Team:UIUC-Illinois-Software/NextVersion
From 2010.igem.org
 | ||
Next Version of BioMORTAR
While BioMORTAR has succeeded in its goal to help automate the design process in bacterial and metabolic engineering, it still can be improved on many fronts.
First and foremost, the next version will include a user registry, coded in Django, that will allow users to create a profile and save their designed plasmids on the server while displaying the plasmids their lab currently stores. This will allow the users of BioMORTAR to be able to more easily share resources they have, and access the resources they need to create their desired plasmid. It will also allow BioMORTAR to match parts of stored plasmids to plasmids generated by the plasmid designer, then reference them to the user so that the user can have easy access to the resources to implement the designed plasmid in vivo.
The next step of development of BioMORTAR 2.0 will be a security feature that will deter users from using this tool for malicious purposes. The registry in place will log activity from each user and store the data on the server, allowing for easy access to records in case of need from law enforcement.
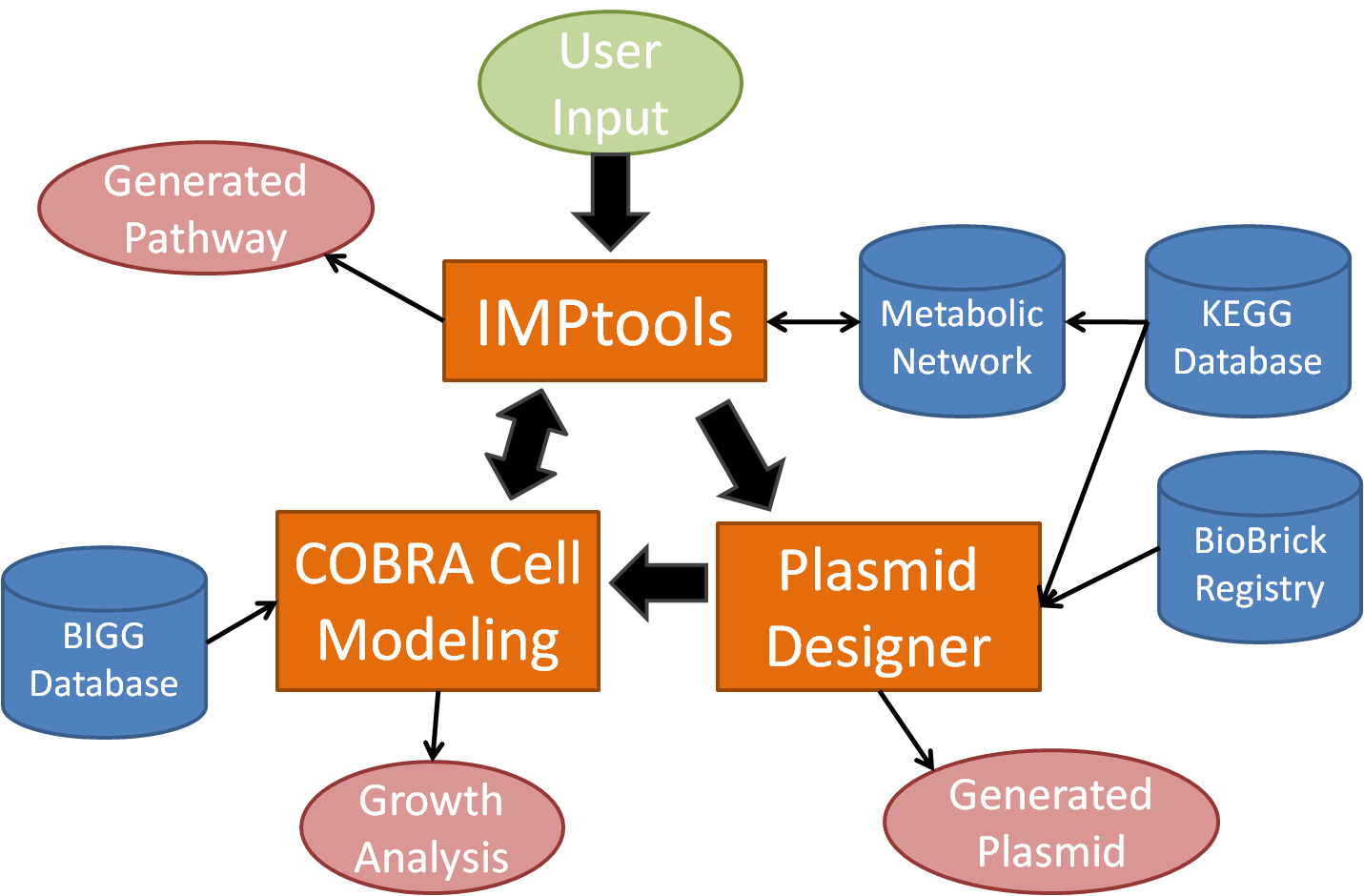
One of the most important new functions in the next version will be the ability for the COBRA cell modeling tool to directly interact with the IMPtools module in order to automatically optimize the metabolic pathway for cell growth. With the implementation of the new features in the next version, BioMORTAR will be able to go above and beyond its initial goal in the automation of cell design.
 "
"