Team:TU Delft/project/rbs characterization
From 2010.igem.org
(Difference between revisions)
(→RBS Characterization) |
(→RBS Characterization) |
||
| Line 9: | Line 9: | ||
|GFP | |GFP | ||
|- | |- | ||
| - | |[http://partsregistry.org/Part: | + | |[http://partsregistry.org/Part:BBa_G00100 G00100] |
| - | | | + | |VF2 primer binding site |
|- | |- | ||
| - | |[http://partsregistry.org/Part: | + | |[http://partsregistry.org/Part:BBa_G00101 G00101] |
| - | | | + | |VR primer binding site |
|- | |- | ||
|[http://partsregistry.org/Part:BBa_J61117 J61117] | |[http://partsregistry.org/Part:BBa_J61117 J61117] | ||
Revision as of 13:01, 10 August 2010
RBS Characterization
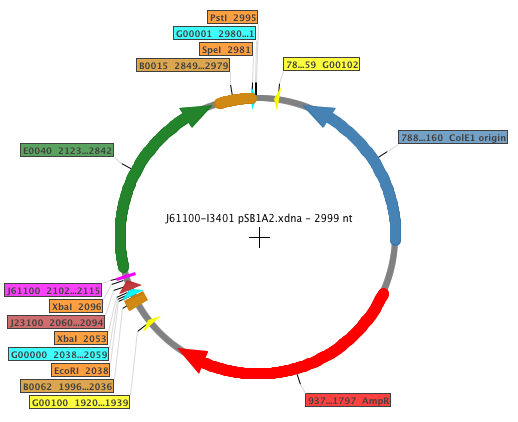
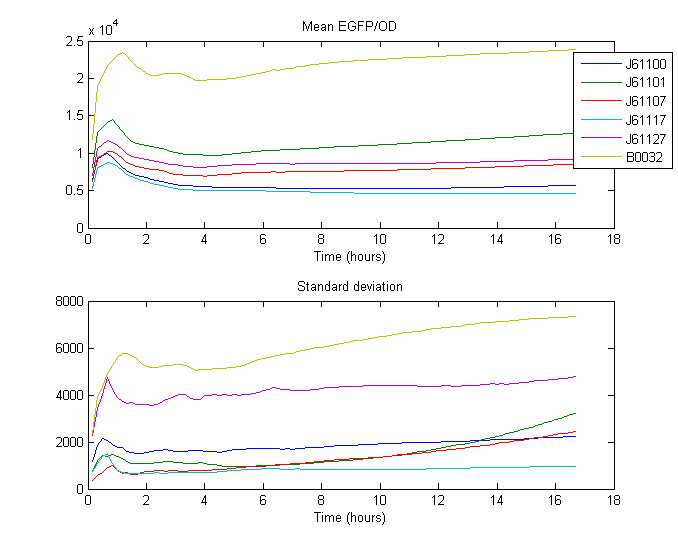
For our RBS characterization project, 5 different RBS sequences from the Anderson RBS family (J61100, J61101, J61107, J61117, J61127) and one known RBS (B0032) where placed in front of GFP and measured over 18 hours using a Gen5 fluorenscence and absorbance plate reader. The general map of the construction is shown below, where the RBS is displayed in fucsia.
| Feature | Function |
| E0040 | GFP |
| G00100 | VF2 primer binding site |
| G00101 | VR primer binding site |
| J61117 | 0.038518 |
| J61127 | 0.087334 |
| B0032 | 0.300000 |
| RBS | Strength |
| J61100 | 0.047513 |
| J61101 | 0.119831 |
| J61107 | 0.065454 |
| J61117 | 0.038518 |
| J61127 | 0.087334 |
| B0032 | 0.300000 |
We've also looked at mRNA folded shapes using mfold to see if there was a common pattern in the Anderson RBS shapes. This might have been usable in predicting RBS strength for the other untested Anderson RBS sequences. Unfortunately, this didn't seem to be the case, as all the tried RBS sequences had very different mfold shapes.
 "
"