Team:TU Delft/project/rbs characterization
From 2010.igem.org
RBS Characterization
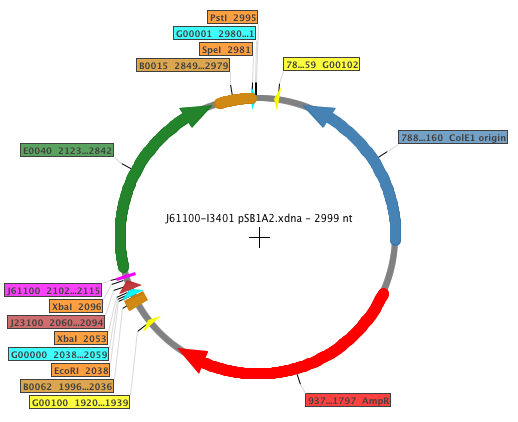
For our RBS characterization project, five different RBS sequences from the Anderson RBS family (J61100, J61101, J61107, J61117, J61127) and the standard RBS B0032 were placed in front of the standard GFP coding sequence. The Biobricks generated in order to perform the experiments were: K398500, K398501, K398502, K398503, K398504. The general map of the construction is shown below, where the RBS is displayed in fucsia.
| Feature | Function |
| AmpR | Ampicillin resistance |
| B0015 | Transcriptional (double) terminator |
| B0062 | Transcriptional terminator |
| E0040 | GFP |
| G00000 | Standard prefix |
| G00001 | Standard suffix |
| G00100 | VF2 primer binding site |
| G00101 | VR primer binding site |
| J61100 | RBS Anderson family |
| J23100 | Promoter |
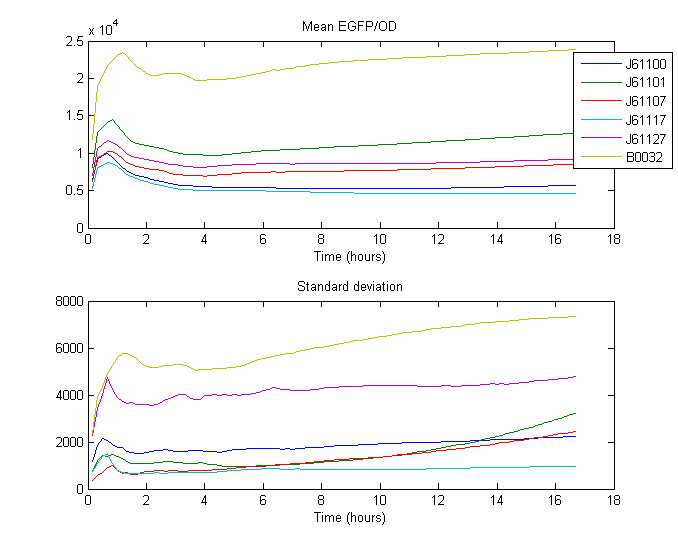
Note: Fluorescence (y-axis) is reported as Arbitrary Fluorescence Units.
The RBS strength was calculated by taking the mean of the ratio between the expression of the standard RBS (B0032) and expression of Anderson RBS over time. The Expression is defined as the quotient of the measured fluorescence divided by measured biomass (OD at 600nm).
| RBS | Strength |
| J61100 | 0.047513 |
| J61101 | 0.119831 |
| J61107 | 0.065454 |
| J61117 | 0.038518 |
| J61127 | 0.087334 |
| B0032 | 0.300000 |
The result of this experimental set up was a simple characterization of five of the Anderson RBS sequences in relation to other standard well-characterized RBS. The given relative strengths are displayed assuming B0034 as the unit.
We're also trying to look into mRNA folded shapes using mfold to see if there is a common pattern in the Anderson RBS shapes. This might be usable in predicting RBS strength for the other untested Anderson RBS sequences. Unfortunately, this seems not to be the case, as all the RBS sequences have very different mfold shapes.
 "
"