Team:Toronto/Project
From 2010.igem.org
| Home | Project | Design | Protocols | Notebook | Results | Parts Submitted to the Registry | Modeling | Software Used | Human Practices | Safety | Team | Official Team Profile | FAQ | Acknowledgments |
|---|
Contents |
Project Description
Background
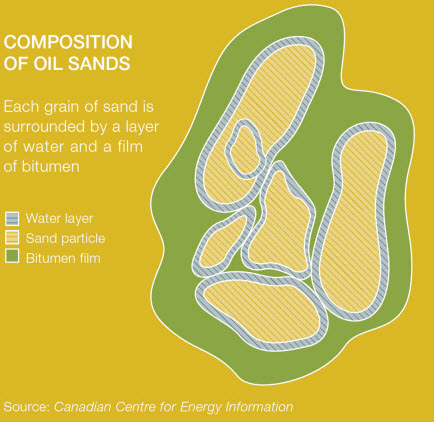
Oil Sands
Oil sands are naturally occurring mixtures of sand or clay, water and an extremely dense and viscous form of petroleum called bitumen.
Bitumen is a collection of heavy hydrocarbons, which results from preferential microbial degradation of lighter hydrocarbons in oil reserves over geological time scales. These heavy oil reserves are some of the largest fossil fuel stores in the world with more stored energy than traditional oil reserves.Oil sands represent approximately 67% of the world’s total petroleum resource, with the two largest deposits being in Canada and Venezuela. Together they contain approximately 3.6 trillion barrels of oil, compared to 1.75 trillion barrels of conventional oil worldwide. The oil sands in the Athabasca Basin in northeastern Alberta, Canada represent the world’s largest resource estimated to contain over 1.7 trillion barrels of bitumen.
Tailings Ponds
Oil sands are developed either through open-pit mining or deep underground (in situ) production. In open-pit mining, hot water is used to separate bitumen from the mixture of sand and clay.
Each cubic meter of mined oil sands requires up to 3 m3 of water and produces about 4 m3 of slurry wastes containing sand, clay, water, unrecovered bitumen and dissolved inorganic and organic compounds. This mixture is then sent to a discontinued mine pit, referred to as a tailings pond. It is a large man-made collection of waste, which has a negative impact on the environment.
Over 70% of the water demand is met by recycling the recovered water. Even so, these wastewaters are accumulating at a rate of about 0.1-0.2 m3 per ton of oil sands processed. Recycling of the water back to the extraction process further concentrates contaminants as more waste material is added to the already contaminated water. It has been estimated that over 1 billion m3 of tailings pond water will be generated by the year 2025.
They are composed of a wide variety of chemical toxins. Amongst these are polycyclic aromatic hydrocarbons (PAHs), with Naphthalenes, Anthracenes and Fluorenes present at the highest concentrations. These compounds are toxic to the surrounding aquatic ecosystems, and have been shown to have an adverse effect on human health. PAHs are carcinogenic and are also known to have a negative impact on immune function, kidney function, liver function, reproduction and development.
Bioremediation
Bioremediation is the use of microorganisms to return the natural environment altered by contaminants to its original condition. It has been proposed as a possible method for the reclamation of land and water impacted by oil sands development. Although natural examples of these processes exist, they tend to be characterized by slow reaction rates and/or undesirable side reactions.
Over expression of rate limiting enzymes has enjoyed moderate success in overcoming some pathway limitations. However, because enzymes operate within complex, highly organized systems, there is increasing recognition within the metabolic engineering community that spatial organization offers an additional route for optimizing pathway efficiency.
Catechol
Pseudomonas putida, a microbe naturally present in tailings ponds, has the metabolic machinery needed to metabolize many toxins within the tailings ponds. This includes the capability to degrade certain PAHs in tailings ponds such as naphthalenes, anthracenes and fluorenes. These three major PAHs are degraded to form a common compound called catechol. However, this process is slow and inefficient. Since catechol is the common breakdown product in the breakdown of these PAHs, enhancing the breakdown of catechol may lead to an increase in the rate of PAH degradation. This project will look to evaluate how we can increase the degradation rate of catechol in order to achieve faster PAH breakdown.
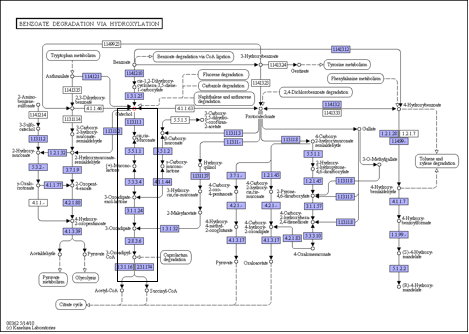
The metabolic network present in P. putida for degrading catechol is highlighted by the black box in Figure 1. (http://www.genome.jp/kegg-bin/show_pathway?ko00362+C00090).
Figure 1. The degradation of catechol to common cellular metabolites, acetyl-CoA and Succinyl-CoA , involves five enzyme catalyzed reactions in Pseudomonas putida.
Metabolic Channeling
Metabolic channeling is the process whereby metabolic intermediates are channelled between the active sites of consecutive enzymes in a biochemical pathway. This serves to increase reaction efficiency by limiting the diffusion of metabolites into the surrounded cellular intracellular environment. Channelling provides a biochemical system with benefits such as:
- Enhancing unfavourable reactions
- Sequestering toxic intermediates
- Protecting unstable intermediates
- Preventing inhibition of other enzymes by the channelling intermediate
Channeling has been utilized in bioengineering to enhance the efficiency of desired biochemical systems. Some methods which have been utilized to mimic examples of metabolic channelling in nature are through:
- Immobilized Enzymes
- Fusion Proteins
- Scaffolds
Immobilized Enzymes Fusion Protein Scaffolding
The Project
This current project aims to take advantage of the catechol degradation pathway present in P. Putida by introducing channelling to increase the efficiency with which catechol is degraded. We anticipate that degrading catechol, an intermediate formed in polyaromatic hydrocarbon (PAH) degradation, will result in an imbalance in cellular catechol concentrations. In turn, we believe there will be increased PAH degradation in order to restore the original cellular catechol concentrations.
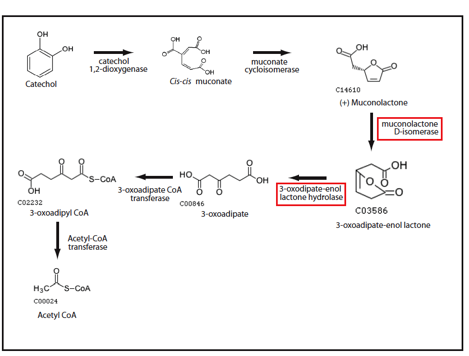
In order to identify enzymes in the catechol degradation pathway that benefit from the introduction of metabolic channelling, we performed in silico analyses with software developed in our lab (Sanford et al.). Preliminary results from our simulations suggests that the 3rd (muconolactone D-isomerase) and 4th (3-oxodipate-enol lactone hydrolase) enzymes in the pathway would benefit most from channelling (Figure 2). Muconolactone D-isomerase catalyzes an energetically unfavourable reaction, while 3-oxodipate-enol lactone hydrolase catalyzes a very energetically favourable reaction. By introducing channeling between these two enzymes, the energetically unfavourable product of the mucanolactone D-isomerase-catalyzed reaction would be converted to the product of 3-oxodipate-enol lactone hydrolase before mucanolactone D-isomerase converts the energetically unfavourable product back into the energetically favourable substrate. The net effect would be an increased flux through the catechol degradation pathway.
Figure 2: Ortho degradation of catechol (1,2-Benzenediol) proceeds in six steps. Only the last enzyme is present in E. coli. Enzyme candidates with the largest predicted channeling effect as predicted by Cell++ are boxed in red.
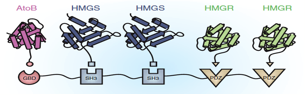
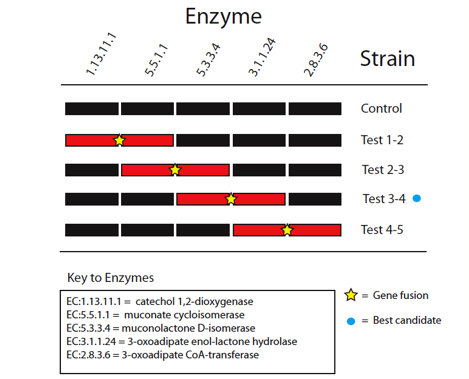
The five enzymes in the catechol degradation pathway not naturally present in E. coli will be introduced into E. coli as BioBrick parts. The parts will then be used to construct five recombinant strains of E. coli (Figure 3). These include a full reconstruction of the pathway (control strain) and several partial reconstructions (test strains). The latter will contain pathway 'holes' corresponding to all combinations of sequential pairs of enzymes, with complementary fusion pairs. The test strain with the fusion of enzymes 3 and 4 will be of prime interest, while the other test strains will be used validate the predictions made by Cell++.
Figure 3: Diagramatic representation of proposed gene fusion experiments. Pathway reconstructions are shown in black for the control and test strains with pathway holes for the partial reconstructions shown in red. Yellow stars mark proposed gene fusions to be tested to validate computational predictions. The blue circle marks the best candidate for enzyme channeling based on in silico experiments
In order to demonstrate the breakdown of catechol by these recombinant strains, catechol will be added into the growth media of these strains. A colorimetric assay, previously developed to quantify the amount of catechol in solution, will be utilized to assay the rate of catechol degradation by the different strains (Tao et al.). In addition, a survival assay will be used to determine the threshold for catechol exposure for non recombinant E. coli and to evaluate the effect that the recombinantly-expressed enzymes in the catechol pathway will have on E. coli survival.
Future Goals
Once we complete the characterization of the “Encapsulator” micro-compartment from the 2009 iGEM project, we will evaluate whether compartmentalization can further enhance the effects of metabolic channeling as compared to gene fusions. Ultimately, channeling will be introduced to target pathways in endogenous tailings ponds species (e.g. P. Putida) for application in the bioremediation of the tailings ponds.
 "
"