Team:WashU/Notebook/Biobricks
From 2010.igem.org
(New page: {{WashUHeader}} ==Week of 7/26== ===7/12=== Ordered the following primers to biobrick parts p18 Forward KanMX4 ATT CTT AGA ATT CGC GGC CGC TTC TAG CCC GCC GCC ACC ATG GGT AAG GAA AAG AC...)
Newer edit →
Revision as of 17:37, 4 October 2010

Week of 7/26
7/12
Ordered the following primers to biobrick parts
p18 Forward KanMX4 ATT CTT AGA ATT CGC GGC CGC TTC TAG CCC GCC GCC ACC ATG GGT AAG GAA AAG ACT CAC GTT TCG p19 Reverse KanMX4 GCC GCC CTC TGC AGC GGC CGC TAC TAG TAT TAG AAA AAC TCA TCG AGC ATC AAA TGA AAC TG p20 Forward NatMX4 GTT TCT TCG AAT TCG CGG CCG CTT CTA GCC CGC CGC CAC CAT GGG TAC CAC TCT TGA CGA CAC G p21 Reverse NatMX4 GTT TCT TCC TGC AGC GGC CGC TAC TAG TAT TAG GGG CAG GGC ATG CTC ATG TAG p22 Forward Ura3-1 GCACAGAACAATAACCTGCTGGAAACGAAGATAAATCgaagacGATTACTTCGCGTTATGCAGGC p23 Reverse Ura3-1 GCATCTTCTCAAATATGCTTCCCAGCCTGCTTATCcttctgAAATTCTGCCTCGTGATACGCC p24 Forward Ura3-2 tactagtagcggccgctgcagTCTTAACCCAACTGCACAGAACAATAACCTGCTGGAAACG p25 Reverse Ura3-2 ctctagaagcggccgcgaattcTTAGTATTGCTGGCCGCATCTTCTCAAATATGCTTCCCAGCC p26 Forward His3-1 GAACAGGCCACACAATCGCAAGTGATTAACgaagacGATTACTTCGCGTTATGCAGGC p27 Reverse His3-1 CCTTGAACGCACTCTCACTACGGTGATGATCActtctgAAATTCTGCCTCGTGATACGCC p28 Forward His3-2 tactagtagcggccgctgcagAGAGGGAGAAGCAGTAGCAGAACAGGCCACACAATCGCAAG p29 Reverse His3-2 ctctagaagcggccgcgaattcTTATGGCAACCGCAAGAGCCTTGAACGCACTCTCACTACGG p30 Biobrick Forward tgccacctgacgtctaagaa p31 Biobrick Reverse attaccgcctttgagtgagc p32 Forward Ura3 Check CGG TAA TCT CCG AGC AGA AGG AAG AAC G p33 Reverse Ura3 Check CAT TAC GAC CGA GAT TCC CGG GTA ATA ACT G p34 Forward His3 Check GAG CAG AAA GCC CTA GTA AAG CGT ATT ACA AAT G p35 Reverse His3 Check CTA CAT AAG AAC ACC TTT GGT GGA GGG AAC p36 Forward SxL gtttcttcgaattcgcggccgcttctagagcccgccgccaccatgtacg p37 Reverse SxL gtttcttcctgcagcggccgctactagtattattataccttgcgctttttcttgggg p38 Vector internal Sequencing 1 - Forward 1400 start ACA CCC GTC CTG TGG ATC p39 Vector internal Sequencing 2 - Reverse 1523 start GAAGTGGCGAGCCCGATCTTC p40 Vector internal Sequencing 3 2301 start CCACCTCGACCTAACTCGAGTTAC
7/21
The following PCr reactions were run and then column Purified.
Number Product Template Forward Primer Reverse Primer Temperature 1 Kan BB KanMX4 p18 p19 58 2 Nat BB NatMX4 p20 p21 58 3 Kan BB KanMX4 p18 p19 60 4 Nat BB NatMX4 p20 p21 60 5 Kan BB KanMX4 p18 p19 64 6 Nat BB NatMX4 p20 p21 64
The following digestions of were run.
K58 N58 K60 N60 K64 N64 P1 H20 12.5 12.5 12.5 12.5 12.5 12.5 22.5 Buffer 2 5 5 5 5 5 5 5 BSA 0.5 0.5 0.5 0.5 0.5 0.5 0.5 DNA 30 30 30 30 30 30 20 EcoRI 1 1 1 1 1 1 1 PstI 1 1 1 1 1 1 1
The digestions were run for 6 hours at 27 degC and then kept at 4 degC o/n
Made Chloramphenicol Plates at a concentration of 34μg/ml
Week of 7/19
Week of 8/2
Miniprepped the following from overnight cultures made on 8/21/10:
DNA Concentration (ng/μl) 260/280 20/230 Kan BB in pSB1C3 37.8 1.92 2.15 pSB1AT3 with BBa_J04450 insert 102.1 1.92 2.26 pSB1C3 with BBa_J04450 insert 196.8 1.91 2.31
The following PCR reaction was run with an elongation time of 3.5 minutes:
Number Product Template Forward Primer Reverse Primer Annealing Temperature 1 His3 Intermediate Vector pSB1AT3 w/ insert p27 p29 58 2 Ura3 intermediate Vector pSB1AT3 w/ insert p23 p25 58 3 His3 intermediate Vector pSB1AT3 w/ insert p27 p29 60 4 Ura3 intermediate Vector pSB1AT3 w/ insert p23 p25 60 5 His3 intermediate Vector pSB1AT3 w/ insert p27 p29 62 6 Ura3 intermediate Vector pSB1AT3 w/ insert p23 p25 62
The following Digestion was run:
pSB1C3 w/ insert H20 40 DNA 2.5 NEB-3 5 BSA 0.5 EcoRI 1 PstI 1
The following Gel was run:
1 1kb Ladder 2 H58 intermediate 3 U58 intermediate 4 H60 intermediate 5 U60 intermediate 6 H62 intermediate 7 U62 intermediate 8 pSB1C3 Digestion product
The gel was cut and the products were gel purified using the Sigma Kit
The following PCR reaction was let run over night with an elongation time of 3.5 minutes:
Number Product Template Forward Primer Reverse Primer Annealing Temperature 1 His3 Final Vector H58 Intermediate Vector p27 p29 58 2 Ura3 Final Vector U58 Intermediate Vector p23 p25 58 3 His3 Final Vector H60 Intermediate Vector p27 p29 60 4 Ura3 Final Vector U60 Intermediate Vector p23 p25 60 5 His3 Final Vector H62 Intermediate Vector p27 p29 62 6 Ura3 Final Vector U62 Intermediate Vector p23 p25 62
7/23
Nanodropped Over night PCR product:
Number Label Concentration ng/μL 260/280 260/230 1 H58 10.2 2.34 1.22 2 U58 6.6 2.31 1.13 3 H60 10.2 1.81 1.14 4 U60 5.5 2.33 1.16 5 H62 7.4 2.00 1.18 6 H62 0.4 -0.65 0.17
Preformed the following digestions at 37 degC for 2 hours and heat inactivation at 80 degC for 20 minutes:
H58 U58 H60 U60 H62 pSB1C3 w/ insert ADH1 Promoter Kan_BB Linear pSB1T3 pSB1AT3 w/ insert H20 12.5 12.5 12.5 12.5 12.5 40 40 29.3 32.5 335.6 DNA 30 30 30 30 30 2.5 2.5 13.2 10 4.9 NEB-2 - - - - - - 5 5 5 5 NEB-3 5 5 5 5 5 5 - - - - BSA 0.5 0.5 0.5 0.5 0.5 0.5 0.5 0.5 0.5 0.5 1μL E1 EcoRI EcoRI EcoRI EcoRI EcoRI EcoRI EcoRI XbaI EcoRI EcoRI 1μL E2 PstI PstI PstI PstI PstI PstI SpeI PstI PstI PstI 1μL E3 XbaI 1μL E4 SpeI
Preformed the following ligations following the Biobrick Manual Protocol
H58 U58 H60 U60 H62 AHD1+K_BB+pSB1T3 ADH1+K_BB+pSB1AT3 H20 11 11 11 11 11 11 11 Component 1 H58 U58 H60 U60 H62 ADH1 ADH1 Component 2 pSB1C3 pSB1C3 pSB1C3 pSB1C3 pSB1C3 K_BB K_BB Component 3 Linear pSB1T3 pSB1AT3 10x T4 ligase buffer 2 2 2 2 2 2 2 T4 DNA Ligase 1 1 1 1 1 1 1 1
Preformed the following transformations:
1 H58 Amp 2 U58 Amp 3 H60 Amp 4 U60 Amp 5 H62 Amp 6 N58+pSB1C3 Chlor 7 N60+pSB1C3 Chlor 8 N64+pSB1C3 Chlor 9 ADH1+Kan_BB+pSB1T3 Tet 10 ADH1+Kan_BB+pSB1AT3 Amp 11 Tet - Control Tet 12 Chlor - Control Chlor
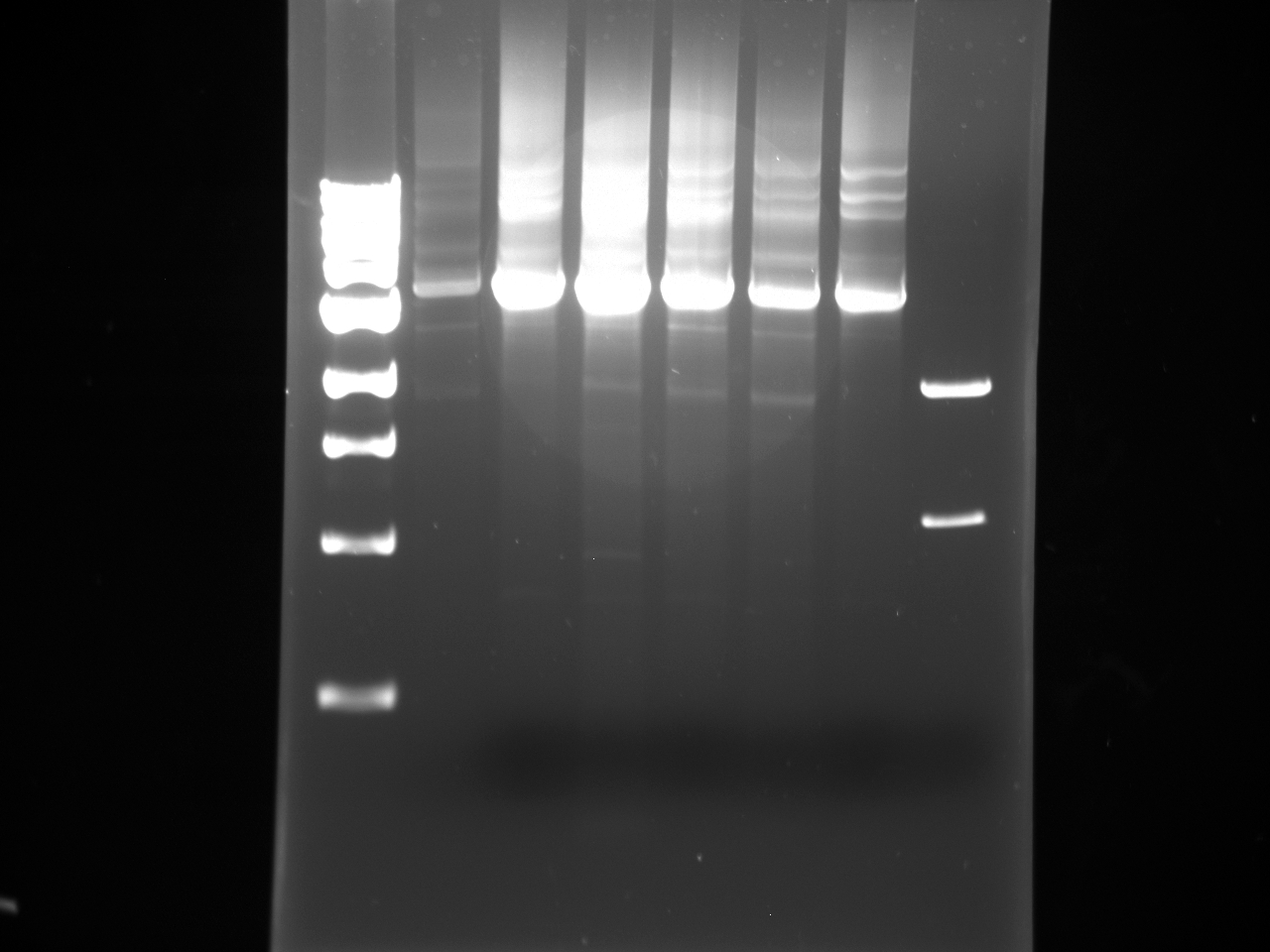
Ran the following Gel:
1 2 3 4 5 6 7 8 1000bp Ladder H58 U58 H60 U60 H62 U62 1000bp
(IMAGE ON WEBSITE https://sites.google.com/site/wuigem/lab-notebook/z/august-23-mondaY)
 "
"