Team:TU Delft/Protocols
From 2010.igem.org
(→General gas chromatography program for alkanes and alkanols) |
(→Protocols) |
||
| (53 intermediate revisions not shown) | |||
| Line 163: | Line 163: | ||
#Wash QIAprep spin column by adding 0.75 mL Buffer PE and centrifuging for 30–60 seconds. | #Wash QIAprep spin column by adding 0.75 mL Buffer PE and centrifuging for 30–60 seconds. | ||
#Discard the flow-through, and centrifuge for an additional 1 min to remove residual wash buffer. Important: Residual wash buffer will not be completely removed unless the flow-through is discarded before this additional centrifugation. Residual ethanol from Buffer PE may inhibit subsequent enzymatic reactions. | #Discard the flow-through, and centrifuge for an additional 1 min to remove residual wash buffer. Important: Residual wash buffer will not be completely removed unless the flow-through is discarded before this additional centrifugation. Residual ethanol from Buffer PE may inhibit subsequent enzymatic reactions. | ||
| - | #Place the QIAprep column in a clean 1.5 | + | #Place the QIAprep column in a clean 1.5 mL microcentrifuge tube. To elute DNA, add 30 μL Buffer EB (10 mM Tris·Cl, pH 8.5) or water to the center of each QIAprep spin column, let stand for 1 minute, and centrifuge for 1 minute. |
#Measure DNA concentration on the Nanodrop | #Measure DNA concentration on the Nanodrop | ||
| Line 289: | Line 289: | ||
Reaction for one sample: | Reaction for one sample: | ||
{| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | {| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | ||
| + | |'''Component''' | ||
| + | |'''Sample''' | ||
| + | |- | ||
|DNA | |DNA | ||
|x μL (up to 1,0 μg) | |x μL (up to 1,0 μg) | ||
| Line 342: | Line 345: | ||
Reaction for one sample: | Reaction for one sample: | ||
{| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | {| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | ||
| + | |'''Component''' | ||
| + | |'''Sample''' | ||
| + | |- | ||
|DNA insert | |DNA insert | ||
|x μL | |x μL | ||
| Line 399: | Line 405: | ||
#Keep solutions on ice | #Keep solutions on ice | ||
#Add to a thin walled PCR tube, on ice, for a 50 μL reaction: | #Add to a thin walled PCR tube, on ice, for a 50 μL reaction: | ||
| - | |||
1× pre-mix | 1× pre-mix | ||
{| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | {| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | ||
| Line 433: | Line 438: | ||
|} | |} | ||
| - | + | <html><ol start="4"<li>Gently vortex the sample and briefly centrifuge (5 sec) to collect all droplets at the bottom of the tube</li></ol></html> | |
PCR program: | PCR program: | ||
| Line 529: | Line 534: | ||
|} | |} | ||
| - | + | <html><ol start="4"<li>Gently vortex the sample and briefly centrifuge (5 sec) to collect all droplets at the bottom of the tube</li></ol></html> | |
PCR program: | PCR program: | ||
| Line 628: | Line 633: | ||
|} | |} | ||
| - | + | <html><ol start="4"<li>Add 25 μL of pre-mix to each PCR tube.</li> | |
| - | + | <li>Pick a sterile toothpick into a colony, dip it into a PCR tube, put it into a 15 mL culture tube containing 5 mL LB medium + antibiotics and then grow overnight for mini-prep cultures and -80 °C stocks. Repeat this for all the colonies. Incubate the mini-prep cultures at 37 °C.</li> | |
| - | + | <li>Keep everything ice cold until you put the tubes in the preheated PCR machine</ol></html> | |
PCR program: | PCR program: | ||
| Line 687: | Line 692: | ||
#After the gel slice has dissolved completely, check that the color of the mixture is yellow (similar to Buffer QG without dissolved agarose). If the color of the mixture is orange or violet, add 10 μL of 3 M sodium acetate, pH 5.0, and mix. The color of the mixture will turn to yellow. The adsorption of DNA to the QIAquick membrane is efficient only at pH <7.5. Buffer QG contains a pH indicator which is yellow at pH <7.5 and orange or violet at higher pH, allowing easy determination of the optimal pH for DNA binding. | #After the gel slice has dissolved completely, check that the color of the mixture is yellow (similar to Buffer QG without dissolved agarose). If the color of the mixture is orange or violet, add 10 μL of 3 M sodium acetate, pH 5.0, and mix. The color of the mixture will turn to yellow. The adsorption of DNA to the QIAquick membrane is efficient only at pH <7.5. Buffer QG contains a pH indicator which is yellow at pH <7.5 and orange or violet at higher pH, allowing easy determination of the optimal pH for DNA binding. | ||
#Add 1 gel volume of isopropanol to the sample and mix. This step increases the yield of DNA fragments <500 bp and >4 kb. For DNA fragments between 500 bp and 4 kb, addition of isopropanol has no effect on yield. Do not centrifuge the sample at this stage. | #Add 1 gel volume of isopropanol to the sample and mix. This step increases the yield of DNA fragments <500 bp and >4 kb. For DNA fragments between 500 bp and 4 kb, addition of isopropanol has no effect on yield. Do not centrifuge the sample at this stage. | ||
| - | #Place a QIAquick spin column in a provided 2 | + | #Place a QIAquick spin column in a provided 2 mL collection tube. |
#To bind DNA, apply the sample to the QIAquick column, and centrifuge for 1 min. The maximum volume of the column reservoir is 800 μL. For sample volumes of more than 800 μL, simply load and spin again. | #To bind DNA, apply the sample to the QIAquick column, and centrifuge for 1 min. The maximum volume of the column reservoir is 800 μL. For sample volumes of more than 800 μL, simply load and spin again. | ||
#Discard flow-through and place QIAquick column back in the same collection tube. | #Discard flow-through and place QIAquick column back in the same collection tube. | ||
| - | #To wash, add 0.75 | + | #To wash, add 0.75 mL of Buffer PE to QIAquick column and centrifuge for 1 min. |
#Discard the flow-through and centrifuge the QIAquick column for an additional 1 min at 10,000 x g (~13,000 rpm). IMPORTANT: Residual ethanol from Buffer PE will not be completely removed unless the flow-through is discarded before this additional centrifugation. | #Discard the flow-through and centrifuge the QIAquick column for an additional 1 min at 10,000 x g (~13,000 rpm). IMPORTANT: Residual ethanol from Buffer PE will not be completely removed unless the flow-through is discarded before this additional centrifugation. | ||
| - | #Place QIAquick column into a clean 1.5 | + | #Place QIAquick column into a clean 1.5 mL microcentrifuge tube. |
#To elute DNA, add 50 μL of Buffer EB (10 mM Tris·Cl, pH 8.5) or H2O to the center of the QIAquick membrane and centrifuge the column for 1 min at maximum speed. Alternatively, for increased DNA concentration, add 30 μl elution buffer to the center of the QIAquick membrane, let the column stand for 1 min, and then centrifuge for 1 min. Important: Ensure that the elution buffer is dispensed directly onto the QIAquick membrane for complete elution of bound DNA. The average eluate volume is 48 μL from 50 μL elution buffer volume, and 28 μL from 30 μL. Elution efficiency is dependent on pH. The maximum elution efficiency is achieved between pH 7.0 and 8.5. When using water, make sure that the pH value is within this range, and store DNA at –20°C as DNA may degrade in the absence of a buffering agent. The purified DNA can also be eluted in TE (10 mM Tris·Cl, 1 mM EDTA, pH 8.0), but the EDTA may inhibit subsequent enzymatic reactions. | #To elute DNA, add 50 μL of Buffer EB (10 mM Tris·Cl, pH 8.5) or H2O to the center of the QIAquick membrane and centrifuge the column for 1 min at maximum speed. Alternatively, for increased DNA concentration, add 30 μl elution buffer to the center of the QIAquick membrane, let the column stand for 1 min, and then centrifuge for 1 min. Important: Ensure that the elution buffer is dispensed directly onto the QIAquick membrane for complete elution of bound DNA. The average eluate volume is 48 μL from 50 μL elution buffer volume, and 28 μL from 30 μL. Elution efficiency is dependent on pH. The maximum elution efficiency is achieved between pH 7.0 and 8.5. When using water, make sure that the pH value is within this range, and store DNA at –20°C as DNA may degrade in the absence of a buffering agent. The purified DNA can also be eluted in TE (10 mM Tris·Cl, 1 mM EDTA, pH 8.0), but the EDTA may inhibit subsequent enzymatic reactions. | ||
| Line 702: | Line 707: | ||
*96% ethanol | *96% ethanol | ||
*70% cold ethanol | *70% cold ethanol | ||
| - | * | + | *milliQ |
*microcentrifuge | *microcentrifuge | ||
*heat block at 37 °C | *heat block at 37 °C | ||
| Line 757: | Line 762: | ||
'''Materials:''' | '''Materials:''' | ||
*2 mg/mL BSA | *2 mg/mL BSA | ||
| - | *bradford | + | *bradford reagents |
*spectrophotometer | *spectrophotometer | ||
'''Protocol:''' | '''Protocol:''' | ||
| - | #Make a dilution serie of standard 2 mg/mL bovine serum albumin (BSA). Use 0 | + | #Make a dilution serie of standard 2 mg/mL bovine serum albumin (BSA). Use 0, 20, 40, 60, 80, 100 μL |
#Add MillQ to a total volume of 800 μL | #Add MillQ to a total volume of 800 μL | ||
#Add 200 μL Bradford reagens and mix | #Add 200 μL Bradford reagens and mix | ||
#Measure after minimal 5 min and max 60 min the OD595 | #Measure after minimal 5 min and max 60 min the OD595 | ||
| - | |||
| - | ===Emulsifier | + | |
| + | ===Emulsifier Assay=== | ||
'''Materials:''' | '''Materials:''' | ||
| Line 797: | Line 802: | ||
*Shaking incubator | *Shaking incubator | ||
*Sonicator (Branson Sonifier 250) | *Sonicator (Branson Sonifier 250) | ||
| - | * | + | *centrifuge |
| - | + | ||
'''Protocol''' | '''Protocol''' | ||
| - | #Pick a single colony from a freshly streaked selective plate and inoculate a starter culture of 50 mL LB medium containing the appropriate selective antibiotic. Incubate at 37°C, 160 rpm. | + | #Pick a single colony from a freshly streaked selective plate and inoculate a starter culture of 50 mL LB medium containing the appropriate selective antibiotic. Incubate at 37°C, 160 rpm. |
| - | * | + | #*''Chef's tips and tricks'': |
| - | **You can also inoculate directly from your cell bank (cell stock with glycerol 40%v/v in fresh medium). Then you can | + | #** You can also inoculate directly from your cell bank (cell stock with glycerol 40%v/v in fresh medium). Then you can start with a previous overnight culture (5 mL) and then inoculate 50 mL of fresh medium with 1 mL, in that way you will reach your desired OD around 2-3 hours you started the culture. |
| - | **Normally, we use 5 | + | #** Normally, we use 5 μL to inoculate 5 mL. Some people may think that this is a lot, specially because of the fact that cells are concentrated in the cell bank. .But, when we used a different amount we had some nasty surprises like no overnight growth. Spin down the cultures at an OD of +/- 0.6 in 50 mL tubes for 10 min at 4.000 rpm using a chilled (4 °C) centrifuge. |
| - | + | #Resuspend the cells in 5 mL of 50 mM Tris buffer (with the appropriate pH) and keep on ice until sonication. | |
| - | + | #*''Chef's tips and tricks'': | |
| - | #Resuspend the cells in 5 mL of 50 mM Tris buffer (with the appropriate pH) and keep on ice until sonication. | + | #**Sometimes we use a washing step, specially when we want to prepare a extract of 100 mL of culture (2 flasks of 50 mL). Then after you resuspend your cells, spin them down and resuspend them again. If you think that this could be futile check the amount of LB remaining next your cells, it's around 200 μL. Just in case you don't want this extra source of pH disturbance, you can do this washing step before you kill your cells. |
| - | * | + | #Sonicate the suspension (placing the tip as close as possible to the bottom of the tube, without touching) for 3 minutes at a 40% duty cycle and an output control of 4; all the while keeping the suspension on ice. |
| - | **Sometimes we use a washing step, specially when we want to prepare a extract of | + | #*''Chef's tips and tricks'': |
| - | + | #**BE CAREFUL, place the tip of your sonicator in the middle of your tube next to the bottom, but not touching the tube otherwise you can break it. If you place the tip somewhere else, you will end up with a nice ''E. coli'' milkshake with zero acitivity. Presence of a lot of foam during the disruption is bad sign; that nice foam it's mainly denaturated protein. So please, BE CAREFUL. | |
| - | #Sonicate the suspension (placing the tip as close as possible to the bottom of the tube, without touching) for 3 minutes at a 40% duty cycle and an output control of 4; all the while keeping the suspension on ice. | + | #**You need ice all the time. Sonication produces a lot of heat, if you have a heat sensitive protein or you don't want to mess up with you negative control (''E. coli''), then KEEP YOUR CELL SUSPENTION ALL THE TIME ON ICE. |
| - | * | + | #**If you see a nice reduction of O.D. by naked eye inspection, then you are in the good way. If that's not the case you can give to the cell suspension some extra minutes (1 or 2) in the sonicator. There should be a nice difference between disrupted cells and normal cell suspension. Disrupted cells produce a transparent(ish) suspension, whereas normal cells have a milky aspect. |
| - | **BE CAREFUL, place the tip of your sonicator in the middle of your tube next to the bottom, but not touching the tube otherwise you can break it. If you place the tip somewhere else, you will end up with a nice | + | #Spin down the suspension for 5 minutes at 4.000 rpm in a chilled centrifuge. |
| - | ** | + | #*''Chef's tips and tricks'': |
| - | **If you see a nice reduction of O.D. by naked eye inspection, then you are in the good way. If that's not the case you can give to the cell suspension some extra minutes (1 or 2) in the sonicator. There should be a nice difference between disrupted cells and normal cell suspension. Disrupted cells produce a transparent(ish) suspension, whereas normal cells have a milky aspect. | + | #**You can see a nice reduction of you cell pellet, and that could be use as a criterion for keep on going with your experiments, sometimes the reduction of the pellet is not that considerable but you may see a viscous supernatant (compared to your buffer). That's a sign of a nice disruption; if you cannot see these things happening to your suspension then something went wrong. You can do an extra sonication step. |
| + | #Your supernatant now contains your enzyme of interest. You can do protein determination assays. ''Recomendation'': DON'T BE TOO GREEDY!!! If you have some protein don't do an extra sonication step in order to obtain more. If you make a mistake or you give extra energy to your extract you may lose the sample and you have to start all over again. | ||
| - | + | '''Evaluating''' | |
| - | + | ||
| - | + | ||
| - | + | ''Use the following checking list in order to know if you have to do an extra disruprtion step or start all over again:'' | |
| - | + | ||
| - | Use the following checking list in order to know if you have to do an extra disruprtion step or start all over again: | + | |
# Is the pellet is smaller compared to the one obtained before disruption? If yes continue to 2. | # Is the pellet is smaller compared to the one obtained before disruption? If yes continue to 2. | ||
# Did the supernatant look like more viscous than water when I transfer the supernatant?If yes continue to 3. | # Did the supernatant look like more viscous than water when I transfer the supernatant?If yes continue to 3. | ||
| Line 829: | Line 830: | ||
# Do the bradford or protein analysis results show that I have more signal in my disruption cells than in the control (buffer)? If yes go to Successful disruption and smile, if not go to Extra step and cry if you feel like that. | # Do the bradford or protein analysis results show that I have more signal in my disruption cells than in the control (buffer)? If yes go to Successful disruption and smile, if not go to Extra step and cry if you feel like that. | ||
| - | + | '''Troubleshooting''' | |
| + | #''Extra step'': Your disruption went really wrong, repeat the sonication step and use the checklist again. If you still have problems (like no protein or a mediocre amount of protein) then ask the person in charge of the sonicator or someone more qualified than you to check the machine. | ||
| + | #''Doubtful results'': Sometimes your activities are lower than they are in reality, which will give you a false positive in case that you want to use your protein for activity assays. If this is your first time with the protocol, don't be scared; keep going on with your tests but take into consideration the advice that YOU HAVE TO REPEAT IT in order to proof that your activities are correct. You can use two or three ''E. coli'' cultures in the same experiment, so that if you mess up with one you have other(s) ready to be used as a confirmation of your results. | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
===Tolerance assay=== | ===Tolerance assay=== | ||
'''Materials''' | '''Materials''' | ||
| - | *Overnight culture | + | *Overnight culture |
*LB medium with the appropriate antibiotics | *LB medium with the appropriate antibiotics | ||
| - | *15 | + | *15 mL tubes |
| - | + | *Appropriate concentration of the agitant (e.g. salt or hydrocarbon) | |
| - | *Appropriate concentration of the agitant (e.g. salt or hydrocarbon) | + | |
*96 well plate | *96 well plate | ||
*Plate reader | *Plate reader | ||
'''Protocol''' | '''Protocol''' | ||
| - | #Subculture 10 | + | #Subculture 10 μL of the overnight culture into 5 mL of LB (with antibiotic) in a 15 mL tube. |
#Grow to an OD600 of 0.3-0.4. | #Grow to an OD600 of 0.3-0.4. | ||
#Dilute back to an OD600 of 0.1. | #Dilute back to an OD600 of 0.1. | ||
#Store on ice. | #Store on ice. | ||
| - | #Load the 96 well plate with LB (with antibiotic | + | #Load the 96 well plate with LB (with antibiotic) and the agitant you wish to measure. Note: do all the experiments in triplicate. |
#Measure the growth curves using the plate reader | #Measure the growth curves using the plate reader | ||
| + | |||
===Alcohol/Aldehyde dehydrogenase activity assays=== | ===Alcohol/Aldehyde dehydrogenase activity assays=== | ||
| + | |||
'''Materials''' | '''Materials''' | ||
| - | + | *Cell extracts | |
| - | *Cell extracts | + | *Buffer 57 mM glycine, 1mM NAD, pH 9.5 |
| - | *Buffer 57 mM glycine 1mM NAD pH 9.5 | + | *Alcohol/Aldehyde |
| - | *Alcohol/Aldehyde | + | |
*96 well plate | *96 well plate | ||
*Plate reader | *Plate reader | ||
'''Protocol''' | '''Protocol''' | ||
| + | #Determine the protein content (mg/mL) of your cell extract. Use whatever method you prefer. | ||
#Keep your extracts on ice. | #Keep your extracts on ice. | ||
| - | #Add buffer to the wells that you are going to use. | + | #Add glycine buffer to the wells that you are going to use, usually we used 180 μL per well. |
| - | #Add the alcohol/aldehyde to be tested to the appropriate wells. | + | #Add the alcohol/aldehyde to be tested to the appropriate wells, usually we added 5 μL. ''WARNING'': Each well with alcohol/aldehyde MUST HAVE A CONTROL WITHOUT SUBSTRATE. Don't forget to prepare a BLANK, which will be a mixture of buffer and substrate without enzyme. |
| - | + | #*''Chef's tips and tricks'': | |
| - | #Warm the plate up for 10-20 min. Keep in mind that this is a biphasic system; therefore we need a previous step in which you let the system to reach a equilibrium | + | #**Long-chain alcohols are not in a liquid form at room temperature, keep them in some warm place before you start your assay, so that they will be melted whenever you will use them. |
| + | #Warm the plate up for 10-20 min at 37ºC. Keep in mind that this is a biphasic system; therefore we need a previous step in which you let the system to reach a equilibrium. Otherwise your first measurements will be noisy and you have to extend the assay, in case that you have a really powerful enzyme that may mean that you will repeat the experiment because the NAD will dissappear very quick. In case that your enzyme is not that fast, you can skip this step and add the enzyme; if you have noisy results at the beggining of the measurement, discard them and analyze the rest (at least 20 min of analysis, preferable more than 5 data points). | ||
| + | #Add the cell extract to your reaction mixture, usually we added 40 uL. | ||
| + | #Measure in your plate reader at wavelength of 340 nm, it is better if you run your experiment for 1 hour or so. Try to do data acquisition each 2-3 minutes, in that way you will have more than enough data to work with. | ||
| + | |||
| + | '''Data analysis''' | ||
| + | #Divide each data point by the extinction coefficient of NADH (6220 M^-1 cm^-1) and by the path length; in our case we had reaction volumes of 225 μL (buffer+enzyme+substrate) and 220 μL (buffer+enzyme). Each well had a length of 1 cm and a total volume of 300 μL. That means that we had path lengths of 0.75 cm and 0.7333 cm. The number that you will get is the MOLAR CONCENTRATION of NADH in you well. | ||
| + | #Multiply the molar concentration of NADH in your well by the reaction volume in order to get the TOTAL AMOUNT OF NADH MOLES in your well. | ||
| + | #Rest the NADH moles from the BLANK (buffer+substrate WITHOUT enzyme) to all your wells. | ||
| + | #For each data point do the following operation: NADH moles produced = NADH moles (buffer+enzyme+substrate) - NADH moles (buffer+enzyme) | ||
| + | #Plot on the x-axis the time and in the y-axis the NADH moles produced. Perform a linear regression and report the slopes (moles/min) of each well. One Unit of alcohol/aldehyde dehydrogenase is defined as the amount of enzyme required for producing 1 umol of NADH in one minute. | ||
| + | #Divide the amount of Units of enzyme by the amount of TOTAL PROTEIN added to the reaction mixture. That will give you a result in U/mg of protein. | ||
| + | |||
===RBS characterization experiment=== | ===RBS characterization experiment=== | ||
| + | |||
'''Materials''' | '''Materials''' | ||
*-80 stocks or plates of strains carrying the RBS of interest, a reference and an 'empty' plasmid | *-80 stocks or plates of strains carrying the RBS of interest, a reference and an 'empty' plasmid | ||
| Line 884: | Line 892: | ||
'''Protocol''' | '''Protocol''' | ||
#Inoculate 5 mL of LB+Amp with your strains of interest, reference strain (B0032) and empty plasmid strain (I13401). Do this in triplicate. Grow up o/n. | #Inoculate 5 mL of LB+Amp with your strains of interest, reference strain (B0032) and empty plasmid strain (I13401). Do this in triplicate. Grow up o/n. | ||
| - | #The following day subculture 2 | + | #The following day subculture 2 μL of o/n culture in 200 μL of LB+Amp (1:100 dilution) on 96 well plates, all in quadruplicate (you will have 12 wells per strain now, a full row!) |
#Culture the cells up to an OD of about 0.3 and dilute 1:100 onto another 96 well plates in the same order as before usig LB+Amp. | #Culture the cells up to an OD of about 0.3 and dilute 1:100 onto another 96 well plates in the same order as before usig LB+Amp. | ||
#Hurry your plate over to the plate reader and start measuring GFP fluorescence and biomass absorbance! | #Hurry your plate over to the plate reader and start measuring GFP fluorescence and biomass absorbance! | ||
#The plate is read by an automatically repeating protocol with shaking at medium speed. The program does the following: | #The plate is read by an automatically repeating protocol with shaking at medium speed. The program does the following: | ||
| - | *Sets the temperature to 37°C | + | #*Sets the temperature to 37°C |
| - | *In a kinetic loop of a fixed time (16 hour 30 mins) the following measurements are taken with a time interval of 10 minutes with shaking: absorbance (600 nm filter) and GFP fluorescence (485 nm excitation and 520 nm emission). | + | #*In a kinetic loop of a fixed time (16 hour 30 mins) the following measurements are taken with a time interval of 10 minutes with shaking: absorbance (600 nm filter) and GFP fluorescence (485 nm excitation and 520 nm emission). |
| - | *followed by a delay of 100 seconds. | + | #*followed by a delay of 100 seconds. |
| + | |||
| + | |||
| + | ===pCaiF characterization experiment=== | ||
| + | |||
| + | '''Materials''' | ||
| + | *-80 stocks or plates of strains carrying the biobrick BBa_K398331, a reference and an 'empty' plasmid | ||
| + | *96-well plate + lid | ||
| + | *96-well plate fluorescence + absorbance reader (BioTek Synergy) | ||
| + | |||
| + | '''Protocol''' | ||
| + | #Inoculate 5 mL of M9 (10 g/L of glc) with your strains of interest and empty plasmid strain (J13002). Grow them overnight. | ||
| + | #*''Chef's tips and tricks'': | ||
| + | #**Be aware that normal M9 contains 2 g/L. In order to prepare this new medium we just added 5 times the specified amount of glucose for normal M9 medium. | ||
| + | #In the next morning subculture 50 μL of o/n culture in another 5 mL of M9 (10 g/L of glc). | ||
| + | #Check the O.D. at 600nm of the cultures after some hours (usually 3 h), if this parameter is in the range of 0.15-0.3 you can start your assay, otherwise wait until you have at least an O.D. of 0.2. | ||
| + | #*''Chef's tips and tricks'': | ||
| + | #**We use to check the O.D. 2 hours after we started the culture and we try to adjust the O.D. to the lowest measurement obtained, once they are in the same O.D. when they are diluted the greater are the chances that you don't have to adjust the O.D. at the end of the culture. | ||
| + | #Pour into your plate 100 μL of fresh medium, use different glucose concentrations. You can do this by diluting your M9 medium (10g/L glc) with M9 WITHOUT glucose. | ||
| + | #*''Chef's tips and tricks'': | ||
| + | #**In our case we used 10 g/L, 5 g/L, 2 g/L, 1 g/L with a secondary carbon source: Laurate (final concentration of 5mM). | ||
| + | #Add 5 μL of you culture according to your personal scheme. | ||
| + | #Now your plate is ready to be measured! Go to your favorite plate reader and program your sample. Set the temperature to 37°C. In a kinetic loop of a fixed time (17 hour 30 mins) the following measurements are taken with a time interval of 10 minutes with CONSTANT shaking: absorbance (600 nm filter) and GFP fluorescence (485 nm excitation and 520 nm emission). Followed by a delay of 100 seconds. | ||
| + | |||
| + | '''Data analysis''' | ||
| + | #Average you Biomass concentration for empty plasmid and your sample. Substract the blank value to your measurements. | ||
| + | #Average the fluorescence produced by your empty plasmid and your sample. Substract the fluorescence of the empty plasmid to the fluorescence given by your strain of interest. The BLANK is already included in this operation if it is all the time lower than the fluorescence measured in your empty plasmid. | ||
| + | #Plot the data points obtained and analyze the amount of GFP produced by your strain. | ||
| + | |||
===Protocol for growth of ''E.coli'' K12 on alkanes=== | ===Protocol for growth of ''E.coli'' K12 on alkanes=== | ||
| + | |||
'''Materials''' | '''Materials''' | ||
| - | *-80 | + | *-80 °C stock or plate containing strain of interest |
*25 mL closed-cap glass flasks | *25 mL closed-cap glass flasks | ||
*The to-be-tested alkanes | *The to-be-tested alkanes | ||
| Line 903: | Line 940: | ||
'''Protocol''' | '''Protocol''' | ||
#Inoculate 5 mL of LB+antibiotic with your strains of interest. Grow up o/n. | #Inoculate 5 mL of LB+antibiotic with your strains of interest. Grow up o/n. | ||
| - | #The following day subculture 500 | + | #The following day subculture 500 μL of o/n culture in 50 mL of LB+antibiotic (1:100 dilution). |
#Culture the cells up to an OD of about 0.3, spin down (4000 rpm, 10 min) and resuspend in 5 mL of wash buffer. | #Culture the cells up to an OD of about 0.3, spin down (4000 rpm, 10 min) and resuspend in 5 mL of wash buffer. | ||
| - | #Spin down (4000 rpm, 5 min), resuspend in 5 mL P-buffer and subculture 50 | + | #Spin down (4000 rpm, 5 min), resuspend in 5 mL P-buffer and subculture 50 μL into 5 mL of M9-medium + E2 salts + 1% alkane. |
#Measure OD600 at the start and compare to OD600 attained in time. | #Measure OD600 at the start and compare to OD600 attained in time. | ||
| + | |||
===Resting-cell assays for ''E.coli''=== | ===Resting-cell assays for ''E.coli''=== | ||
| + | |||
'''Materials''' | '''Materials''' | ||
| - | *-80 | + | *-80 °C stock or plate containing strain of interest |
| - | *100 | + | *100 mL sterile erlenmeyer flasks |
*25 mL closed-cap glass flasks | *25 mL closed-cap glass flasks | ||
*The to-be-tested alkanes | *The to-be-tested alkanes | ||
*LB-medium + antibiotic | *LB-medium + antibiotic | ||
*Phosphate buffer of pH 7.4 (aka wash buffer) | *Phosphate buffer of pH 7.4 (aka wash buffer) | ||
| - | *Phosphate buffer of pH 7.4 + E2 salts + 0.66% v/v glycerol ( | + | *Phosphate buffer of pH 7.4 + E2 salts + 0.66% v/v glycerol (also known as RSA buffer) |
| - | *Incubator | + | *Incubator at 37 °C |
'''Protocol''' | '''Protocol''' | ||
| - | + | ||
| + | ''Don't forget to think about sterile conditions where needed!'' | ||
#Inoculate 5 mL of LB+antibiotic with your strains of interest. Grow up o/n. | #Inoculate 5 mL of LB+antibiotic with your strains of interest. Grow up o/n. | ||
| - | #The following day subculture 500 | + | #The following day subculture 500 μL of o/n culture in 50 mL of LB+antibiotic (1:100 dilution). |
| - | #Culture the cells up to an OD of about 0.3, spin down ( | + | #Culture the cells up to an OD of about 0.3, spin down (4.000 rpm, 10 min) and resuspend in 5 mL of wash buffer. |
#Spin down (4000 rpm, 5 min) and resuspend in 20 mL of RSA buffer. | #Spin down (4000 rpm, 5 min) and resuspend in 20 mL of RSA buffer. | ||
#Measure the OD600 (for reference to the OD600 at the end of the experiment). | #Measure the OD600 (for reference to the OD600 at the end of the experiment). | ||
#Make cell-mixture aliquots of 6 mL in 25 mL closed-cap glass flasks and add 100 nmoles of the desired alkane. | #Make cell-mixture aliquots of 6 mL in 25 mL closed-cap glass flasks and add 100 nmoles of the desired alkane. | ||
| - | #Incubate the mixtures at the optimal enzyme temperature (below 37 | + | #Incubate the mixtures at the optimal enzyme temperature (below 37 °C!) for the desired reaction time |
#Measure the OD600 afterwards. | #Measure the OD600 afterwards. | ||
| + | |||
===Ethyl acetate extraction protocol=== | ===Ethyl acetate extraction protocol=== | ||
| + | |||
'''Materials''' | '''Materials''' | ||
*Aqueous mixture from which hydrocarbons will be extracted | *Aqueous mixture from which hydrocarbons will be extracted | ||
| Line 939: | Line 981: | ||
'''Protocol''' | '''Protocol''' | ||
| - | + | ||
| + | ''Make sure all steps are performed using glassware! EtOAc dissolves plastics'' | ||
#Prepare adequate amounts of EtOAc + 0.1% internal standard. | #Prepare adequate amounts of EtOAc + 0.1% internal standard. | ||
#Add 2.5 mL of the above preparation to 6 mL of the aqueous mixture | #Add 2.5 mL of the above preparation to 6 mL of the aqueous mixture | ||
| Line 945: | Line 988: | ||
#If a detergent (such as Triton-X100) had been added to the aqueous mixture it is recommended to spin down the samples for 10 minutes @4000 rpm in order to get a proper two-phasic system | #If a detergent (such as Triton-X100) had been added to the aqueous mixture it is recommended to spin down the samples for 10 minutes @4000 rpm in order to get a proper two-phasic system | ||
#Remove the organic layer (top) and dry the solvent using MgSO4 | #Remove the organic layer (top) and dry the solvent using MgSO4 | ||
| - | #Filter off the MgSO4 using a 0.2 micron filter and store at -20 | + | #Filter off the MgSO4 using a 0.2 micron filter and store at -20 °C in GC-vials. |
| + | |||
===General gas chromatography program for alkanes and alkanols=== | ===General gas chromatography program for alkanes and alkanols=== | ||
| + | |||
'''Materials''' | '''Materials''' | ||
*Hydrocarbons in EtOAc to be analysed (dried over MgSO4) | *Hydrocarbons in EtOAc to be analysed (dried over MgSO4) | ||
*GC-vials | *GC-vials | ||
| - | *GC-machine with CP- | + | *GC-machine with CP-SIL 5CB column (length= 50 m) |
*Operation system to run the machine | *Operation system to run the machine | ||
'''Protocol''' | '''Protocol''' | ||
| + | |||
| + | ''Don't forget to fill the rinse vials with acetonitrile and methanol before use!'' | ||
#Place GC-vials containing sample in the proper position on the GC rack. | #Place GC-vials containing sample in the proper position on the GC rack. | ||
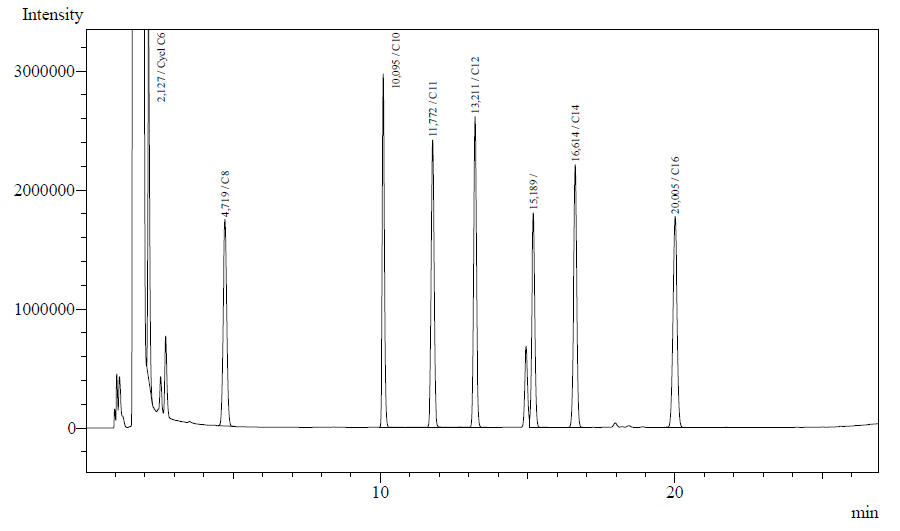
| - | #The | + | #The column temperature program (shown below) has yielded nice peak separations for the alkanes/alkanols we've used. The calibration GC graph for the alkanes we've used is given below. |
| - | {| | + | #Run batch and save results as pdf. |
| + | [[Image:TUDelft_Calibration_graph.png|500px|thumb|left|GC-calibration curve for 0.125% of alkanes used for our project]] | ||
| + | {| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | ||
|<b>Rate</b> | |<b>Rate</b> | ||
|<b>Temperature [C]</b> | |<b>Temperature [C]</b> | ||
| Line 972: | Line 1,021: | ||
|50 | |50 | ||
|110 | |110 | ||
| - | |2 | + | |2.0 |
|- | |- | ||
|50 | |50 | ||
|130 | |130 | ||
| - | |2 | + | |2.0 |
|- | |- | ||
|50 | |50 | ||
|145 | |145 | ||
| - | |2 | + | |2.0 |
|- | |- | ||
|50 | |50 | ||
|160 | |160 | ||
| - | |2 | + | |2.0 |
|- | |- | ||
|50 | |50 | ||
|170 | |170 | ||
| - | |2 | + | |2.0 |
|- | |- | ||
|50 | |50 | ||
|185 | |185 | ||
| - | |2 | + | |2.0 |
|- | |- | ||
|50 | |50 | ||
|210 | |210 | ||
| - | |2 | + | |2.0 |
|- | |- | ||
|50 | |50 | ||
|250 | |250 | ||
| - | |2 | + | |2.0 |
|- | |- | ||
|50 | |50 | ||
|320 | |320 | ||
| - | |1 | + | |1.0 |
|} | |} | ||
===Enzyme activity assay for LadA by GC=== | ===Enzyme activity assay for LadA by GC=== | ||
| + | |||
'''Materials''' | '''Materials''' | ||
*Lysates obtained from desired strains using our [https://2010.igem.org/Team:TU_Delft#Preparing_cell_lysates_for_enzyme_kinetics_measurements sonication protocol] | *Lysates obtained from desired strains using our [https://2010.igem.org/Team:TU_Delft#Preparing_cell_lysates_for_enzyme_kinetics_measurements sonication protocol] | ||
| Line 1,016: | Line 1,066: | ||
*Triton X-100 | *Triton X-100 | ||
*25 mL closed-cap flasks | *25 mL closed-cap flasks | ||
| - | *Incubator at 37 and/or at 60 | + | *Incubator at 37 °C and/or at 60 °C |
*Boiling water | *Boiling water | ||
*Desired alkane (0.1% of total volume) | *Desired alkane (0.1% of total volume) | ||
'''Protocol''' | '''Protocol''' | ||
| - | + | ||
| + | ''Make sure all steps are performed using glassware! Alkanes may dissolve plastics.'' | ||
#Add 0.1% v/v of the alkane to the Tris buffer and heat in boiling water for 5 minutes. Sonicate for 1 minute while still warm until a homogenous, viscous mixture is obtained. | #Add 0.1% v/v of the alkane to the Tris buffer and heat in boiling water for 5 minutes. Sonicate for 1 minute while still warm until a homogenous, viscous mixture is obtained. | ||
#Add up to 1mM of NADH, 1mM FMN, 1mM MgSO4 and 0.01& v/v Triton X-100 | #Add up to 1mM of NADH, 1mM FMN, 1mM MgSO4 and 0.01& v/v Triton X-100 | ||
#Make 6 mL aliquots in 25 mL closed-cap flasks | #Make 6 mL aliquots in 25 mL closed-cap flasks | ||
| - | #Add adequate amounts of lysates (depends on Bradford assays, | + | #Add adequate amounts of lysates (depends on Bradford assays, 1 mL taken for our experiments) |
#Extract using the [https://2010.igem.org/Team:TU_Delft#Ethyl_acetate_extraction_protocol extraction protocol] after desired reaction time (o/n in our experiments). | #Extract using the [https://2010.igem.org/Team:TU_Delft#Ethyl_acetate_extraction_protocol extraction protocol] after desired reaction time (o/n in our experiments). | ||
===Enzyme activity assay for LadA by NADH absorption=== | ===Enzyme activity assay for LadA by NADH absorption=== | ||
| + | |||
'''Materials''' | '''Materials''' | ||
| - | |||
*Cell extracts of desired strain | *Cell extracts of desired strain | ||
*Buffer 50 mM Tris containing 1mM NADH and 100nM FMN at pH 7.5 | *Buffer 50 mM Tris containing 1mM NADH and 100nM FMN at pH 7.5 | ||
| - | *Alkane ( | + | *Alkane (30 μL per well of heptadecane or hexadecane) |
*96 well plate | *96 well plate | ||
*Plate reader | *Plate reader | ||
| Line 1,040: | Line 1,091: | ||
#Keep your extracts on ice. | #Keep your extracts on ice. | ||
#Add buffer to the wells that you are going to use. | #Add buffer to the wells that you are going to use. | ||
| - | #Add the alkane to be tested to the appropriate wells.KEEP IN MIND: Each well with alkane MUST HAVE A CONTROL WITHOUT SUBSTRATE. Besides you must have a blank (mixture without enzyme). | + | #Add the alkane to be tested to the appropriate wells. KEEP IN MIND: Each well with alkane MUST HAVE A CONTROL WITHOUT SUBSTRATE. Besides you must have a blank (mixture without enzyme). |
| - | #Warm the plate up for 10-20 min at 37 | + | #Warm the plate up for 10-20 min at 37 °C (in the case of heptadecane). Keep in mind that this is a biphasic system; therefore we need a previous step in which you let the system reach an equilibrium. |
| - | #Add appropriate amounts of cell extracts to the buffer. Remember you are adding cell extract to a highly reduced compound which will easily be oxidized by anything else what is present in the lysate. In order to have enough measurement point per well (before everything going to 0) you must add very small amounts of lysate. For our experiments | + | #Add appropriate amounts of cell extracts to the buffer. Remember you are adding cell extract to a highly reduced compound which will easily be oxidized by anything else what is present in the lysate. In order to have enough measurement point per well (before everything going to 0) you must add very small amounts of lysate. For our experiments 5 μL was sufficient to yield slopes that could be further analysed. Parallel to this we've heat shocked the lysate at 60 °C for 10 minutes. This deactivates most of the ''E.coli'' oxidoreductases and should leave LadA in its native form, thus decreasing ''E.coli'' background 'noise'. |
#Start absorption measurment at 340 nm immediately after adding lysates! | #Start absorption measurment at 340 nm immediately after adding lysates! | ||
| Line 1,117: | Line 1,168: | ||
NOTE: Autoclave the solutions 1 M MgSO4, 1M CaCl2, 20% glucose (w/v) on a liquid cycle. Filter the solution Thiamine-HCl 1M using a 0.22 μm membrane, store the solution in a cold room when is not required. | NOTE: Autoclave the solutions 1 M MgSO4, 1M CaCl2, 20% glucose (w/v) on a liquid cycle. Filter the solution Thiamine-HCl 1M using a 0.22 μm membrane, store the solution in a cold room when is not required. | ||
| - | Store in cold room if the medium is not used. | + | Store in cold room if the medium is not used. |
| + | |||
| + | ''Chef's tips and tricks:'' | ||
| + | |||
| + | Additionally we added 1 mL of E2 salts to 1 Liter of our M9 medium, we did this expecting that we will eventually grow our cells on alkanes and some of our enzymes required cofactors that are present in this solution. | ||
| + | |||
| + | |||
| + | ===E2 Salts=== | ||
| + | For 1 liter dissolve in H2O: | ||
| + | |||
| + | {| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | ||
| + | |'''Compound/Ingredient''' | ||
| + | |'''Amount required (1L of medium)''' | ||
| + | |- | ||
| + | |FeSO4.7H2O | ||
| + | |2.8 g | ||
| + | |- | ||
| + | |MnCl2 | ||
| + | |0.19g | ||
| + | |- | ||
| + | |CoCl2.6H2O | ||
| + | |2.8 g | ||
| + | |- | ||
| + | |CaCl2.4H2O | ||
| + | |1.84g | ||
| + | |- | ||
| + | |ZnSO4.7H2O | ||
| + | |0.28 g | ||
| + | |- | ||
| + | |CuSO4 | ||
| + | |0.16g | ||
| + | |} | ||
| + | |||
| + | Filter this solution using a 0.25 um membrane, and add 1mL/L of M9 medium. | ||
| + | |||
===Antibiotics=== | ===Antibiotics=== | ||
| Line 1,144: | Line 1,229: | ||
For 1 liter dissolve in H2O: | For 1 liter dissolve in H2O: | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | Adjust pH to 7.0 | + | {| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" |
| - | Before use add 5 mL of 2 mM MgCl2 | + | |'''Compound/Ingredient''' |
| + | |'''Amount required (1L of medium)''' | ||
| + | |- | ||
| + | |Bacto tryptone | ||
| + | |20 g | ||
| + | |- | ||
| + | |Yeast extract | ||
| + | |5 g | ||
| + | |- | ||
| + | |NaCl | ||
| + | |0.5 | ||
| + | |- | ||
| + | |250 mM KCl | ||
| + | |10 mL | ||
| + | |} | ||
| + | |||
| + | Adjust pH to 7.0. Before use add 5 mL of 2 mM MgCl2 | ||
| Line 1,179: | Line 1,275: | ||
Adjust volume to 1 liter with H2O | Adjust volume to 1 liter with H2O | ||
| + | |||
| + | ===Potassium Laurate 500 mM=== | ||
| + | Mix in the same tube: | ||
| + | |||
| + | {| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | ||
| + | |'''Compound/Ingredient''' | ||
| + | |'''Amount required''' | ||
| + | |- | ||
| + | |KOH 2M | ||
| + | |2500 µL | ||
| + | |- | ||
| + | |Lauric acid (melted at 50ºC) | ||
| + | |1136 µL | ||
| + | |- | ||
| + | |miliQ water | ||
| + | |6.364 mL | ||
| + | |} | ||
| + | |||
| + | ''Chef's tips and tricks:'' | ||
| + | |||
| + | Warm the mixture up, DON'T SHAKE IT VIGOROUSLY otherwise you'll get a lot of foam. Mix gently and keep it at 50ºC until you won't see any crystal or second phase (around 1 hour after the addition of lauric acid). After your mixture becomes a transparent liquid, you can store it at 4ºC or keep it at room temperature without problems. | ||
| + | |||
| + | |||
| + | ===Glycine 570 mM=== | ||
| + | *Weight 2.1375 g of anhydrous glycine. | ||
| + | *Add the miliQ water required for 50 mL of solution. | ||
| + | |||
| + | |||
| + | ===Tris 500 mM=== | ||
| + | *Weight 3.039 g of anhydrous Tris. | ||
| + | *Add the miliQ water required for 50 mL of solution. | ||
| + | |||
| + | |||
| + | ===NAD 50 mg/mL=== | ||
| + | *Weight 50 mg of anhydrous NAD. | ||
| + | *Add the miliQ water required for 1 mL of solution. | ||
| + | |||
| + | |||
| + | ===Disruption buffer=== | ||
| + | *Add 5 mL of solution 500 mM of Tris | ||
| + | *Add 40 mL of miliQ water | ||
| + | *Adjust the pH by adding 1M HCl until you reach the desired pH (7.4-7.5) | ||
| + | *Add miliQ water in order to complete a volume of 50 mL of solution | ||
| + | |||
| + | |||
| + | ===ADH/ALDH reaction buffer=== | ||
| + | For 25 mL solution Glycine 57mM, NAD 1mM, pH 9.5 dissolve in 20 mL H2O: | ||
| + | |||
| + | {| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | ||
| + | |'''Compound/Ingredient''' | ||
| + | |'''Amount required''' | ||
| + | |- | ||
| + | |570 mM glycine 570 mM of glycine solution | ||
| + | |2.5 mL | ||
| + | |- | ||
| + | |50 mg/mL NAD solution | ||
| + | |331.5 µL | ||
| + | |} | ||
| + | |||
| + | *Adjust the pH using a solution of KOH 2M | ||
| + | *Add miliQ water in order to complete a volume of 25 mL of solution | ||
==Primers== | ==Primers== | ||
Latest revision as of 22:17, 27 October 2010
Protocols
Use of provided BioBricks
From the iGEM organization each team has received a distribution kit containing selected biobricks from the previous years. These biobricks are provided in three 384 well plates containing dried DNA with which competent cells can transformed.
The biobricks are supplied as the DNA in a plasmid with a certain antibiotic resistance, which is denoted by a letter in the plasmid name. All the possible antibiotic resistances that are used within the Biobrick standard are ampicillin (A), chloramphenicol (C), erythromycin (E), gentamycin (G), kanamycin (K), neomycin (N), nalidixic acid (Na), rifampicin (R), spectinomycin (S), streptomycin (St), tetracycline (T), trimethoprim (Tm) and zeocin (Z).
Before use, the dried DNA must be resuspended. This protocol is based on Part registry
Materials
- BioBrick distribution plate
- milliQ
Protocol:
- Punch a hole with a pipette tip through the foil cover into the corresponding well of the desired BioBrick part
- Add 10 μL of milliQ
- Pipette up and down several times, let sit for a few minutes
Making competent cells
Materials:
- bacterial culture
- 0.1 M MgCl2
- 0.1 M CaCl2
- 80% glycerol
- microcentrifuge
- spectrophotometer
Protocol:
- Cultivate bacterial cells in 100 mL LB medium, 37 °C o/n (140 rpm)
- Cultivate 50 mL further in 80 mL LB, ~ 2 hours at 37 °C with shaking (140 rpm) until OD601 = 0.4
- Centrifuge at 4,500 rpm for 6 min at 4 °C
- Resuspend the pellet in 80 mL ice-cold 0.1 M MgCl2
- Centrifuge at 4,500 rpm for 6 min at 4 °C
- Resuspend the pellet in 50 mL ice-cold 0.1 M CaCl2 (do not vortex, stir with Pasteur pipet, solution should look like milk)
- Incubate 15 min on ice
- Centrifuge at 4,500 rpm for 6 min at 4 °C
- Resuspend the pellet in 40 mL 0.1 M CaCl2 (do not vortex, stir with Pasteur pipet)
- Incubate 60 min on ice
- Centrifuge at 4,500 rpm for 10 min at 4 °C
- Add 1 à 2 mL ice-cold 80% glycerol (depends on the amount of pellet)
- Divide in parties of 30 µL and quickly put them in the liquid nitrogen
- Freeze in -80 °C
Transformation Home-Made competent cells
Materials:
- competent cells
- LB medium (warmed to room temperature)
- Plasmid DNA or DNA ligation mix
- LB agar plates containing 15-100 μg/mL antibiotic of choice, pre-warmed to 37 °C
- water bath at 37 °C
- shaking incubator at 37 °C.
Protocol:
- Add 50-100 ng DNA into a 30 μL competent E.coli, and mix gently. Do not mix by pipetting up and down!
- Incubate tube vial on ice for 15 minutes
- Heat-shocks the cells for 5 minutes at 37 °C without shaking
- Immediately transfer the tubes back to ice for 2 minutes
- Add 800 μL of room temperature LB medium
- Cap tube tightly and shake tube horizontally (225 rpm) at 37 °C for 1 hour
- Spin tube (2,000 rpm, 2 minutes), discard supernatant to leave no more than 100 μL, vortex and plate on an agar plate containing antibiotic
- Incubate plates overnight at 37 °C
Transformation commercial competent cells
Materials:
- competent cells
- SOC medium (warmed to room temperature)
- Plasmid DNA or DNA ligation mix
- LB agar plates containing 15-100 μg/mL antibiotic of choice, pre-warmed to 37 °C
- water bath at 42 °C
- shaking incubator at 37 °C.
Protocol:
- Add 50-100 ng DNA into a 20 μL competent E.coli, and mix gently. Do not mix by pipetting up and down!
- Incubate tube vial on ice for 30 minutes
- Heat-shocks the cells for 30 seconds at 42 °C without shaking
- Immediately transfer the tubes back to ice for 2 minutes
- Add 250 μL of room temperature SOC medium
- Cap tube tightly and shake tube horizontally (225 rpm) at 37 °C for 1 hour
- Plate from each tube 100 μL on an agar plate containing antibiotic.
- Incubate plates overnight at 37 °C
Transformation efficiency (CFU/µG)
For example, if 10 pg of pUC19 yield 50 colonies when 50 µL of a 1:100 dilution is plated, then:
Birnboim Plasmid Isolation
This plasmid isolation is used to obtain plasmid DNA that can be used to test the transformed colonies for the right insert.
Materials:
- bacterial culture
- milliQ
- BB solution I (10 mg/mL glucose; 25 mM Tris-HCl pH 8,0; 10 mM EDTA, 0,1 mg/mL RNAse A)
- BB solution II (0.2 NaOH; 1% SDS)
- BB solution III (3 M NaAc, pH 4,8 = 100 mL: 40,8 g NaAc.3H2O, 38 mL HAc, 52 mL H2O)
- BB solution IV (0.3 m NaAc)
- cold 70% ethanol
- cold 100% ethanol
- microcentrifuge
- nanodrop
Protocol:
- Pour 1 mL overnight bacterial culture into Eppendorf tube, centrifuge for 1 minute at maximum speed (13,000 rpm). Carefully suck off supernatant
- Resuspend the bacterial pellet in 100 μL cold solution I, suspend well (vortex) and incubate for 15 min on ice
- Vortex shortly and add immediately 200 μL fresh solution II (make fresh, every two weeks, keep at room temperature), mix vigorously (no vortexing) and leave 1 minute at room temperature
- Add 150 μL cold solution III, mix gently until white precipitate appears and incubate on ice for 30 minutes
- Centrifuge at maximum speed for 5 minutes at room temperature (Eppendorf centrifuge)
- Take 400 μL of the supernatant (beware of taking along some pellet) and add to 800 μL of cold (-20 °C) 100% ethanol
- Mix, leave at -20 °C for 30 minutes and centrifuge at room temperature for 10 minutes maximum speed (Eppendorf centrifuge)
- Carefully pipet off the supernatant
- Add 100 μL solution IV, mix by vortexing and add 200 μL ice cold 100% ethanol. Incubate at -20 °C for 30 minutes. Centrifuge 10 minutes full speed (Eppendorf centrifuge), carefully discard supernatant (pipet)
- Wash DNA pellet with 1 mL of ice cold 70% ethanol and centrifuge for 10 minutes maximum speed (Eppendorf centrifuge) can carefully pipet off the supernatant
- Air-dry the pellet for 5 minutes, and redissolve the DNA in 50 μL water
- Measure DNA concentration on the Nanodrop
QIAGEN Mini-prep plasmid isolation
This protocol is based on QIAGEN® Plasmid Purification Handbook.
This protocol is designed for preparation of up to 20 µg of high-copy plasmid or cosmid DNA using the Qiagen Plasmid Mini Kit. Qiagen plasmid isolation is used to obtain very pure plasmid DNA.
Maximum recommended culture volumes for the Qiagen-tip 20:
High-copy plasmids 1-5 mL
Materials:
- bacterial culture
- Qiagen colums
- buffer P1 (100 μg/mL RNAse A, 50 mM Tris/HCl, 10 mM EDTA, pH 8.0)
- buffer P2 (200 mM NaOH, 1% SDS)
- buffer P3 (3 M KAc, pH 5.5)
- buffer PE
- milliQ pH 8.0
- centrifuge
- nanodrop
Protocol:
- Pick a single colony from a freshly streaked selective plate and inoculate a starter culture of 2–5 mL LB medium containing the appropriate selective antibiotic. Incubate for approximately 8 h at 37°C with vigorous shaking (approx. 300 rpm)
- Harvest the 5 mL bacterial cells by centrifugation at 13,000 rpm for 1 min at 20°C (microcentrifuge tube). If you wish to stop the protocol and continue later, freeze the cell pellets at –20°C
- Resuspend pelleted bacterial cells in 250 μL Buffer P1. Ensure that RNase A has been added to Buffer P1. No cell clumps should be visible after resuspension of the pellet.
- Add 250 μL Buffer P2 and mix thoroughly by inverting the tube 4–6 times. Mix gently by inverting the tube. Do not vortex, as this will result in shearing of genomic DNA. If necessary, continue inverting the tube until the solution becomes viscous and slightly clear. Do not allow the lysis reaction to proceed for more than 5 minutes.
- Add 350 μL Buffer N3 and mix immediately and thoroughly by inverting the tube 4–6 times. To avoid localized precipitation, mix the solution thoroughly, immediately after addition of Buffer N3. Large culture volumes (e.g. ≥5 mL) may require inverting up to 10 times. The solution should become cloudy.
- Incubate at -20 °C for 15 minutes.
- Centrifuge for 10 min at 13,000 rpm in a table-top microcentrifuge. A compact white pellet will form.
- Apply the supernatants from step 7 to the QIAprep spin column by decanting or pipetting.
- Centrifuge for 30–60 seconds. Discard the flow-through.
- Wash QIAprep spin column by adding 0.75 mL Buffer PE and centrifuging for 30–60 seconds.
- Discard the flow-through, and centrifuge for an additional 1 min to remove residual wash buffer. Important: Residual wash buffer will not be completely removed unless the flow-through is discarded before this additional centrifugation. Residual ethanol from Buffer PE may inhibit subsequent enzymatic reactions.
- Place the QIAprep column in a clean 1.5 mL microcentrifuge tube. To elute DNA, add 30 μL Buffer EB (10 mM Tris·Cl, pH 8.5) or water to the center of each QIAprep spin column, let stand for 1 minute, and centrifuge for 1 minute.
- Measure DNA concentration on the Nanodrop
QIAGEN Midi-prep plasmid isolation
This protocol is based on QIAGEN® Plasmid Purification Handbook.
This protocol is designed for preparation of up to 100 µg of high- or low-copy plasmid or cosmid DNA using the Qiagen Plasmid Midi Kit. Qiagen plasmid isolation is used to obtain very pure plasmid DNA.
Maximum recommended culture volumes for the Qiagen-tip 100:
High-copy plasmids 25 mL
Low-copy plasmids 100 mL
Materials:
- bacterial culture
- Qiagen colums
- buffer P1 (100 μg/mL RNAse A, 50 mM Tris/HCl, 10 mM EDTA, pH 8.0)
- buffer P2 (200 mM NaOH, 1% SDS)
- buffer P3 (3 M KAc, pH 5.5)
- buffer QBT (750 mM NaCl, 50 mM MOPS, 15% Ethanol, pH 7.0, 0.15% Triton X-100)
- buffer QC (1 M NaCl, 50 mM MOPS, 15% Ethanol, pH 7.0)
- buffer QF (1.25 M NaCl, 50 mM Tris/HCl, 15% Ethanol, pH 8.5)
- isopropanol
- milliQ
- centrifuge
- nanodrop
Protocol:
- Pick a single colony from a freshly streaked selective plate and inoculate a starter culture of 25 to 100 mL LB medium containing the appropriate selective antibiotic. Incubate for approximately 8 h at 37°C with vigorous shaking (approx. 300 rpm)
- Harvest the 25 to 100 mL bacterial cells by centrifugation at 4,000 rpm for 15 min at 4°C ( Sorvall buckets). If you wish to stop the protocol and continue later, freeze the cell pellets at –20°C
- Resuspend the bacterial pellet in 4 mL of chilled Buffer P1 (containing RNAse) and pour into a blue cap 50 mL tube
- Add 4 mL of Buffer P2, mix gently but thoroughly by inverting 4–6 times, and incubate at room temperature for 5 minutes
- Add 4 mL of chilled Buffer P3, mix gently by inverting 4–6 times, and incubate on ice for 15 minutes
- Centrifuge 30 min at 4,000 rpm and pour supernatant through cheesecloth into a fresh blue cap 50 mL tube
- During centrifugation equilibrate a Qiagen-tip 100 by applying 4 mL Buffer QBT, and allow the column to empty by gravity flow
- Apply the supernatant from step 6 onto the Qiagen-tip and allow it to enter the resin by gravity flow
- Wash the Qiagen-tip with 2 × 10 mL Buffer QC
- Elute DNA with 5 mL Buffer QF into 15 mL blue cap tube
- Precipitate DNA by adding 3.5 mL (0.7 volumes) room-temperature isopropanol to the eluted DNA
- Mix and centrifuge immediately at 4,000 rpm for 30 minutes at 4 °C
- Proceed immediately when centrifuge stops and carefully decant the supernatant
- Wash DNA pellet with 2 mL of cold 70% ethanol, and centrifuge at 4,000 rpm for 10 minutes at 4 °C
- Carefully decant the supernatant without disturbing the pellet
- Air-dry the pellet for 5–10 min (has to be dry), and redissolve the DNA in a suitable volume of milliQ H2O
- Measure DNA concentration on the Nanodrop
QIAGEN Maxi-prep plasmid isolation
This protocol is based on QIAGEN® Plasmid Purification Handbook.
This protocol is designed for preparation of up to 100 µg of high- or low-copy plasmid or cosmid DNA using the Qiagen Plasmid Midi Kit. Qiagen plasmid isolation is used to obtain very pure plasmid DNA.
Maximum recommended culture volumes for the Qiagen-tip 100:
High-copy plasmids 100 mL
Low-copy plasmids 500 mL
Materials:
- bacterial culture
- Qiagen colums
- buffer P1 (100 μg/mL RNAse A, 50 mM Tris/HCl, 10 mM EDTA, pH 8.0)
- buffer P2 (200 mM NaOH, 1% SDS)
- buffer P3 (3 M KAc, pH 5.5)
- buffer QBT (750 mM NaCl, 50 mM MOPS, 15% Ethanol, pH 7.0, 0.15% Triton X-100)
- buffer QC (1 M NaCl, 50 mM MOPS, 15% Ethanol, pH 7.0)
- buffer QF (1.25 M NaCl, 50 mM Tris/HCl, 15% Ethanol, pH 8.5)
- isopropanol
- milliQ
- centrifuge
- nanodrop
Protocol:
- Pick a single colony from a freshly streaked selective plate and inoculate a starter culture of 100 to 500 mL LB medium containing the appropriate selective antibiotic. Incubate for approximately 8 h at 37°C with vigorous shaking (approx. 300 rpm)
- Harvest the 100 to 500 mL bacterial cells by centrifugation at 4,000 rpm for 15 min at 4°C (Sorvall buckets). If you wish to stop the protocol and continue later, freeze the cell pellets at –20°C
- Resuspend the bacterial pellet in 10 mL of chilled Buffer P1 (containing RNAse) and pour into a blue cap 50 mL tube
- Add 10 mL of Buffer P2, mix gently but thoroughly by inverting 4–6 times, and incubate at room temperature for 5 minutes
- Add 10 mL of chilled Buffer P3, mix gently by inverting 4–6 times, and incubate on ice for 20 minutes
- Centrifuge 30 min at 4,000 rpm and pour supernatant through cheesecloth into a fresh blue cap 50 mL tube
- During centrifugation equilibrate a Qiagen-tip 100 by applying 10 mL Buffer QBT, and allow the column to empty by gravity flow
- Apply the supernatant from step 6 onto the Qiagen-tip and allow it to enter the resin by gravity flow
- Wash the Qiagen-tip with 2 × 30 mL Buffer QC
- Elute DNA with 15 mL Buffer QF into 50 mL blue cap tube
- Precipitate DNA by adding 10.5 mL (0.7 volumes) room-temperature isopropanol to the eluted DNA
- Mix and centrifuge immediately at 4,000 rpm for 30 minutes at 4 °C
- Proceed immediately when centrifuge stops and carefully decant the supernatant
- Wash DNA pellet with 5 mL of cold 70% ethanol, and centrifuge at 4,000 rpm for 10 minutes at 4 °C
- Carefully decant the supernatant without disturbing the pellet
- Air-dry the pellet for 5–10 min (has to be dry), and redissolve the DNA in a suitable volume of milliQ H2O
- Measure DNA concentration on the Nanodrop
Freezing of bacterial stocks
Materials:
- bacterial culture
- LB medium
- 80% glycerol
- centrifuge
Protocol:
- Take 5 mL bacterial cells from the Erlenmeyer of a freshly grown culture and spin in a 15 mL tube for 10 minutes at 2,000 rpm (Eppendorf centrifuge)
- Decant the supernatant without disturbing the pellet
- Pipet on the pellet 0.5 mL 1% LB medium and 0.5 mL 80% glycerol and mix by vortexing and save in -80 °C freezer
Restriction enzyme digestion
Materials:
- plasmid DNA or PCR product
- restriction enzymes (Roche and BioLabs)
- buffer (10x)
- H2O
- water bath at 37 °C
- heat block or water bath at 65 °C
Protocol:
Digestions (cutting plasmid DNA) were performed at the appropriate temperature with the appropriate buffer in the appropriate concentration, according to the supplier.
Reaction for one sample:
| Component | Sample |
| DNA | x μL (up to 1,0 μg) |
| Buffer (10x) | x μL (for 1×) |
| Restriction enzymes | x μL (10 units/μg DNA = 1 µL) |
| H2O | x μL |
| 20-25 μL |
Incubate for (at least) one hour at 37 °C. Inactivate the restriction endonucleases by heat, incubation at 80°C for 10 minutes and centrifuge shortly.
Used Buffers:
Buffer H (Roche): 50 mM Tris-HCl, 1 M NaCl, 100 mM MgCl2, 10 mM DTE, pH 7.5 at 37 °C
Buffer M (Roche): 100 mM Tris-HCl, 500 mM NaCI, 100 mM MgCl2, 10 mM DTE, pH 7.5 at 37 °C
Buffer 1 (BioLabs): 10 mM Bis-Tris-Propane-HCl, 10 mM MgCl2, 1 mM DTE,pH 7.0 at 25°C
Buffer 2 (BioLabs): 50 mM NaCl, 10 mM Tris-HCl, 10 mM MgCl2, 1 mM DTE, pH 7.9 at 25°C
Buffer 3 (BioLabs): 100 mM NaCl, 50 mM Tris-HCl, 10 mM MgCl2, 1 mM DTE, pH 7.9 at 25°C
Buffer 4 (BioLabs): 50 mM CH3CO2K, 20 mM TAE, 10 mM Mg(CH3COO)2, 1 mM DTE, pH 7.9 at 25°C
Note: some of the restriction enzymes of New England BioLabs required the addition of 100 µg/mL BSA
Ligation
Materials:
- digested plasmid DNA or PCR product
- T4 ligation buffer (10x) (Fermentas)
- T4 ligase (Fermentas)
- H2O
- water bath at 16 °C
Protocol:
Ligations (pasting plasmid DNA) were performed at the appropriate temperature with the appropriate buffer in the appropriate concentration, according to the supplier.
Reaction for one sample:
| Component | Sample |
| DNA insert | x μL |
| DNA vector | x μL |
| T4 Ligation buffer (10×) | x μL (for 1×) |
| T4 Ligase | 1.0 μL |
| H2O | x μL |
| 10-15 μL |
The final concentration is preferably ~100 ng/μL. Incubate at 16 °C for at least 3 hours. For transformation use circa half of the ligation mix.
Transform circa half of the ligation mix. Incubate at 16 °C o/n
Agarose gel
Materials:
- agarose
- TBE buffer
- SYBRSafe (Invitrogen)
Protocol:
- Dissolve 1 g of agarose in 100 mL 1x TBE buffer for 1% gel (can be stored in the 70 °C stove)
- Add 5 µL of SYBRSafe
PCR
Materials:
- Pfx polymerase (Invitrogen)
- 10x Pfx Buffer (Invitrogen)
- enhancer (Invitrogen)
- 50 mM MgSO4 (Invitrogen)
- 10 mM dNTPs
- primer solutions 5 mol/mL
- template DNA (plasmid at 50 pg – 1 ng/μL)
- PCR machine
Protocol:
First make sure that there is a PCR machine available for you. Take the solutions from the freezer and thaw them on ice. Preparation of reaction mixture:
- Gently vortex and briefly centrifuge all solutions after thawing
- Keep solutions on ice
- Add to a thin walled PCR tube, on ice, for a 50 μL reaction:
1× pre-mix
| Component | Sample |
| Pfx polymerse | 0.6 μL |
| 10x Pfx buffer | 5 μL |
| Enhancer | 5 μL |
| 10 mM dNTPs | 1.5 μL |
| 50 mM MgSO4 | 1 μL |
| Primer 1 | 3 μL |
| Primer 2 | 3 μL |
| DNA template | 1 μL |
| H20 | 29.9 μL |
- Gently vortex the sample and briefly centrifuge (5 sec) to collect all droplets at the bottom of the tube
PCR program:
| Step | Annealing Temperature | Time, min:sec | Number of cycles |
| Initial denaturation | 95 °C | 2:00 | 1 |
| Annealing | x °C * | 1:00 | 1 |
| Extension | 68 °C | 1:00-2:00 | 1 |
| Denaturation | 95 °C | 1:00 | 25 |
| Annealing | x °C * | 1:00 | 25 |
| Extension | 68 °C | 1:00-2:00 | 25 |
| Final Extension | 68 °C | 10:00 | 1 |
* Annealing temperature dependent on primer. Optimal temperature: 3x G/C + 2x A/T
Overlapping PCR
Materials:
- Pfx polymerase (Invitrogen)
- 10x Pfx Buffer (Invitrogen)
- enhancer (Invitrogen)
- 50 mM MgSO4 (Invitrogen)
- 10 mM dNTPs
- primer solutions 5 mol/mL
- template DNA (plasmid at 50 pg – 1 ng/μL)
- PCR machine
Protocol:
First make sure that there is a PCR machine available for you. Take the solutions from the freezer and thaw them on ice. Preparation of reaction mixture:
- Gently vortex and briefly centrifuge all solutions after thawing
- Keep solutions on ice
- Add to a thin walled PCR tube, on ice, for PCR reaction:
1× pre-mix
| Component | Sample |
| Pfx polymerse | 0.6 μL |
| 10x Pfx buffer | 5 μL |
| Enhancer | 5 μL |
| 10 mM dNTPs | 1.5 μL |
| 50 mM MgSO4 | 1 μL |
| DNA template | 1 μL |
| H20 | 29.9 μL |
- Gently vortex the sample and briefly centrifuge (5 sec) to collect all droplets at the bottom of the tube
PCR program:
| Step | Annealing Temperature | Time, min:sec | Number of cycles |
| Initial denaturation | 95 °C | 2:00 | 1 |
| Annealing | X °C* | 1:00 | 1 |
| Extension | 68 °C | 1:00 | 1 |
| Denaturation | 95 °C | 1:00 | 5 |
| Annealing | X °C * | 1:00 | 5 |
| Extension | 68 °C | 1:00 | 5 |
| add primers (3 μL Primer 1 + 3 μL Primer 2) | |||
| Denaturation | 95 °C | 1:00 | 25 |
| Annealing | X °C * | 1:00 | 25 |
| Extension | 68 °C | 1:00-2:00 (1min/kb) | 25 |
| Final Extension | 68 °C | 10:00 | 1 |
* Annealing temperature dependent on primer. Optimal temperature: 3x G/C + 2x A/T
Colony PCR
Materials:
- Taq PCR Master Mix (Qiagen) is premixed solution containing Taq DNA Polymerase, PCR Buffer and dNTPs. The solution provides a final concentration of 1.5 mM MgCl2 and 200 μM each dNTP.
- primer solutions 5 mol/mL
- PCR machine
Protocol:
First make sure that there is a PCR machine available for you. Take the solutions from the freezer and thaw on ice. Preparation of reaction mixture:
- Gently vortex and briefly centrifuge all solutions after thawing
- Keep solutions on ice
- Make a pre-mix for the amount of colonies (to be analyzed) + 1, add to an Eppendorf tube:
1× pre-mix
| Component | Sample |
| Taq PCR Master Mix (Qiagen) | 12.5 μL |
| Primer 1 | 1.5 μL |
| Primer 2 | 1.5 μL |
| H20 | 9.5 μL |
- Add 25 μL of pre-mix to each PCR tube.
- Pick a sterile toothpick into a colony, dip it into a PCR tube, put it into a 15 mL culture tube containing 5 mL LB medium + antibiotics and then grow overnight for mini-prep cultures and -80 °C stocks. Repeat this for all the colonies. Incubate the mini-prep cultures at 37 °C.
- Keep everything ice cold until you put the tubes in the preheated PCR machine
PCR program:
| Step | Annealing Temperature | Time, min:sec | Number of cycles |
| Initial denaturation | 94 °C | 2:00 | 1 |
| Denaturation | 94 °C | 1:00 | 30 |
| Annealing | x °C * | 0:45 | 30 |
| Extension | 72 °C | 1:00 | 30 |
| Final Extension | 72 °C | 5:00 | 1 |
* Annealing temperature dependent on primer. Optimal temperature: 3x G/C + 2x A/T
QIAGEN Gel Extraction
This protocol is based on QIAGEN® Gel Extraction Handbook.
Materials:
- QIAquick columns
- buffer QG
- buffer PE
- isopropanol
- milliQ
- microcentrifuge
- heat block at 50 °C
Protocol:
- Excise the DNA fragment from the agarose gel with a clean, sharp scalpel. Minimize the size of the gel slice by removing extra agarose.
- Weigh the gel slice in a colorless tube. Add 3 volumes of Buffer QG to 1 volume of gel (100 mg ~ 100 μL). For >2% agarose gels, add 6 volumes of Buffer QG. The maximum amount of gel slice per QIAquick column is 400 mg; for gel slices >400 mg use more than one QIAquick column.
- Incubate at 50°C for 10 min (or until the gel slice has completely dissolved). To help dissolve gel, mix by vortexing the tube every 2–3 min during the incubation. IMPORTANT: Solubilize agarose completely. For >2% gels, increase incubation time.
- After the gel slice has dissolved completely, check that the color of the mixture is yellow (similar to Buffer QG without dissolved agarose). If the color of the mixture is orange or violet, add 10 μL of 3 M sodium acetate, pH 5.0, and mix. The color of the mixture will turn to yellow. The adsorption of DNA to the QIAquick membrane is efficient only at pH <7.5. Buffer QG contains a pH indicator which is yellow at pH <7.5 and orange or violet at higher pH, allowing easy determination of the optimal pH for DNA binding.
- Add 1 gel volume of isopropanol to the sample and mix. This step increases the yield of DNA fragments <500 bp and >4 kb. For DNA fragments between 500 bp and 4 kb, addition of isopropanol has no effect on yield. Do not centrifuge the sample at this stage.
- Place a QIAquick spin column in a provided 2 mL collection tube.
- To bind DNA, apply the sample to the QIAquick column, and centrifuge for 1 min. The maximum volume of the column reservoir is 800 μL. For sample volumes of more than 800 μL, simply load and spin again.
- Discard flow-through and place QIAquick column back in the same collection tube.
- To wash, add 0.75 mL of Buffer PE to QIAquick column and centrifuge for 1 min.
- Discard the flow-through and centrifuge the QIAquick column for an additional 1 min at 10,000 x g (~13,000 rpm). IMPORTANT: Residual ethanol from Buffer PE will not be completely removed unless the flow-through is discarded before this additional centrifugation.
- Place QIAquick column into a clean 1.5 mL microcentrifuge tube.
- To elute DNA, add 50 μL of Buffer EB (10 mM Tris·Cl, pH 8.5) or H2O to the center of the QIAquick membrane and centrifuge the column for 1 min at maximum speed. Alternatively, for increased DNA concentration, add 30 μl elution buffer to the center of the QIAquick membrane, let the column stand for 1 min, and then centrifuge for 1 min. Important: Ensure that the elution buffer is dispensed directly onto the QIAquick membrane for complete elution of bound DNA. The average eluate volume is 48 μL from 50 μL elution buffer volume, and 28 μL from 30 μL. Elution efficiency is dependent on pH. The maximum elution efficiency is achieved between pH 7.0 and 8.5. When using water, make sure that the pH value is within this range, and store DNA at –20°C as DNA may degrade in the absence of a buffering agent. The purified DNA can also be eluted in TE (10 mM Tris·Cl, 1 mM EDTA, pH 8.0), but the EDTA may inhibit subsequent enzymatic reactions.
Precipitation of PCR products
Materials:
- 3 M sodium acetate pH 4.8
- 96% ethanol
- 70% cold ethanol
- milliQ
- microcentrifuge
- heat block at 37 °C
- nanodrop
Protocol:
- Add 1/10 volumes of 3 M sodium acetate pH 4.8 to the PCR mixture
- Add 2 volumes 96% ethanol
- Incubate 15 minutes at -80 °C or 30 minutes at -80 °C
- Centrifuge at 14.000 rpm for 10-15 minutes at 4 °C
- Proceed immediately when centrifuge stops and carefully decant the supernatant
- Wash DNA pellet with 100 μL of ice cold 70% and centrifuge at 14.000 rpm for 10 minutes at 4 °C
- Carefully decant the supernatant without disturbing the pellet
- Air-dry the pellet and redissolve the DNA in water and incubate for 10 minutes at 37 °C
- Measure DNA concentration on the Nanodrop
Protein Purification
Materials:
- bacterial culture on plates
- LB medium
- 25 mM TEA (triethanolamine buffer), pH 8.0
- 1% lysozyme in TEA
- 4 M Urea
- 10% streptomycin in TEA
- 25 mM Tris buffer, pH 8.0
- glass beads
- centrifuge
- spectrophotometer
Protocol:
- Cultivate bacterical cells on plates, 37 °C o/n (140 rpm)
- Pick a single colony from a freshly streaked selective plate and inoculate a starter culture of 250 mL LB medium containing the appropriate selective antibiotic. Incubate at 37°C with vigorous shaking
- Induce production of protein with 1 mM IPTG when OD600 is reached (check with 1.5 mL sample)
- Harvest after 2 hours the 250 mL bacterial cells by centrifugation at 10.000 rpm for 10 min (Sorvall buckets)
- Collect the free supernatant and store it
- Wash pellet with TEA buffer, pH 8.0
- Freeze pellet in -80 °C for 15 minutes
- Resuspend pellet in 30 mL TEA with 1% lysozyme (final concentration 0.02%)
- Incubate for 10 min at RT
- Disrupt cells with glass beads and vortex for 10 minutes
- Centrifuge at 14 .000 rpm for 30 min at 4°C (protein is in supernatant)
- Aliquot 1.8 mL supernatant into fresh eppendorf tubes and add 200 μL 10% streptomycin in TEA (final concentration 1%).
- Centrifuge supernatant 14 .000 rpm for 30 min at 4°C (protein is in pellet)
- Resuspend pellet in 4 M Urea
- Centrifuge supernatant 14.000 rpm for 30 min at 4°C (protein is in supernatant)
- Dialyse supernatant from step 14 overnight 20 mM Tris buffer
- Dialyse supernatant from step 5 overnight 20 mM Tris buffer
Protein content measurements (Bradford Assay)
Materials:
- 2 mg/mL BSA
- bradford reagents
- spectrophotometer
Protocol:
- Make a dilution serie of standard 2 mg/mL bovine serum albumin (BSA). Use 0, 20, 40, 60, 80, 100 μL
- Add MillQ to a total volume of 800 μL
- Add 200 μL Bradford reagens and mix
- Measure after minimal 5 min and max 60 min the OD595
Emulsifier Assay
Materials:
- 25 mM Tris buffer, pH 8
- emulsifier sample
- 0.1% Sudan II in 50% EtOH
- shaking incubator at 30 °C
- spectrophotometer
Protocol:
- Determine protein concentration by Bradford assay
- Aliquot 0.1-0.5 mL of protein sample in a cuvet
- Add Tris buffer up to 1 mL
- Add 20 μL 0.1% Sudan II
- Vortex 15 seconds at max speed
- Let sample rest for 1 minute
- Measure absorbance at OD493
- Dilute sample to get in the right absorbance range if neccesary
Preparing cell lysates for enzyme kinetics measurements
Materials:
- bacterial culture on plates
- LB medium
- 50 mM Tris buffer, pH 7.5 or 7.4 (or other, e.g. enzyme's optimal pH)
- 100 mL glass flasks
- 50 mL centrifuge tubes
- Shaking incubator
- Sonicator (Branson Sonifier 250)
- centrifuge
Protocol
- Pick a single colony from a freshly streaked selective plate and inoculate a starter culture of 50 mL LB medium containing the appropriate selective antibiotic. Incubate at 37°C, 160 rpm.
- Chef's tips and tricks:
- You can also inoculate directly from your cell bank (cell stock with glycerol 40%v/v in fresh medium). Then you can start with a previous overnight culture (5 mL) and then inoculate 50 mL of fresh medium with 1 mL, in that way you will reach your desired OD around 2-3 hours you started the culture.
- Normally, we use 5 μL to inoculate 5 mL. Some people may think that this is a lot, specially because of the fact that cells are concentrated in the cell bank. .But, when we used a different amount we had some nasty surprises like no overnight growth. Spin down the cultures at an OD of +/- 0.6 in 50 mL tubes for 10 min at 4.000 rpm using a chilled (4 °C) centrifuge.
- Chef's tips and tricks:
- Resuspend the cells in 5 mL of 50 mM Tris buffer (with the appropriate pH) and keep on ice until sonication.
- Chef's tips and tricks:
- Sometimes we use a washing step, specially when we want to prepare a extract of 100 mL of culture (2 flasks of 50 mL). Then after you resuspend your cells, spin them down and resuspend them again. If you think that this could be futile check the amount of LB remaining next your cells, it's around 200 μL. Just in case you don't want this extra source of pH disturbance, you can do this washing step before you kill your cells.
- Chef's tips and tricks:
- Sonicate the suspension (placing the tip as close as possible to the bottom of the tube, without touching) for 3 minutes at a 40% duty cycle and an output control of 4; all the while keeping the suspension on ice.
- Chef's tips and tricks:
- BE CAREFUL, place the tip of your sonicator in the middle of your tube next to the bottom, but not touching the tube otherwise you can break it. If you place the tip somewhere else, you will end up with a nice E. coli milkshake with zero acitivity. Presence of a lot of foam during the disruption is bad sign; that nice foam it's mainly denaturated protein. So please, BE CAREFUL.
- You need ice all the time. Sonication produces a lot of heat, if you have a heat sensitive protein or you don't want to mess up with you negative control (E. coli), then KEEP YOUR CELL SUSPENTION ALL THE TIME ON ICE.
- If you see a nice reduction of O.D. by naked eye inspection, then you are in the good way. If that's not the case you can give to the cell suspension some extra minutes (1 or 2) in the sonicator. There should be a nice difference between disrupted cells and normal cell suspension. Disrupted cells produce a transparent(ish) suspension, whereas normal cells have a milky aspect.
- Chef's tips and tricks:
- Spin down the suspension for 5 minutes at 4.000 rpm in a chilled centrifuge.
- Chef's tips and tricks:
- You can see a nice reduction of you cell pellet, and that could be use as a criterion for keep on going with your experiments, sometimes the reduction of the pellet is not that considerable but you may see a viscous supernatant (compared to your buffer). That's a sign of a nice disruption; if you cannot see these things happening to your suspension then something went wrong. You can do an extra sonication step.
- Chef's tips and tricks:
- Your supernatant now contains your enzyme of interest. You can do protein determination assays. Recomendation: DON'T BE TOO GREEDY!!! If you have some protein don't do an extra sonication step in order to obtain more. If you make a mistake or you give extra energy to your extract you may lose the sample and you have to start all over again.
Evaluating
Use the following checking list in order to know if you have to do an extra disruprtion step or start all over again:
- Is the pellet is smaller compared to the one obtained before disruption? If yes continue to 2.
- Did the supernatant look like more viscous than water when I transfer the supernatant?If yes continue to 3.
- Was the cell extract foamy (like a milkshake) before the last centrifugation? If yes, go to Doubtful results and pray, continue to 4.
- Do the bradford or protein analysis results show that I have more signal in my disruption cells than in the control (buffer)? If yes go to Successful disruption and smile, if not go to Extra step and cry if you feel like that.
Troubleshooting
- Extra step: Your disruption went really wrong, repeat the sonication step and use the checklist again. If you still have problems (like no protein or a mediocre amount of protein) then ask the person in charge of the sonicator or someone more qualified than you to check the machine.
- Doubtful results: Sometimes your activities are lower than they are in reality, which will give you a false positive in case that you want to use your protein for activity assays. If this is your first time with the protocol, don't be scared; keep going on with your tests but take into consideration the advice that YOU HAVE TO REPEAT IT in order to proof that your activities are correct. You can use two or three E. coli cultures in the same experiment, so that if you mess up with one you have other(s) ready to be used as a confirmation of your results.
Tolerance assay
Materials
- Overnight culture
- LB medium with the appropriate antibiotics
- 15 mL tubes
- Appropriate concentration of the agitant (e.g. salt or hydrocarbon)
- 96 well plate
- Plate reader
Protocol
- Subculture 10 μL of the overnight culture into 5 mL of LB (with antibiotic) in a 15 mL tube.
- Grow to an OD600 of 0.3-0.4.
- Dilute back to an OD600 of 0.1.
- Store on ice.
- Load the 96 well plate with LB (with antibiotic) and the agitant you wish to measure. Note: do all the experiments in triplicate.
- Measure the growth curves using the plate reader
Alcohol/Aldehyde dehydrogenase activity assays
Materials
- Cell extracts
- Buffer 57 mM glycine, 1mM NAD, pH 9.5
- Alcohol/Aldehyde
- 96 well plate
- Plate reader
Protocol
- Determine the protein content (mg/mL) of your cell extract. Use whatever method you prefer.
- Keep your extracts on ice.
- Add glycine buffer to the wells that you are going to use, usually we used 180 μL per well.
- Add the alcohol/aldehyde to be tested to the appropriate wells, usually we added 5 μL. WARNING: Each well with alcohol/aldehyde MUST HAVE A CONTROL WITHOUT SUBSTRATE. Don't forget to prepare a BLANK, which will be a mixture of buffer and substrate without enzyme.
- Chef's tips and tricks:
- Long-chain alcohols are not in a liquid form at room temperature, keep them in some warm place before you start your assay, so that they will be melted whenever you will use them.
- Chef's tips and tricks:
- Warm the plate up for 10-20 min at 37ºC. Keep in mind that this is a biphasic system; therefore we need a previous step in which you let the system to reach a equilibrium. Otherwise your first measurements will be noisy and you have to extend the assay, in case that you have a really powerful enzyme that may mean that you will repeat the experiment because the NAD will dissappear very quick. In case that your enzyme is not that fast, you can skip this step and add the enzyme; if you have noisy results at the beggining of the measurement, discard them and analyze the rest (at least 20 min of analysis, preferable more than 5 data points).
- Add the cell extract to your reaction mixture, usually we added 40 uL.
- Measure in your plate reader at wavelength of 340 nm, it is better if you run your experiment for 1 hour or so. Try to do data acquisition each 2-3 minutes, in that way you will have more than enough data to work with.
Data analysis
- Divide each data point by the extinction coefficient of NADH (6220 M^-1 cm^-1) and by the path length; in our case we had reaction volumes of 225 μL (buffer+enzyme+substrate) and 220 μL (buffer+enzyme). Each well had a length of 1 cm and a total volume of 300 μL. That means that we had path lengths of 0.75 cm and 0.7333 cm. The number that you will get is the MOLAR CONCENTRATION of NADH in you well.
- Multiply the molar concentration of NADH in your well by the reaction volume in order to get the TOTAL AMOUNT OF NADH MOLES in your well.
- Rest the NADH moles from the BLANK (buffer+substrate WITHOUT enzyme) to all your wells.
- For each data point do the following operation: NADH moles produced = NADH moles (buffer+enzyme+substrate) - NADH moles (buffer+enzyme)
- Plot on the x-axis the time and in the y-axis the NADH moles produced. Perform a linear regression and report the slopes (moles/min) of each well. One Unit of alcohol/aldehyde dehydrogenase is defined as the amount of enzyme required for producing 1 umol of NADH in one minute.
- Divide the amount of Units of enzyme by the amount of TOTAL PROTEIN added to the reaction mixture. That will give you a result in U/mg of protein.
RBS characterization experiment
Materials
- -80 stocks or plates of strains carrying the RBS of interest, a reference and an 'empty' plasmid
- 96-well plate + lid
- 96-well plate fluorescence + absorbance reader (BioTek Synergy)
Protocol
- Inoculate 5 mL of LB+Amp with your strains of interest, reference strain (B0032) and empty plasmid strain (I13401). Do this in triplicate. Grow up o/n.
- The following day subculture 2 μL of o/n culture in 200 μL of LB+Amp (1:100 dilution) on 96 well plates, all in quadruplicate (you will have 12 wells per strain now, a full row!)
- Culture the cells up to an OD of about 0.3 and dilute 1:100 onto another 96 well plates in the same order as before usig LB+Amp.
- Hurry your plate over to the plate reader and start measuring GFP fluorescence and biomass absorbance!
- The plate is read by an automatically repeating protocol with shaking at medium speed. The program does the following:
- Sets the temperature to 37°C
- In a kinetic loop of a fixed time (16 hour 30 mins) the following measurements are taken with a time interval of 10 minutes with shaking: absorbance (600 nm filter) and GFP fluorescence (485 nm excitation and 520 nm emission).
- followed by a delay of 100 seconds.
pCaiF characterization experiment
Materials
- -80 stocks or plates of strains carrying the biobrick BBa_K398331, a reference and an 'empty' plasmid
- 96-well plate + lid
- 96-well plate fluorescence + absorbance reader (BioTek Synergy)
Protocol
- Inoculate 5 mL of M9 (10 g/L of glc) with your strains of interest and empty plasmid strain (J13002). Grow them overnight.
- Chef's tips and tricks:
- Be aware that normal M9 contains 2 g/L. In order to prepare this new medium we just added 5 times the specified amount of glucose for normal M9 medium.
- Chef's tips and tricks:
- In the next morning subculture 50 μL of o/n culture in another 5 mL of M9 (10 g/L of glc).
- Check the O.D. at 600nm of the cultures after some hours (usually 3 h), if this parameter is in the range of 0.15-0.3 you can start your assay, otherwise wait until you have at least an O.D. of 0.2.
- Chef's tips and tricks:
- We use to check the O.D. 2 hours after we started the culture and we try to adjust the O.D. to the lowest measurement obtained, once they are in the same O.D. when they are diluted the greater are the chances that you don't have to adjust the O.D. at the end of the culture.
- Chef's tips and tricks:
- Pour into your plate 100 μL of fresh medium, use different glucose concentrations. You can do this by diluting your M9 medium (10g/L glc) with M9 WITHOUT glucose.
- Chef's tips and tricks:
- In our case we used 10 g/L, 5 g/L, 2 g/L, 1 g/L with a secondary carbon source: Laurate (final concentration of 5mM).
- Chef's tips and tricks:
- Add 5 μL of you culture according to your personal scheme.
- Now your plate is ready to be measured! Go to your favorite plate reader and program your sample. Set the temperature to 37°C. In a kinetic loop of a fixed time (17 hour 30 mins) the following measurements are taken with a time interval of 10 minutes with CONSTANT shaking: absorbance (600 nm filter) and GFP fluorescence (485 nm excitation and 520 nm emission). Followed by a delay of 100 seconds.
Data analysis
- Average you Biomass concentration for empty plasmid and your sample. Substract the blank value to your measurements.
- Average the fluorescence produced by your empty plasmid and your sample. Substract the fluorescence of the empty plasmid to the fluorescence given by your strain of interest. The BLANK is already included in this operation if it is all the time lower than the fluorescence measured in your empty plasmid.
- Plot the data points obtained and analyze the amount of GFP produced by your strain.
Protocol for growth of E.coli K12 on alkanes
Materials
- -80 °C stock or plate containing strain of interest
- 25 mL closed-cap glass flasks
- The to-be-tested alkanes
- LB-medium
- Phosphate buffer of pH 7.4 (wash buffer)
- M9 minimal medium (containing E2-salts)
Protocol
- Inoculate 5 mL of LB+antibiotic with your strains of interest. Grow up o/n.
- The following day subculture 500 μL of o/n culture in 50 mL of LB+antibiotic (1:100 dilution).
- Culture the cells up to an OD of about 0.3, spin down (4000 rpm, 10 min) and resuspend in 5 mL of wash buffer.
- Spin down (4000 rpm, 5 min), resuspend in 5 mL P-buffer and subculture 50 μL into 5 mL of M9-medium + E2 salts + 1% alkane.
- Measure OD600 at the start and compare to OD600 attained in time.
Resting-cell assays for E.coli
Materials
- -80 °C stock or plate containing strain of interest
- 100 mL sterile erlenmeyer flasks
- 25 mL closed-cap glass flasks
- The to-be-tested alkanes
- LB-medium + antibiotic
- Phosphate buffer of pH 7.4 (aka wash buffer)
- Phosphate buffer of pH 7.4 + E2 salts + 0.66% v/v glycerol (also known as RSA buffer)
- Incubator at 37 °C
Protocol
Don't forget to think about sterile conditions where needed!
- Inoculate 5 mL of LB+antibiotic with your strains of interest. Grow up o/n.
- The following day subculture 500 μL of o/n culture in 50 mL of LB+antibiotic (1:100 dilution).
- Culture the cells up to an OD of about 0.3, spin down (4.000 rpm, 10 min) and resuspend in 5 mL of wash buffer.
- Spin down (4000 rpm, 5 min) and resuspend in 20 mL of RSA buffer.
- Measure the OD600 (for reference to the OD600 at the end of the experiment).
- Make cell-mixture aliquots of 6 mL in 25 mL closed-cap glass flasks and add 100 nmoles of the desired alkane.
- Incubate the mixtures at the optimal enzyme temperature (below 37 °C!) for the desired reaction time
- Measure the OD600 afterwards.
Ethyl acetate extraction protocol
Materials
- Aqueous mixture from which hydrocarbons will be extracted
- Ethyl acetate (EtOAc)
- Anhydrous magnesium sulphate (MgSO4)
- Internal standard (undecane in our experiments)
- Vortex @1500 rpm
Protocol
Make sure all steps are performed using glassware! EtOAc dissolves plastics
- Prepare adequate amounts of EtOAc + 0.1% internal standard.
- Add 2.5 mL of the above preparation to 6 mL of the aqueous mixture
- Vortex each sample for 5 seconds @1500 rpm and let the two-phasic equilibrium establish
- If a detergent (such as Triton-X100) had been added to the aqueous mixture it is recommended to spin down the samples for 10 minutes @4000 rpm in order to get a proper two-phasic system
- Remove the organic layer (top) and dry the solvent using MgSO4
- Filter off the MgSO4 using a 0.2 micron filter and store at -20 °C in GC-vials.
General gas chromatography program for alkanes and alkanols
Materials
- Hydrocarbons in EtOAc to be analysed (dried over MgSO4)
- GC-vials
- GC-machine with CP-SIL 5CB column (length= 50 m)
- Operation system to run the machine
Protocol
Don't forget to fill the rinse vials with acetonitrile and methanol before use!
- Place GC-vials containing sample in the proper position on the GC rack.
- The column temperature program (shown below) has yielded nice peak separations for the alkanes/alkanols we've used. The calibration GC graph for the alkanes we've used is given below.
- Run batch and save results as pdf.
| Rate | Temperature [C] | Time [min] |
| 0 | 50 | 7.5 |
| 50 | 90 | 1.0 |
| 50 | 110 | 2.0 |
| 50 | 130 | 2.0 |
| 50 | 145 | 2.0 |
| 50 | 160 | 2.0 |
| 50 | 170 | 2.0 |
| 50 | 185 | 2.0 |
| 50 | 210 | 2.0 |
| 50 | 250 | 2.0 |
| 50 | 320 | 1.0 |
Enzyme activity assay for LadA by GC
Materials
- Lysates obtained from desired strains using our sonication protocol
- 50mM Tris-HCL solution with a pH of 7.5
- NADH,
- FMN,
- MgS04(*7H2O) and
- Triton X-100
- 25 mL closed-cap flasks
- Incubator at 37 °C and/or at 60 °C
- Boiling water
- Desired alkane (0.1% of total volume)
Protocol
Make sure all steps are performed using glassware! Alkanes may dissolve plastics.
- Add 0.1% v/v of the alkane to the Tris buffer and heat in boiling water for 5 minutes. Sonicate for 1 minute while still warm until a homogenous, viscous mixture is obtained.
- Add up to 1mM of NADH, 1mM FMN, 1mM MgSO4 and 0.01& v/v Triton X-100
- Make 6 mL aliquots in 25 mL closed-cap flasks
- Add adequate amounts of lysates (depends on Bradford assays, 1 mL taken for our experiments)
- Extract using the extraction protocol after desired reaction time (o/n in our experiments).
Enzyme activity assay for LadA by NADH absorption
Materials
- Cell extracts of desired strain
- Buffer 50 mM Tris containing 1mM NADH and 100nM FMN at pH 7.5
- Alkane (30 μL per well of heptadecane or hexadecane)
- 96 well plate
- Plate reader
Protocol
- Keep your extracts on ice.
- Add buffer to the wells that you are going to use.
- Add the alkane to be tested to the appropriate wells. KEEP IN MIND: Each well with alkane MUST HAVE A CONTROL WITHOUT SUBSTRATE. Besides you must have a blank (mixture without enzyme).
- Warm the plate up for 10-20 min at 37 °C (in the case of heptadecane). Keep in mind that this is a biphasic system; therefore we need a previous step in which you let the system reach an equilibrium.
- Add appropriate amounts of cell extracts to the buffer. Remember you are adding cell extract to a highly reduced compound which will easily be oxidized by anything else what is present in the lysate. In order to have enough measurement point per well (before everything going to 0) you must add very small amounts of lysate. For our experiments 5 μL was sufficient to yield slopes that could be further analysed. Parallel to this we've heat shocked the lysate at 60 °C for 10 minutes. This deactivates most of the E.coli oxidoreductases and should leave LadA in its native form, thus decreasing E.coli background 'noise'.
- Start absorption measurment at 340 nm immediately after adding lysates!
Buffers and Stock solutions
Luria-bertani (LB)
For 1 liter dissolve in H2O:
| Compound/Ingredient | Amount required (1L of medium) |
| Tryptone | 10 g |
| Yeast extract | 5 g |
| NaCl | 10 g |
| Agar+ | 15 g |
+ Add agar in case a solid medium is required.
Autoclave on a liquid cycle. Once broth reaches room temperature, store in cold room.
M9
For 1 liter dissolve in H2O:
| Compound/Ingredient | Amount required (1L of medium) |
| Na2HPO4 | 6 g |
| KH2PO4 | 3 g |
| NaCl | 0.5 g |
| (NH4)Cl | 1 g |
| Agar+ | 15 g |
+ Add agar in case a solid medium is required.
Once broth reaches room temperature, add to the broth in aseptic conditions the following amount of each solution:
| Compound/Ingredient | Amount required (1L of medium) |
| 1 M MgSO4 | 2 mL |
| 1 M CaCl2 | 0.1 mL |
| 20% Glucose (w/v)* | 10 mL |
* For M9-modified medium no glucose solution is added, instead 10 mL of filtered hydrocarbon/ hydrocarbon mixture are added per liter of medium. Be careful when adding the hydrocarbon/hydrocarbon mixture.
NOTE: Autoclave the solutions 1 M MgSO4, 1M CaCl2, 20% glucose (w/v) on a liquid cycle. Filter the solution Thiamine-HCl 1M using a 0.22 μm membrane, store the solution in a cold room when is not required.
Store in cold room if the medium is not used.
Chef's tips and tricks:
Additionally we added 1 mL of E2 salts to 1 Liter of our M9 medium, we did this expecting that we will eventually grow our cells on alkanes and some of our enzymes required cofactors that are present in this solution.
E2 Salts
For 1 liter dissolve in H2O:
| Compound/Ingredient | Amount required (1L of medium) |
| FeSO4.7H2O | 2.8 g |
| MnCl2 | 0.19g |
| CoCl2.6H2O | 2.8 g |
| CaCl2.4H2O | 1.84g |
| ZnSO4.7H2O | 0.28 g |
| CuSO4 | 0.16g |
Filter this solution using a 0.25 um membrane, and add 1mL/L of M9 medium.
Antibiotics
| Antibiotic | Final concentration in the medium |
| Ampicillin | 100 μg/mL |
| Kanamycin | 50 μg/mL |
| Chlorampenicol | 35 μg/mL |
| Tetracycline | 15 μg/mL |
All stock solutions must be kept at -20 ºC and used under aseptic conditions.
SOB
For 1 liter dissolve in H2O:
| Compound/Ingredient | Amount required (1L of medium) |
| Bacto tryptone | 20 g |
| Yeast extract | 5 g |
| NaCl | 0.5 |
| 250 mM KCl | 10 mL |
Adjust pH to 7.0. Before use add 5 mL of 2 mM MgCl2
SOC (Super optimal broth with catabolite repression)
Add 20 mM glucose to 1 L SOB
10x TBE Buffer
For 1 liter, dissolve in 800 mL H2O:
| Compound/Ingredient | Amount required (1L of medium) |
| Tris | 54 g |
| Boric Acid | 27.5 g |
| 0.5 M EDTA | 4.65 g or 20 mL |
Adjust volume to 1 liter with H2O
Potassium Laurate 500 mM
Mix in the same tube:
| Compound/Ingredient | Amount required |
| KOH 2M | 2500 µL |
| Lauric acid (melted at 50ºC) | 1136 µL |
| miliQ water | 6.364 mL |
Chef's tips and tricks:
Warm the mixture up, DON'T SHAKE IT VIGOROUSLY otherwise you'll get a lot of foam. Mix gently and keep it at 50ºC until you won't see any crystal or second phase (around 1 hour after the addition of lauric acid). After your mixture becomes a transparent liquid, you can store it at 4ºC or keep it at room temperature without problems.
Glycine 570 mM
- Weight 2.1375 g of anhydrous glycine.
- Add the miliQ water required for 50 mL of solution.
Tris 500 mM
- Weight 3.039 g of anhydrous Tris.
- Add the miliQ water required for 50 mL of solution.
NAD 50 mg/mL
- Weight 50 mg of anhydrous NAD.
- Add the miliQ water required for 1 mL of solution.
Disruption buffer
- Add 5 mL of solution 500 mM of Tris
- Add 40 mL of miliQ water
- Adjust the pH by adding 1M HCl until you reach the desired pH (7.4-7.5)
- Add miliQ water in order to complete a volume of 50 mL of solution
ADH/ALDH reaction buffer
For 25 mL solution Glycine 57mM, NAD 1mM, pH 9.5 dissolve in 20 mL H2O:
| Compound/Ingredient | Amount required |
| 570 mM glycine 570 mM of glycine solution | 2.5 mL |
| 50 mg/mL NAD solution | 331.5 µL |
- Adjust the pH using a solution of KOH 2M
- Add miliQ water in order to complete a volume of 25 mL of solution
Primers
| Name | Direction | Sequence |
| G00100 | Forward | 5’ TGCCACCTGACGTCTAAGAA 3’ |
| G00101 | Reverse | 5’ ATTACCGCCTTTGAGTGAGC 3’ |
| MF | Forward | 5’ TTTCCCAGTCACGACGTTG 3’ |
| MR2 | Reverse | 5’ GCAAGGAAACAGCTATGACCATG 3’ |
| PCR F | Forward | 5’ TCTGAATTCGCGGCCGCTTCT 3’ |
| PCR R | Reverse | 5’ GAACTGCAGCGGCCGCTACTAGTA 3’ |
| 304F | Forward | 5’ CCGTAACCTATGGTGAAAATTTTACTAGAGTCACACAGGAAAGTACTAGTAGCG 3’ |
| 304R | Reverse | 5’ CGCTACTAGTACTTTCCTGTGTGACTCTAGTAAAATTTTCACCATAGGTTACGG 3’ |
| 327F | Forward | 5’ CGGTGAAGTTGCATACTATCGATATACTAGAGTCACACAGGAAAGTACTAGTAGCG 3’ |
| 327R | Reverse | 5’ CGCTACTAGTACTTTCCTGTGTGACTCTAGTATATCGATAGTATGCAACTTCACCG 3’ |
 "
"