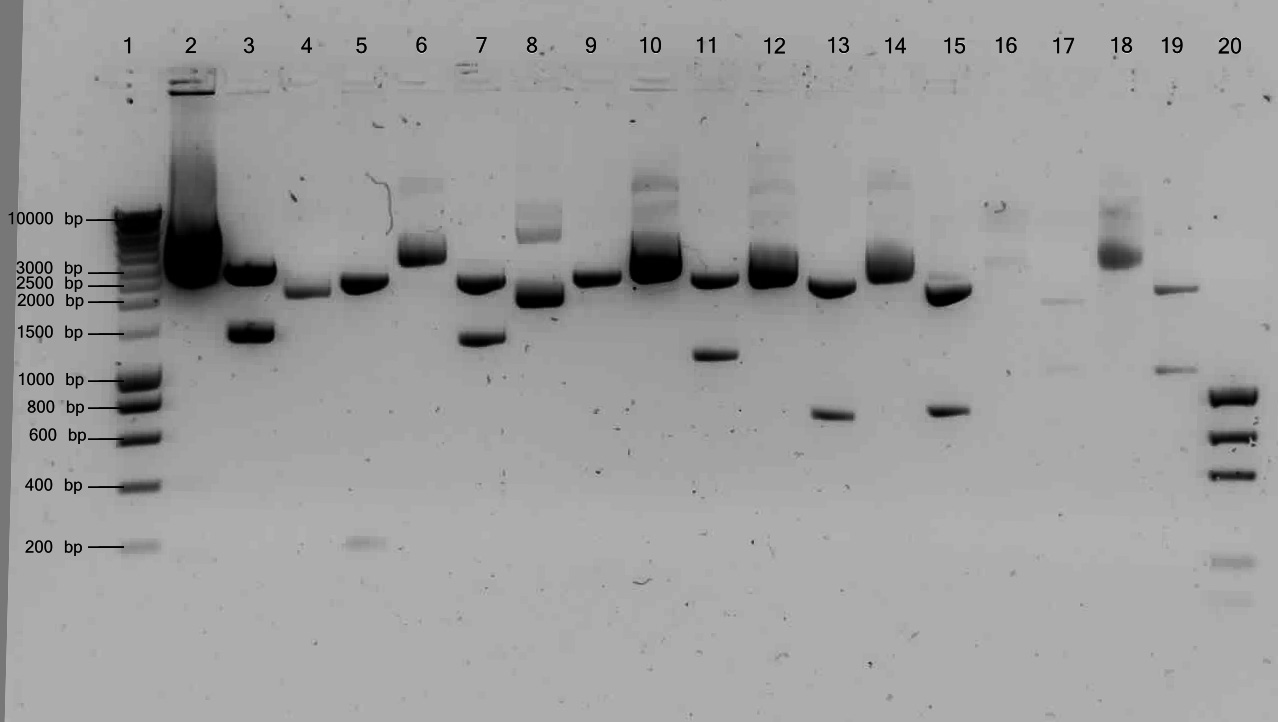
Yesterdays digestions were tested on gel.
Lane description:
| # | Description | Expected Length (bp) | Status |
| 1 | Smartladder | n/a | n/a |
| 2 | Undigested ladA | 4265 | ✓ |
| 3 | ladA + EcoR1 + PstI | 1365, 2900 | ✓ |
| 4 | Undigested RubA3 | 2607 | ✓ |
| 5 | RubA3 + EcoR1 + PstI | 213, 2394 | ✓ |
| 6 | Undigested AlkB2 | 3666 | ✓ |
| 7 | AlkB2 + EcoR1 + PstI | ✓ | |
| 8 | Undigested PalkS | 2516 | ✓ |
| 9 | PalkS + EcoR1 + PstI | 124, 2392 | ✓ |
| 10 | Undigested AlnA | 3469 | ✓ |
| 11 | AlnA + EcoR1 + PstI | 1089, 2380 | ✓ |
| 12 | Undigested OprG | 3122 | ✓ |
| 13 | OprG + EcoR1 + PstI | 726, 2396 | ✓ |
| 14 | Undigested ADH | 3180 | ✓ |
| 15 | ADH + EcoR1 + PstI | 784, 2396 | ✓ |
| 16 | Undigested pSB1C3 | ✓ | |
| 17 | pSB1C3 + EcoR1 + PstI | ✓ | |
| 18 | Undigested pSB1T3 | ✓ | |
| 19 | pSB1T3 + EcoR1 + PstI | ✓ |
At the same time the digestion products were ligated at 16 °C:
| # | BioBrick | Fragment | Recipient vector |
| 1 | K398001 | 8 μL ‘E-alkB2-P’ | 1 μL ‘E–linear pSB1C3–P’ |
| 2 | K398002 | 8 μL ‘E-rubA3-P’ | 1 μL ‘E–linear pSB1C3–P’ |
| 3 | K398000 | 8 μL ‘E-ladA-P’ | 1 μL ‘E–linear pSB1C3–P’ |
| 4 | K398005 | 8 μL ‘E-ADH-P’ | 1 μL ‘E–linear pSB1C3–P’ |
| 5 | K398200 | 8 μL ‘E-AlnA-P’ | 1 μL ‘E–linear pSB1C3–P’ |
| 6 | K398201 | 8 μL ‘E-OprG-P’ | 1 μL ‘E–linear pSB1C3–P’ |
| 7 | K398300 | 8 μL ‘E-AlkS-P’ | 1 μL ‘E–linear pSB1C3–P’ |
| 8 | K398301 | 8 μL ‘E-PalkB-P’ | 1 μL ‘E–linear pSB1C3–P’ |
| 9 | K398303 | 8 μL ‘E-PalkS12-P’ | 1 μL ‘E–linear pSB1C3–P’ |
| 10 | K398400 | 8 μL ‘E-PhPFDα-P’ | 1 μL ‘E–linear pSB1C3–P’ |
| 11 | K398401 | 8 μL ‘E-PhPFDβ-P’ | 1 μL ‘E–linear pSB1C3–P’ |
| 12 | negative control | - | 1 μL pSB1C3 |
After 4 hour the 10 μL of the ligation mixes were transformed in Top10 competent cells. The rest of the ligation mix was overnight incubated at 16 °C.
Solvent Tolerance and Hydrocarbon Sensing
Yesterdays digestions were tested on gel.
Lane description:
| # | Description | Expected Length (bp) | Status |
| 1 | Smartladder | n/a | n/a |
| 2 | Undigested AlkS | 4999 | ✓ |
| 3 | AlkS + EcoR1 + SpeI | 2673, 2326 | ✓ |
| 4 | AlkS + EcoR1 + PstI | 2691, 1618, 690 | ✓ |
| 5 | Undigested PalkB | 2516 | ✓ |
| 6 | PalkB + EcoR1 + SpeI | 2410, 106 | ✓ |
| 7 | PalkB + EcoR1 + PstI | 2392, 124 | ✓ |
| 8 | Undigested PhPFDα | 2890 | ✓ |
| 9 | PhPFDα + EcoR1 + SpeI | 2410, 480 | ✓ |
| 10 | PhPFDα + EcoR1 + PstI | 2392, 498 | ✓ |
| 11 | Undigested PhPFDβ | 2790 | ✓ |
| 12 | PhPFDβ + EcoR1 + SpeI | 2412, 378 | ✓ |
| 13 | PhPFDβ + EcoR1 + PstI | 2394, 396 | ✓ |
| 14 | Undigested B0015 | ✓ | |
| 15 | B0015 + XbaI + PstI | ✓ | |
| 16 | Undigested B0032 | ✓ | |
| 17 | B0032 + XbaI + PstI | ✓ | |
| 18 | Undigested E0422 | ✓ | |
| 19 | E0422 + EcoR1 + PstI | ✓ |
At the same time the digestion products were ligated at 16 °C:
| # | BioBrick | Fragment 1 | Fragment 2 | Recipient vector |
| 1 | K398309 | 8 μL ‘E-AlkS-S’ | 8 μL ‘X-E0422–P’ | 1 μL ‘E-linear pSB1T3–P’ |
| 2 | 8 μL ‘E-AlkS-S’ | 8 μL ‘X-B0015–P’ | 1 μL ‘E-linear pSB1T3–P’ | |
| 3 | 8 μL PhPFDα + | 8 μL ‘X-B0032–P’ | 1 μL ‘E-linear pSB1T3–P’ | |
| 4 | 8 μL PhPFDβ + | 8 μL ‘X-B0015–P’ | 1 μL ‘E-linear pSB1T3–P’ | |
| 5 | negative control | - | 8 μL ‘X-B0015–P’ | 1 μL ‘E-linear pSB1T3–P’ |
| 6 | negative control | - | - | 1 μL ‘E-linear pSB1T3–P’ |
After 4 hour the 10 μL of the ligation mixes were transformed in Top10 competent cells. The rest of the ligation mix was overnight incubated at 16 °C.
Emulsifier
The transformations of yesterday containing the different ligation reactions gave colonies. All plates contained red and white colonies. This is probably due to the fact that we did not purified the backbone before ligation. So some plasmids reassembled. Pieter picked 5 colonies of every plate (no red ones) and 1 from the control plate (a red colony) and performed a colony PCR to check which colonies contained the right insert. At the same time he cultivated the colonies in 5 mL LB medium containing 15 μg/mL tetracycline to grow over night at 37 °C.
The PCR products were run on gel a 1% agarose gel.
Lane Description
| # | Description | Expected lenght (bp) | Primers | Status | Remarks |
| M1 | SmartLadder | n/a | n/a | n/a | |
| 1 | Transformant #1 of Ligation mix: R0011 + B0032 + pSB1T3 | 392 | G00100 + G00101 | ✗ | On gel there are only bands poorly visible at height of 1500 bp |
| 2 | Transformant #2 of Ligation mix: R0011 + B0032 + pSB1T3 | 392 | G00100 + G00101 | ✗ | |
| 3 | Transformant #3 of Ligation mix: R0011 + B0032 + pSB1T3 | 392 | G00100 + G00101 | ✗ | |
| 4 | Transformant #4 of Ligation mix: R0011 + B0032 + pSB1T3 | 392 | G00100 + G00101 | ✗ | |
| 5 | Transformant #5 of Ligation mix: R0011 + B0032 + pSB1T3 | 392 | G00100 + G00101 | ✗ | |
| 6 | Transformant #6 of Ligation mix: OprG + B0015 + pSB1T3 | 1138 | G00100 + G00101 | ✓ | The bands just a little lower are unidentified |
| 7 | Transformant #7 of Ligation mix: OprG + B0015 + pSB1T3 | 1138 | G00100 + G00101 | ✓ | |
| 8 | Transformant #8 of Ligation mix: OprG + B0015 + pSB1T3 | 1138 | G00100 + G00101 | ✗ | |
| 9 | Transformant #9 of Ligation mix: OprG + B0015 + pSB1T3 | 1138 | G00100 + G00101 | ✗ | |
| 10 | Transformant #10 of Ligation mix: OprG + B0015 + pSB1T3 | 1138 | G00100 + G00101 | ✓ | |
| 11 | Transformant #11 of Ligation mix: AlnA + B0015 + pSB1T3 | 1501 | G00100 + G00101 | ? | The bands of AlnA + B0015 and J04450 of the control are about the same size, it is not clear whether this transformation succeeded. Tomorrow I will check the isolated plasmids on gel with a characteristic digesition reaction. |
| 12 | Transformant #12 of Ligation mix: AlnA + B0015 + pSB1T3 | 1501 | G00100 + G00101 | ✗ | |
| 13 | Transformant #13 of Ligation mix: AlnA + B0015 + pSB1T3 | 1501 | G00100 + G00101 | ? | |
| 14 | Transformant #14 of Ligation mix: AlnA + B0015 + pSB1T3 | 1501 | G00100 + G00101 | ? | |
| 15 | Transformant #15 of Ligation mix: AlnA + B0015 + pSB1T3 | 1501 | G00100 + G00101 | ? | |
| 16 | Transformant #16 of Ligation mix: pSB1T3 | 1360 | G00100 + G00101 | ✓ |
Characterization of Anderson RBS sequences
Fluorescence measurements
The first results are in from last night's measurements! The results showed unexpected OD600 curves which were attributed to excessive evaporation from the wells.
Time for attempt #2 (with evaporation restrictions): The TOP 10 strains carrying K398500, K398501, K398502, K398503, K398504 and I13401 were cultured over night in 3 mL LB + 100 μg/mL ampicillin medium (37 ℃, 160 rpm).
Assembly of reference construct
In order to properly characterize the Anderson RBS sequences a 'reference' RBS will be run in parallel with the fluorescence measurements of the Anderson RBS sequences. Well characterized and standardized RBSs are the community RBSs, especially B0030, B0032 and B0034. A useful construct which could be used as the reference in these experiments has the form of: J23100 - B0030 or B0032 or B0034 - GFP - dT. As these constructs are (unfortunately) not readily available from the Spring 2010 distribution plates, they will have to be created:
1. The simplest method involves PCR amplifying K081005, digesting with EcoRI and SpeI and ligation into a pre-cut (EcoRI and XbaI) I13401 in pSB1A2 (which we have in stock).
2. The second method involves the transformation of K081005 into chemically competent Top10 cells, overnight culturing and plasmid isolation. The obtained K081005 in pSB1A2 can be cut by SpeI and PstI and ligated to the PCR amplified and digested (XbaI and PstI) I13401. This is analogous to the method used for obtaining the Anderson RBS constructs K398500-K398504.
3. The third method involves transforming J23100 into chemically competent Top10 cells, overnight culturing and plasmid isolation. The obtained J23100 in J61002 can be cut by SpeI and PstI and ligated to the PCR amplified and digested (XbaI and PstI) E0240.
4. The fourth method is a variation on method 3, where part E0240 is not PCR amplified, but directly cut out of the plasmid and ligated into S-J23100-pSB1A2-P. While this is not the most elegant method, screening for GFP fluorescent colonies will be enough to obtain the positive colony.
Thus, BioBricks K081005 and J23100 were transformed into chemically competent Top10 cells and plated out over ampicillin plates. K081005 was PCR amplified.
Assembly of positive control
In order to also have a working and strong positive control for GFP fluorescence the BioBrick I13522 was transformed into chemically competent Top10 cells and plated out over ampicillin plates.
 "
"