Team:UNIPV-Pavia/Project/solution
From 2010.igem.org
(→How to perform genome integration) |
(→Validation of pir strains to propagate the R6K replication origin) |
||
| Line 310: | Line 310: | ||
===Validation of pir strains to propagate the R6K replication origin=== | ===Validation of pir strains to propagate the R6K replication origin=== | ||
| + | The BW25141 (pir+) and BW23474 (pir-116) ''E. coli'' strains were chosen to propagate the vectors with the R6K replication origin at medium copy (~15 molecules per cell) and high copy (~250) respectively. | ||
| + | The results about their capability to propagate R6K plasmids are shown here: | ||
| + | {|border=1 | ||
| + | |'''Strain''' | ||
| + | |'''Efficiency with no DNA''' | ||
| + | |'''Efficiency with pSB*** (positive control)''' | ||
| + | |'''Efficiency with the self-ligated <partinfo>BBa_K300008</partinfo> (R6K plasmid)''' | ||
| + | |- | ||
| + | |BW25141 (<partinfo>BBa_K300084</partinfo>, pir+) | ||
| + | |0 | ||
| + | |10^5 | ||
| + | |10^5 | ||
| + | |- | ||
| + | |BW23474 (<partinfo>BBa_K300085</partinfo>, pir-116) | ||
| + | |0 | ||
| + | |10^6 | ||
| + | |10^6 | ||
| + | |- | ||
| + | |DH5alpha (<partinfo>BBa_V1001</partinfo>, neg control) | ||
| + | |0 | ||
| + | |10^8 | ||
| + | |0 | ||
| + | |- | ||
| + | |MC1061 (<partinfo>BBa_K300078</partinfo>, neg control) | ||
| + | |0 | ||
| + | |10^6 | ||
| + | |0 | ||
| + | |- | ||
| + | |MG1655 (<partinfo>BBa_V1000</partinfo>, neg control) | ||
| + | |0 | ||
| + | |10^5 | ||
| + | |six small colonies from 2 ng of DNA | ||
| + | |} | ||
| + | |||
| + | These results show that the R6K conditional replication origin can be only propagated in pir+ and pir-116 strains (<partinfo>BBa_K300084</partinfo> and <partinfo>BBa_K300085</partinfo>), while the transformation of other strains with the R6K plasmid yielded no colonies after transformation. The exception was MG1655, for which six small unwanted colonies appeared on the selective plate. Plasmid DNA was purified/HindIII-digested/gel-run for at least one colony for BW25141-<partinfo>BBa_K300008</partinfo> and BW23474-<partinfo>BBa_K300008</partinfo>, while all the six colonies were analyzed for MG1655-<partinfo>BBa_K300008</partinfo> plate. The electrophoresis showed the expected length of the transformed DNA for all the clones except for the MG1655 six colonies, for which a smear was present in the lane (data not shown). Probably, Cm-resistant contaminants were present in the MG1655 culture during the preparation of competent cells. | ||
| + | |||
| + | For this reason, competent cells were prepared again for MG1655 and the transformation procedure was repeated for this strain, yielding no colonies in the <partinfo>BBa_K300008</partinfo> plate as expected. | ||
| + | |||
| + | Miniprep of the BW25141 and BW23474 strains transformed with <partinfo>BBa_K300008</partinfo> yielded a DNA concentration of ~20 ng/ul (qualitatively comparable with medium copy number plasmids) and ~90-100 ng/ul (qualitatively comparable with high copy number plasmids). | ||
| + | The results shown in the table above also show that the R6K plasmid in pir+ and pir-116 strains was transformed with the same efficiency as the pSB*** positive control plasmid, demonstrating that the R6K origin doesn't give any handicap in plasmid transformation. | ||
| + | So, the BW25141 and BW23474 strains can be successfully used to propagate the integrative vector after the excision of the pUC19-derived high copy replication origin, present in the default insert <partinfo>BBa_I52002</partinfo>. | ||
===Integration of the desired BioBrick part into the Phi80 genome locus=== | ===Integration of the desired BioBrick part into the Phi80 genome locus=== | ||
Revision as of 20:07, 22 October 2010
|
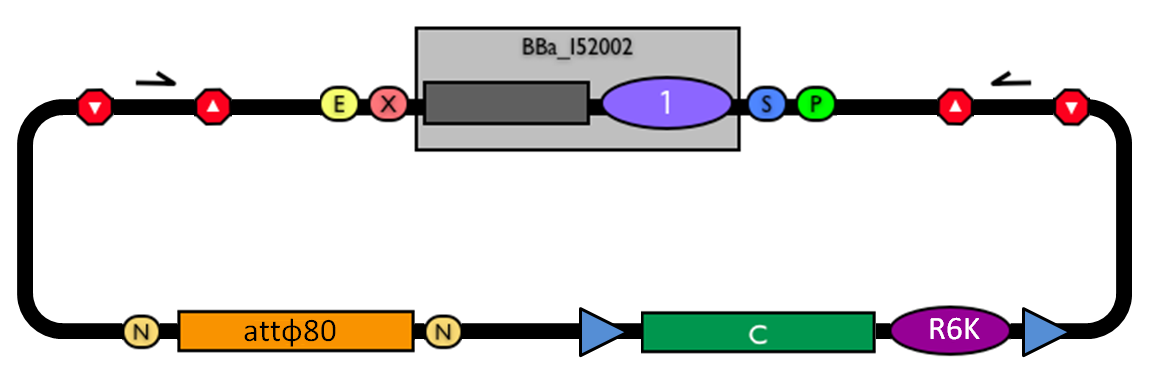
This vector can be considered as a base vector, which can be specialized to target the desired integration site in the host genome. The default version of this backbone has the bacteriophage Phi80 attP (<partinfo>BBa_K300991</partinfo>) as integration site. This vector enables multiple integrations in different positions of the same genome.
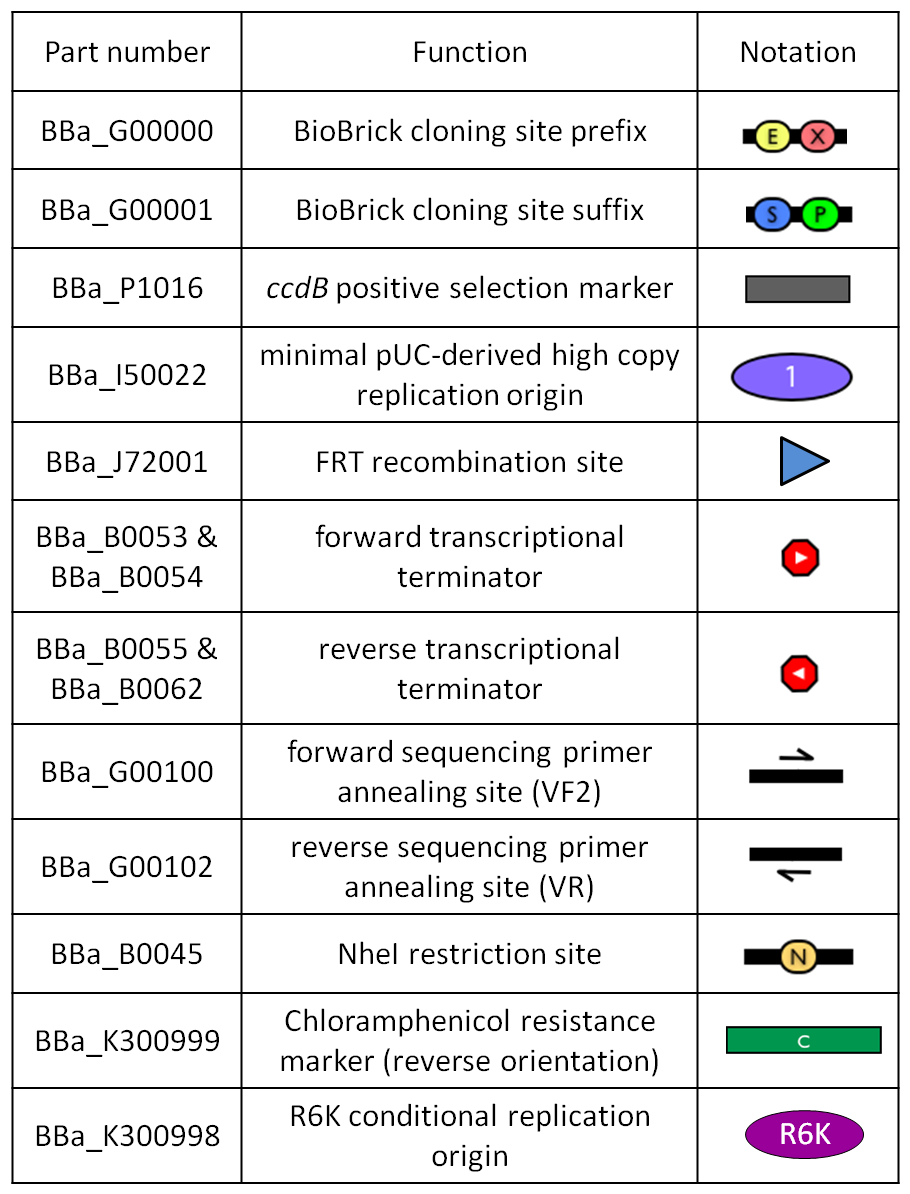
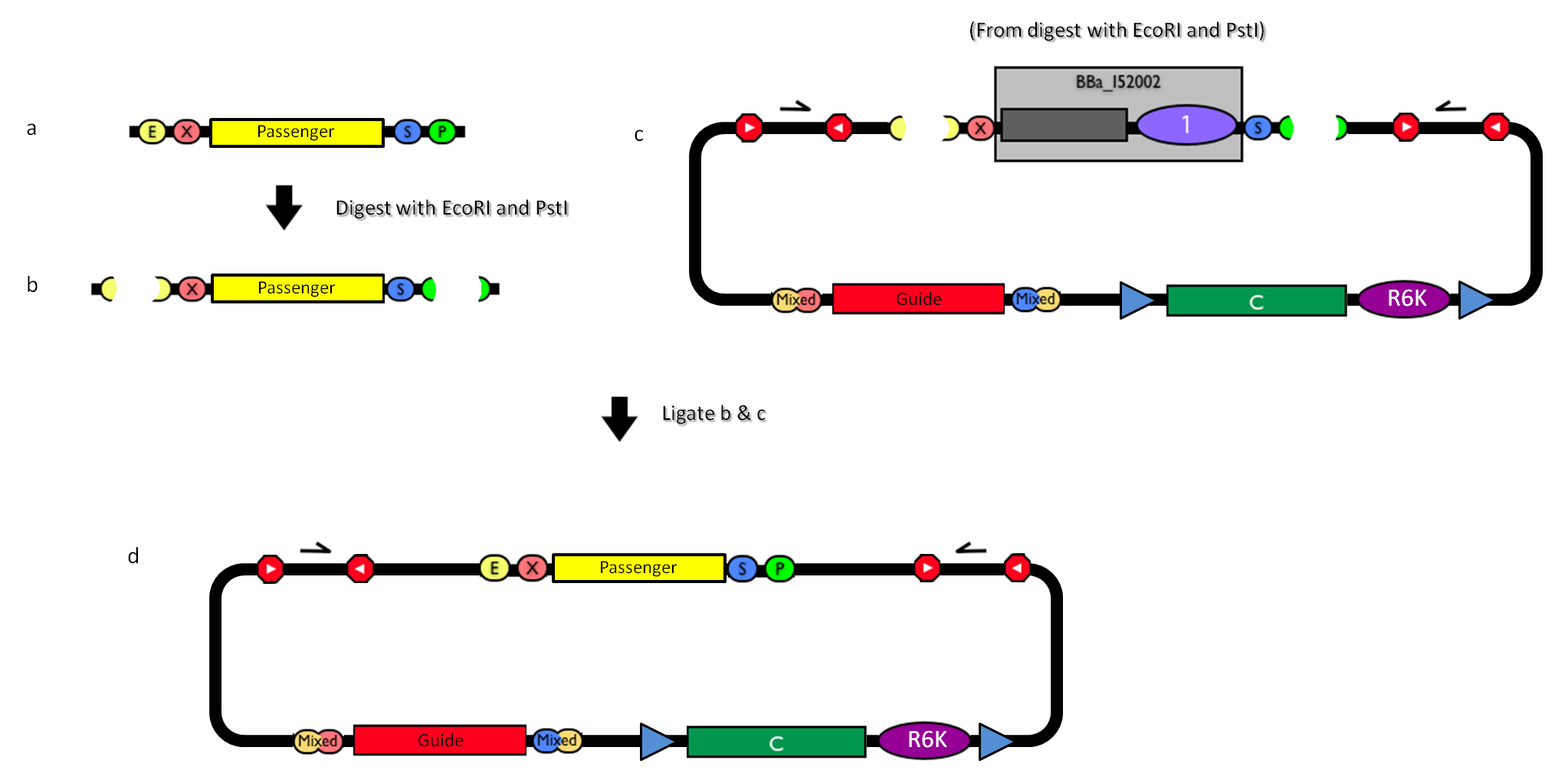
GlossaryThe passenger is the desired DNA part to be integrated into the genome. The guide is the DNA sequence that is used to target the passenger into a specific locus in the genome.
The main design features for vector engineering and for the genome integration of the vector are reported below. Vector engineering features:
Genome integration features:
How to use it<partinfo>BBa_K300000</partinfo> can be:
How to propagate it before performing genome integrationThe default version of this vector contains the <partinfo>BBa_I52002</partinfo> insert, so it *must* be propagated in a ccdB-tolerant strain such as DB3.1 (<partinfo>BBa_V1005</partinfo>). After the insertion of the desired BioBrick part in the cloning site, this vector does not contain a standard replication origin anymore, so it *must* be propagated in a pir+ or pir-116 strain such as BW25141 (<partinfo>BBa_K300984</partinfo>) or BW23474 (<partinfo>BBa_K300985</partinfo>) that can replicate the R6K conditional origin (<partinfo>BBa_J61001</partinfo>).
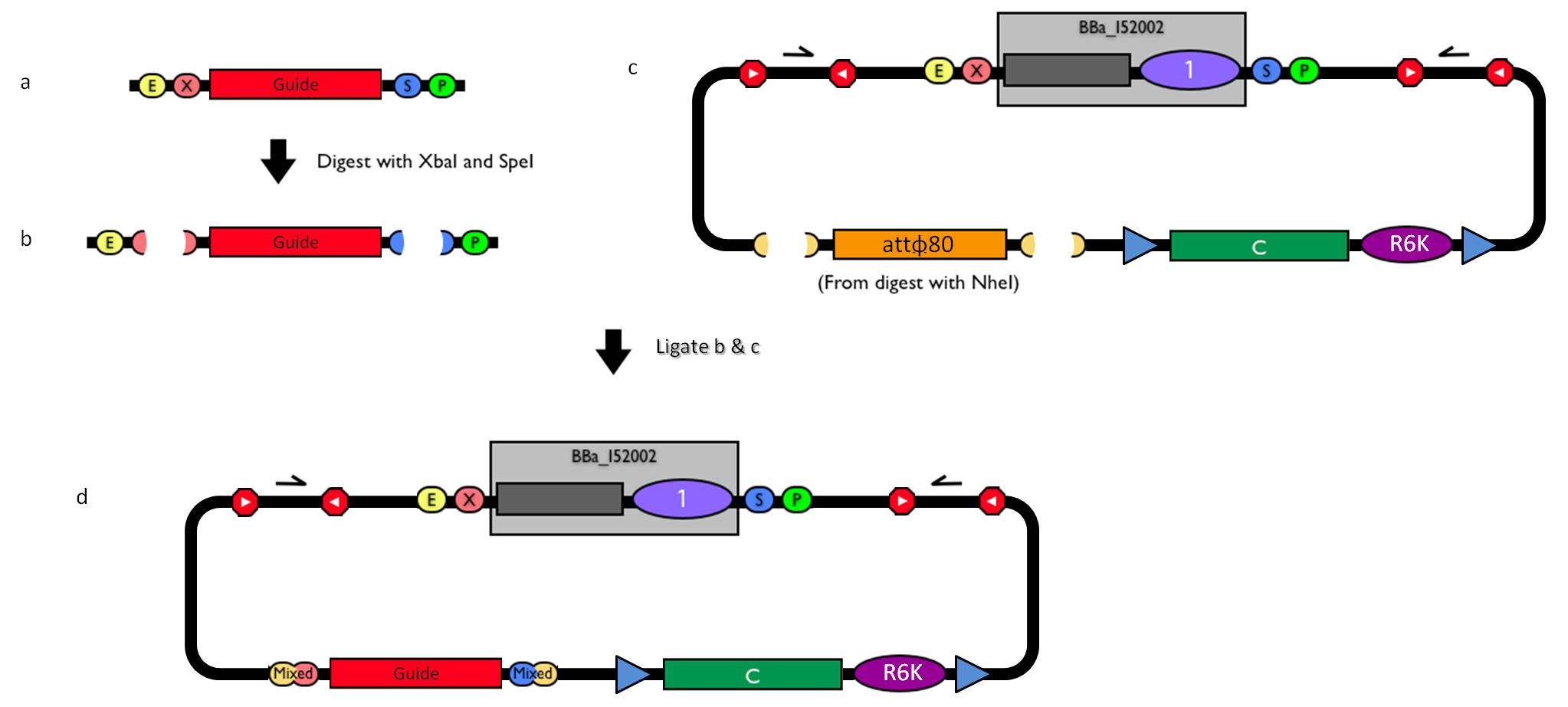
How to engineer itThe DNA guide can be changed as follows:
How to perform genome integrationThe integration into the E. coli chromosome can exploit the bacteriophage attP-mediated integration or the homologous recombination. Detailed protocols about attP-mediated integration can be found here:
Detailed protocols about homologous recombination can be found here:
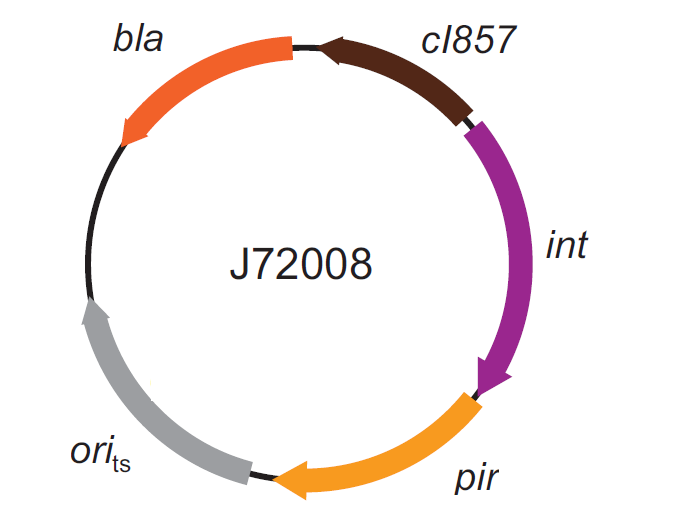
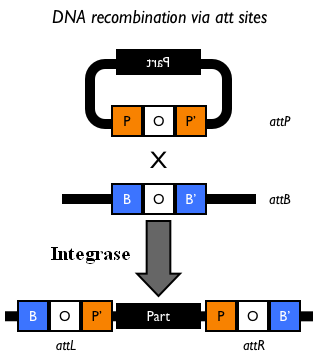
This integration method is applicable when the host strain does not have prophages in the att(Phi80) locus. TOP10 (<partinfo>BBa_V1009</partinfo>) and DH5alpha (<partinfo>BBa_V1001</partinfo>) strains have the Phi80 prophage and so their chromosome cannot be engineered with this procedure. The genomic integration of the desired BioBrick part into the attP(Phi80) locus has to be mediated by co-transforming a helper plasmid (such as the Amp-resistant <partinfo>BBa_J72008</partinfo> plasmid) carrying the IntPhi80 site-specific integrase gene under the control of a thermoinducible promoter (see Fig.5). The helper plasmid also has a heat-sensitive replication origin, whose replication can be inhibited at temperatures of 37-42°C, while a permissive temperature for this vector is 30°C. For this reason, it can be cured at high temperatures, when the integrase expression is triggered at the same time. The Phi80 integrase mediates the site-specific recombination between the attP site in the integrative vector and the attB site in the bacterial genome (for a schematic description of this process, see Fig.6 and http://partsregistry.org/Recombination). Thanks to its R6K conditional replication origin, the integrative vector cannot be replicated in common E. coli strains, so the Chloramphenicol resistant bacteria are actual integrants. In the Materials and Methods section, a detailed protocol to target the desired BioBrick part into the Phi80 locus is reported. How to perform multiple integration in the same genomeWhen this vector is integrated into the genome, the desired passenger should be maintained into the host, as well as the Chloramphenicol resistance marker and the R6K conditional replication origin. The CmR and the R6K can be excised from the genome by exploiting the two FRT recombination sites that flank them. The Flp recombinase protein mediates this recombination event, so it has to be expressed by a helper plasmid, such as pCP20 (CGSC#7629). This enables the sequential integration of several parts using the same antibiotic resistance marker, which can be eliminated each time. Detailed protocols about homologous recombination can be found here:
Materials and MethodsPlasmids and strains: the <partinfo>BBa_J72008</partinfo> helper plasmid was kindly given by Prof. JC Anderson (UC Berkeley). MC1061 (<partinfo>BBa_K300078</partinfo>) and MG1655 (<partinfo>BBa_V1000</partinfo>) E. coli strains and the pCP20 helper plasmid were purchased from the Coli Genetic Stock Center (Yale University). DH5alpha (<partinfo>BBa_V1001</partinfo>) strain was purchased from Invitrogen.
efficiency [CFU/ug of DNA]= # CFU * 1000 ng of DNA / amount of transformed DNA [ng]
Self-ligated <partinfo>BBa_K300008</partinfo> was prepared by digesting <partinfo>pSB1A2</partinfo>-<partinfo>BBa_K300008</partinfo> (yielded by BioBrick Standard Assembly) with XbaI-SpeI. The insert was isolated and purified from a 1% agarose gel. Then, it was self-ligated to generate a Cm-resistant R6K plasmid. The colonies were counted in each plate and the transformation efficiency was estimated as described before.
All the DNA manipulations were performed according to manufacturer's protocols.
Validate the loss of the helper plasmid by inoculating colonies in Cm (at 12.5 ug/ml) media and counterselecting them in Amp (at 50 ug/ml) media. Validate the correct integration position by performing colony PCR with primers P1/P2, P3/P4, P1/P4, P2,P3 and VF2/VR. Validate the phenotype (when possible).
Validate the loss of the helper plasmid by inoculating colonies in Amp (at 100 ug/ml) media and validate the loss of the Cm resistance from the genome by inoculating colonies in Cm (at 12.5 ug/ml) media. Validate the correct length of the integrated part without Cm resistance and R6K origin by performing colony PCR with primers P1/P4 (which amplify the entire Phi80 locus) and VF2/VR (which amplify the integrated part). Validate the phenotype (when possible).
ResultsValidation of pir strains to propagate the R6K replication originThe BW25141 (pir+) and BW23474 (pir-116) E. coli strains were chosen to propagate the vectors with the R6K replication origin at medium copy (~15 molecules per cell) and high copy (~250) respectively. The results about their capability to propagate R6K plasmids are shown here:
These results show that the R6K conditional replication origin can be only propagated in pir+ and pir-116 strains (<partinfo>BBa_K300084</partinfo> and <partinfo>BBa_K300085</partinfo>), while the transformation of other strains with the R6K plasmid yielded no colonies after transformation. The exception was MG1655, for which six small unwanted colonies appeared on the selective plate. Plasmid DNA was purified/HindIII-digested/gel-run for at least one colony for BW25141-<partinfo>BBa_K300008</partinfo> and BW23474-<partinfo>BBa_K300008</partinfo>, while all the six colonies were analyzed for MG1655-<partinfo>BBa_K300008</partinfo> plate. The electrophoresis showed the expected length of the transformed DNA for all the clones except for the MG1655 six colonies, for which a smear was present in the lane (data not shown). Probably, Cm-resistant contaminants were present in the MG1655 culture during the preparation of competent cells. For this reason, competent cells were prepared again for MG1655 and the transformation procedure was repeated for this strain, yielding no colonies in the <partinfo>BBa_K300008</partinfo> plate as expected. Miniprep of the BW25141 and BW23474 strains transformed with <partinfo>BBa_K300008</partinfo> yielded a DNA concentration of ~20 ng/ul (qualitatively comparable with medium copy number plasmids) and ~90-100 ng/ul (qualitatively comparable with high copy number plasmids). The results shown in the table above also show that the R6K plasmid in pir+ and pir-116 strains was transformed with the same efficiency as the pSB*** positive control plasmid, demonstrating that the R6K origin doesn't give any handicap in plasmid transformation. So, the BW25141 and BW23474 strains can be successfully used to propagate the integrative vector after the excision of the pUC19-derived high copy replication origin, present in the default insert <partinfo>BBa_I52002</partinfo>. Integration of the desired BioBrick part into the Phi80 genome locusIntegration of the desired BioBrick part into the Phi80 genome locusValidation of the loss of BBa_J72008 Validation of the actual integration site Validation of the integrants phenotype Chloramphenicol resistance marker excisionValidation of the loss of pCP20 and the resistance marker Validation of the marker-less phenotype
DiscussionA novel integrative vector for E. coli has been successfully designed, constructed and used to integrate two proof of concept protein expression systems in two commonly used E. coli strains. The results showed that the vector is fully functional and can integrate into the correct targeted locus of the host chromosome through the Phi80 site-specific recombination system by using <partinfo>BBa_J72008</partinfo>, an existing BioBrick helper plasmid from the Registry. In most cases, the integration occurs in tandem copies, probably because of the too high Chloramphenicol concentration used during the selection of integrants, which forces multiple integration of Cm-resistant constructs. This concentration was the same used during the pSC101 low copy plasmid (~5 copies per cell) selection. In some cases, it is desirable to have a single copy of the desired BioBrick in the genome, for example when the gene dosage is important. In [Haldimann A and Wanner BL, 2001] the usage of Chloramphenicol at 6 ug/ml yielded a very high percentage of single integrants. However, when tested in our lab, the MG1655 strain could survive on LB plates with Cm at 6 ug/ml and also at 8 ug/ml. For this reason a higher concentration of Cm was chosen for selection. Further studies should investigate the optimal antibiotic concentration to yield the highest single integrants percentage as possible.
Integrative standard vector for yeast
Self-cleaving affinity tags to easily purify proteins
|
 "
"