Team:Tsinghua/project/outline/m1
From 2010.igem.org
(New page: __NOTOC__ {{:Team:Tsinghua/header}} <html> <body id="bd_project"> <div id="content_box"> </html> {{:Team:Tsinghua/leftbar}} <html> <script> navlist=new Array("Background", "Project Outlin...) |
|||
| Line 13: | Line 13: | ||
writenav(navlist, linkl, 1, subs); | writenav(navlist, linkl, 1, subs); | ||
</script> | </script> | ||
| - | <div id="main_content"> | + | <div id="main_content"> |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
<a name="mod1"></a></html> | <a name="mod1"></a></html> | ||
| Line 217: | Line 160: | ||
[[Image:THUProjectFigure8.JPG|650px]] | [[Image:THUProjectFigure8.JPG|650px]] | ||
| - | <html><br><br> | + | <html><br><br> <a href="#top"> <img src="https://static.igem.org/mediawiki/2010/d/dd/TOP.png" />TOP</a> |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | <a href="#top"> <img src="https://static.igem.org/mediawiki/2010/d/dd/TOP.png" />TOP</a> | + | |
</div> | </div> | ||
</div></div></body> | </div></div></body> | ||
</html> | </html> | ||
Revision as of 15:36, 25 October 2010

Module I
Abstract
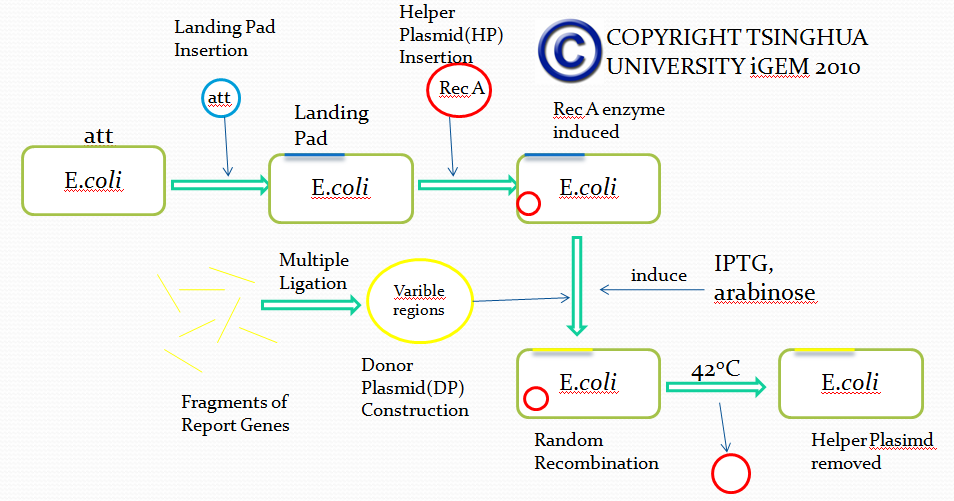
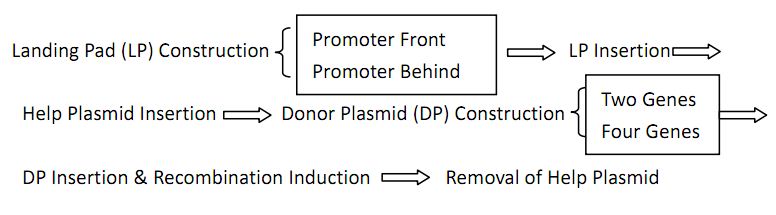
Flow chart
Landing Pad Construction and Insertion
Purpose of this step
Landing pad insertion is the first step of our two-step recombination system. By doing this step we induced our “RSS Sequence” into E.coli.
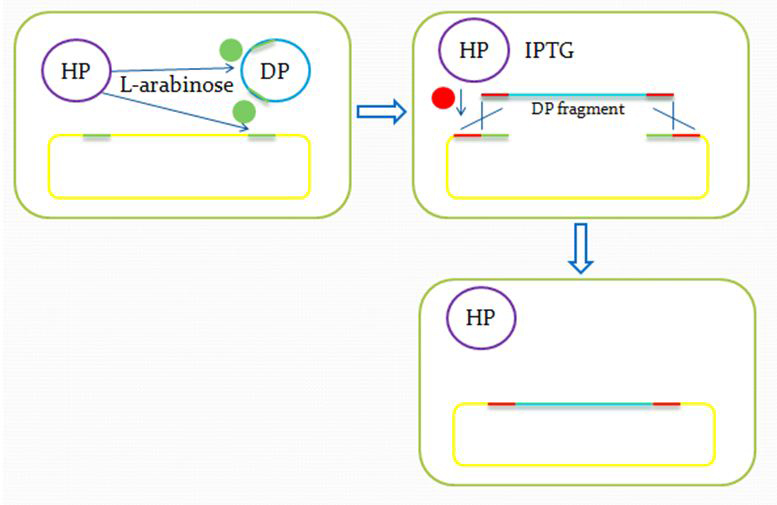
We insert a “landing pad” fragment which includes a promoter (placIQ1) and a tetracycline resistance gene (tetA) flanked by I-SceI recognition sites and 20-bp landing pad regions (LP1 and LP2) into Escherichia coli chromosome via att recombination. Then the helper plasmids encoding I-SceI endonuclease and λ-Red and donor plasmid encoding various antibiotic genes flanked by I-SceI recognition sites and same landing pad regions (LP1 and LP2) are transformed, which will be introduced in detail in next part. I-SceI expression is induced via the addition of L-arabinose. I-SceI recognition sites in the donor plasmid and chromosome are cleaved. Integration of the fragment in donor plasmid is facilitated by IPTG-induced λ-Red expression.
This two-step recombination method allows for the insertion of very large fragments into a specific location in Escherichia coli chromosome and in any orientation.
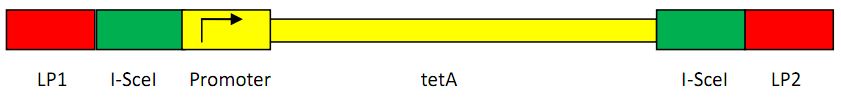
Construction of landing pad
Landing pad consists of the following parts: a promoter (placIQ1), two landing pad regions, two I-SceI recognition sites and a tetracycline resistance gene (tetA).
Another construction strategy is shown as below:
The first construction is used as the four-gene donor plasmid method, while the latter is for the two-gene donor plasmid method. We take the advantage of the former construction in the final recombination, because the promoter will not be cut when the recombination happens, thus saving us lots of time. As the latter construction, we can easily add a promoter before the gene, in two-gene donor plasmid A(shown in the Donor Plasmid Construction Part).
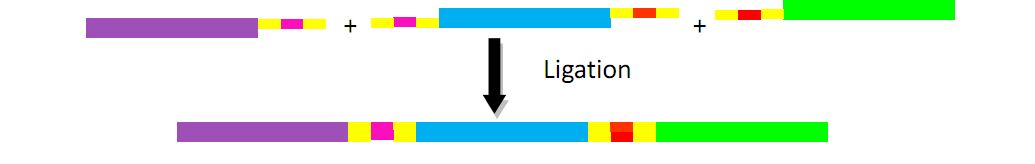
First we obtain two target DNAs – promoter and tetA with PCR. Then We use the overlap extension polymerase chain reaction (or OE-PCR) to link two fragments together. Primer 2 and primer 3 have homologous sequences, so one segment can anneal to the other in certain conditions, in other words, the two segments can overlap into one segment after extension. If necessary, operate PCR again with primer 1 and primer 4 to obtain more products.
Landing pad insertion-- ATT RECOMBINATION
Our approach is based on genome targeting systems that utilize plasmids carrying a conditional-replication origin and a phage attachment (attP) site. We refer to our plasmids as CRIM (conditionalreplication, integration, and modular) plasmids. CRIM plasmids can be integrated into or retrieved from their bacterial attachment (attB) site by supplying phage integrase (Int) without or with excisionase (Xis) in trans.
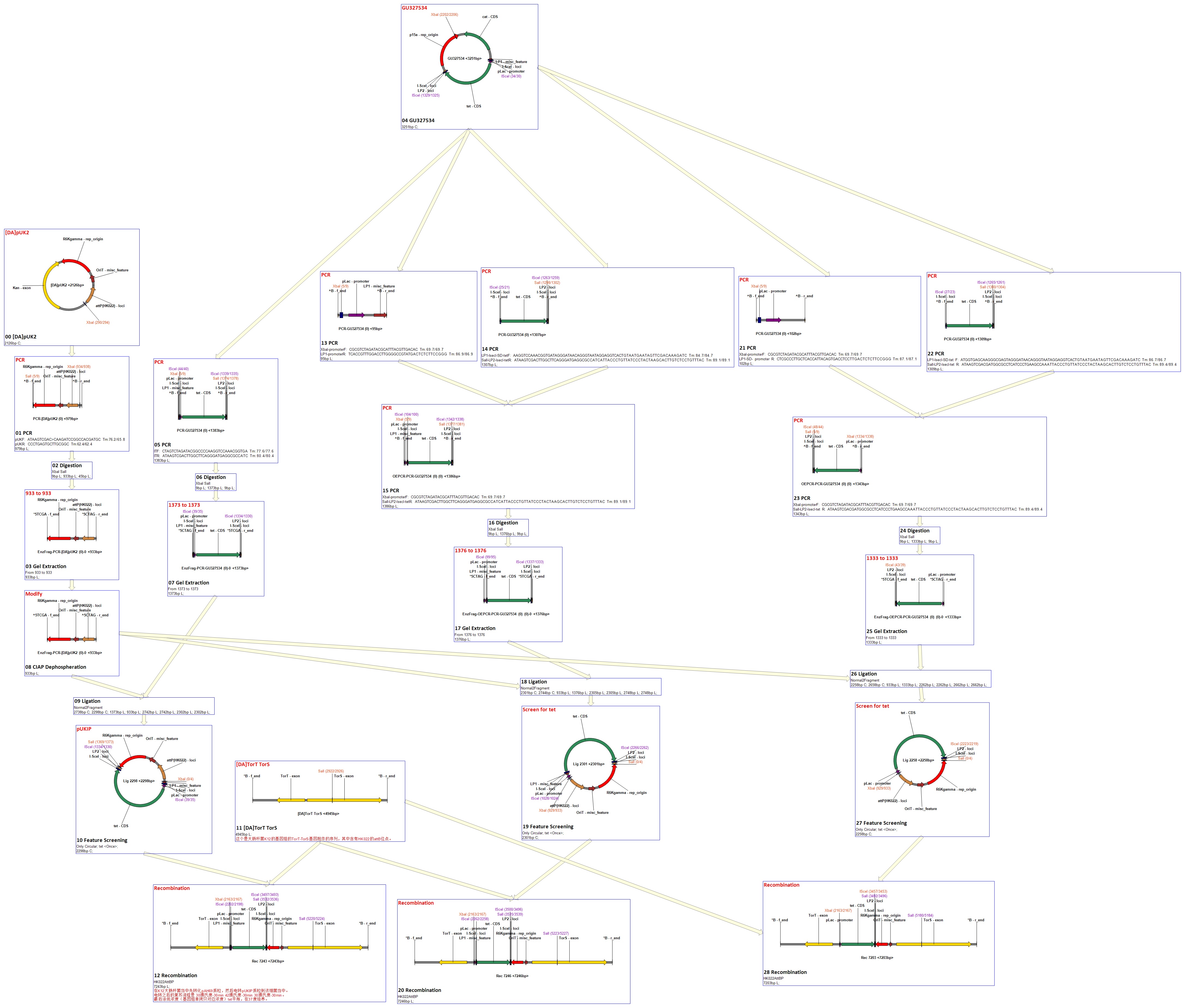
We got a strain with plasmid pUK2 from LAB. Than we develop a E.coli strain contains the helper plasmid AH69. These two plasmids are shown below. In order to match other parts of our whole project, the modification that kan-exon should be replaced with tet-SDS was necessary. We got tet-SDS from another plasmid named pkts-cs. Then we use PCR to get the two fragments as PT (from pkts-cs) and V (from pUK2). We ligate PT and V after digestion and modification to form a new plasmid. This new plasmid was named as pUKIP and contains a promoter region, a tet-SDS region and phage attachment site (attP). As reported in the paper, there exist one bacterial attachment site (attB) of HK002 in TorT-TorS gene of the chromosomal DNAs of E.coli K12 strain. That is to say, the PT fragment will integrate into the cell genome after the plasmid was transferred in cells.
Helper Plasmid(HP) Insertion
CRIM plasmid integration
Cells carrying a CRIM helper plasmid were grown in 20 ml of LB cultures with ampicillin at 30°C to an optical density of 600 nm of ca. 0.6 and then made electrocompetent. Following electroporation, cells were suspended in LB without ampicillin, incubated at 30°C for 30 min, at 42°C for 30 min and at 30°C for 30 min, and then spread onto selective agar (tet) and incubated at 37°C. Colonies were purified once nonselectively and then tested for antibiotic resistance for stable integration and loss of the helper plasmid and by PCR for copy number.
CRIM plasmid excision
Cells were transformed with the respective Xis/Int CRIM helper plasmid and then spread on ampicillin agar media at 30°C. Colonies were purified once or twice nonselectively on plates that were incubated for 1 h at 42°C and overnight at 37°C. They were then tested for antibiotic sensitivities and by PCR for loss of the integrated plasmid.
Helper Plasmid Insertion: “Applying Recombinase Coding Gene”
We transferred our helper plasmid, which contains RecA enzyme coding gene and I-Sel enzyme coding gene, into E.coli. These two enzymes will act exactly as what the recombinase does.
Donor Plasmid(DP) Construction: Constructing Our “Antibody Coding Gene Library”
Purpose of this step:
The construction of DP provides us multiple genes, which vary from each other, forming the library for us to select. After inserting landing pad and helper plasmid to E.coli, we must construct a series of donor plasmids to demonstrate that this system can truly realize the recombinant process in E.coli, thus we can further use this module to simulate the recombination of antibody gene in mammalian B cells. We not only need to test the efficiency of recombination, but also ensure that genes we get from this recombinant process can be expressed correctly and have their original function. So we intend to construct three plasmids to test the system.
Experiment design and expected results:
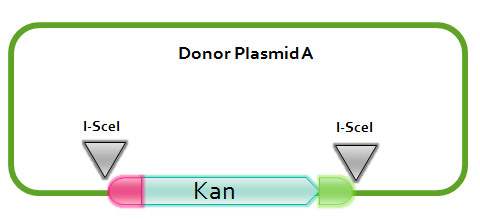
[1] Donor plasmid A
In donor plasmid A, we insert only one gene, kanamycin resistant gene (Kanr) to test our recombination. At the 5’ end of Kanr, we add I-scel recognizing sequence (which is represented by the white arrow) and recombination sequence 1 (which is shown in red). At the 3’ end, we add another recombination sequence (which is shown in blue) and also I-scel recognizing sequence.
After construction, we will transform this donor plasmid to E.coli with landing pad and helper plasmid. After arabinose and IPTG inducing, the restriction enzyme I-scel will cut down Kanr (containing recombination sequences), which can recombine with the bacterial chromosome.
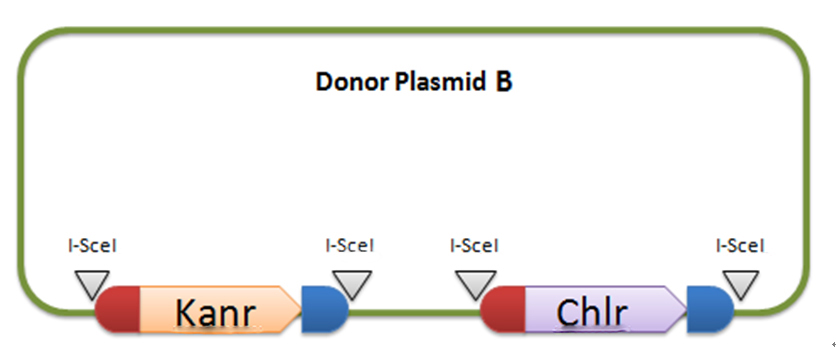
[2] Donor plasmid B
In donor plasmid B, we insert two genes, kanamycin resistant gene (Kanr) and Chloromycetin resistant gene (Chlr). At the 5’ end of these two genes, we add I-scel recognizing sequence (which is represented by the white arrow) and recombination sequence 1 (which is shown in red). At the 3’ end, we add another recombination sequence (which is shown in blue) and also I-scel recognizing sequence.
After construction, we will transform this donor plasmid to E.coli with landing pad and helper plasmid. After arabinose and IPTG inducing, the restriction enzyme I-scel will cut down these two genes (containing recombination sequences), which can recombine with the bacterial chromosome. In our expectation, either kanr or chlr will replace the landing pad, resulting in the bacteria resistance to either kanamycin or chloromycetin, but not both. This process will be random, so we can get as many colonies resistant to kanamycin as those resistant to chloromycetin.
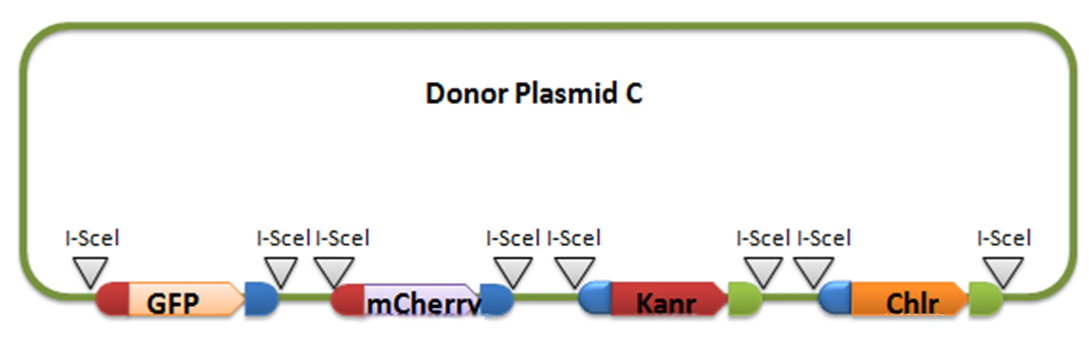
[3] Donor plasmid C
Donor plasmid C includes four genes, GFP, mCherry, Kanr, Chlr, respectively. The same with genes in donor plasmid A, we add recombination sequences and I-scel recognition sequences to the ends of each genes. The recombination sequences of GFP and mCherry are identical, and those of Kanr and Chlr are the same. Note that recombination sequence at 3’ end of GFP (mCherry) and that at 5’ end of Kanr (Chlr) are the same, so we can get a random recombination of two genes, one is a fluorescence gene and the other is resistant gene, creating 2X2=4 different results.
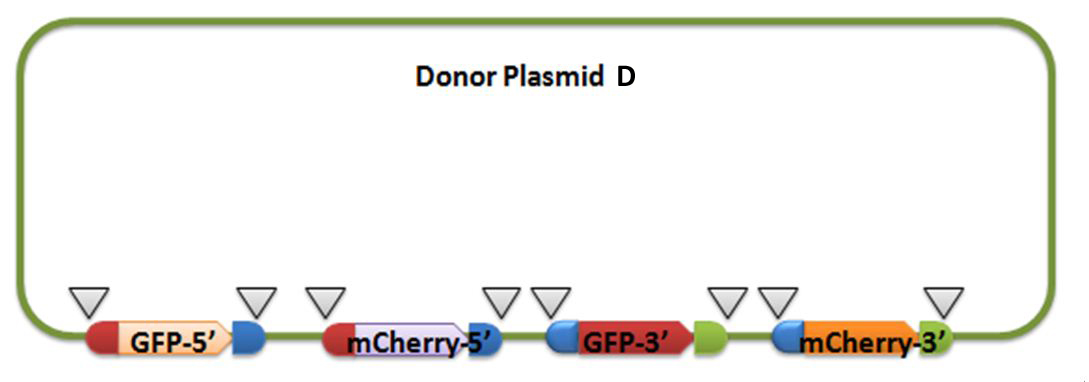
[4] Donor plasmid D
For constructing donor plasmid D, we first cut the GFP and mCherry to 2 fragments respectively. Then we insert these four fragments into the plasmid in the order shown in the above picture. We expect that we can see either green or red fluorescence after transforming and inducing. Through this experiment, we can tell whether or not recombination sequence will affect the normal function of genes, further demonstrate that antibody producing by our system will be effective. On the other hand, we should note that the sequence length is another significant reason of antibody diversity, so the effect of recombination sequence on antibody will much small than that on fluorescence genes.
Strategies
Traditionally, we can construct these three plasmids by inserting genes into the plasmid one by one. However, considering the huge number of antibody fragments, we try our best to seek other strategies to complete the ligation of multiple fragments, which can be done more quickly and efficiently.
Principle
The fragments include different landing pad regions and endoclease recognition site, and are grouped together by sharing the same landing pad region.
Method
In the demand of larger amount of fragments and constructed plasmids, we find two ways to realize such purpose, regarded as the key point of our whole project-guarantee the large variation of antibody.
Removal of Helper Plasmid(HP)
Helper plasmid (HP) includes a temperature sensitive pSC101 replication origin, which maintains the plasmid at low copy number. This plasmid is thus easily removed by growth at 42℃ and screening against spectinomycin resistance. Of more concern is the donor plasmid, which is cured by I-SceI cleavage, and this process is very efficient, with only about 1% of cells retaining the donor plasmid.
Donor Plasmid(DP) Insertion & Recombination Induction
Propose of this step:
After the reform of the E.coli genome and the construction of the donor plasmid, we need to test our module’s function. First we should insert the donor plasmid(DP) into the E.coli, and then induce the recombination by adding IPTG and arabinose. Arabinose active the I-Sel restriction enzyme, then cut DP and genome at the same site. After the digestion, we add IPTG to help the recombination, using the homologous sequence near the cohesive end.
DP Insertion:
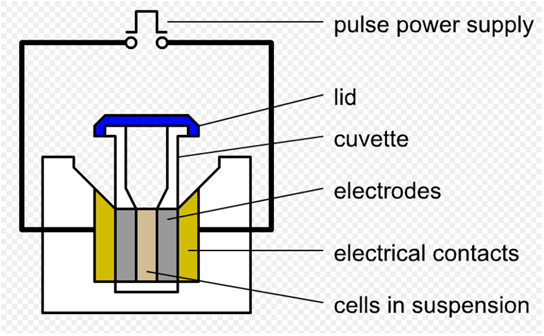
As we said before, we use a pre-altered template to amplify landing pad fragments using the landing pad regions as standardized priming sites. Here we used the conventional electroporation method to transform the DP into E.coli genome, then incubate the plate at 37°overnight. Electroporation is a significant increase in the electrical conductivity and permeability of the cell plasma membrane caused by an externally applied electrical field. It is a dynamic phenomenon that depends on the local transmembrane voltage (we used 1 V) at each point on the cell membrane. If E.coli and DP are mixed together, the plasmids can be transferred into the cell after electroporation. This procedure is highly efficient than chemical transformation. (Partly from wikipedia)
Recombination Induction:
Individual colonies were inoculated into 5 ml of EZ-Rich Defined Medium (RDM; Teknova) +0.5% glycerol, 2mM IPTG, and 0.2% w/v L-arabinose. After growing at 37°C for 1 h in a shaking water bath, we transfer the medium to 30_C shaking water bath for 4 h, then 100 mg/ml spectinomycin was added. At first the I-Sel enzyme is expressed to cut the genome and DP at the same site. And this step is used to constitutive express the Rec A enzyme, thus initiating the recombination by the homologous region. The appropriate antibiotic for the given insertion fragment was then added (25 mg/ml kanamycin, 34 mg/ml chloramphenicol), and the cultures were grown overnight. The next day, 100 ml sample was plated on LB plates with the appropriate antibiotic and grown at 37°C. We test the sample by screening it on LB plates containing 100 mg/ml ampicillin or 10 mg/ml tetracycline to verify the loss of the landing pad and donor plasmid.
 "
"







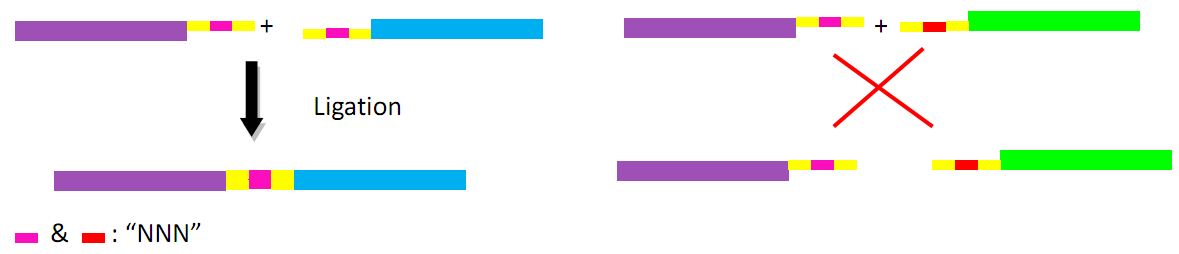
 TOP
TOP