Team:TU Delft/Project/rbs-characterization/results
From 2010.igem.org
(→The Results) |
(→The Results) |
||
| Line 28: | Line 28: | ||
<html><center><img src="https://static.igem.org/mediawiki/2010/0/00/TU_Delft_project_navigation.jpg" usemap="#projectnavigation" border="0" /></center><map id="projectnavigation" name="projectnavigation"><area shape="rect" alt="Characterization" title="" coords="309,3,591,45" href="https://2010.igem.org/Team:TU_Delft#page=Project/rbs-characterization/characterization" target="" /><area shape="rect" alt="Results" title="" coords="609,3,891,44" href="https://2010.igem.org/Team:TU_Delft#page=Project/rbs-characterization/results" target="" /><area shape="rect" alt="Parts" title="" coords="9,3,290,44" href="https://2010.igem.org/Team:TU_Delft#page=Project/rbs-characterization/parts" target="" /></map></html> | <html><center><img src="https://static.igem.org/mediawiki/2010/0/00/TU_Delft_project_navigation.jpg" usemap="#projectnavigation" border="0" /></center><map id="projectnavigation" name="projectnavigation"><area shape="rect" alt="Characterization" title="" coords="309,3,591,45" href="https://2010.igem.org/Team:TU_Delft#page=Project/rbs-characterization/characterization" target="" /><area shape="rect" alt="Results" title="" coords="609,3,891,44" href="https://2010.igem.org/Team:TU_Delft#page=Project/rbs-characterization/results" target="" /><area shape="rect" alt="Parts" title="" coords="9,3,290,44" href="https://2010.igem.org/Team:TU_Delft#page=Project/rbs-characterization/parts" target="" /></map></html> | ||
==The Results== | ==The Results== | ||
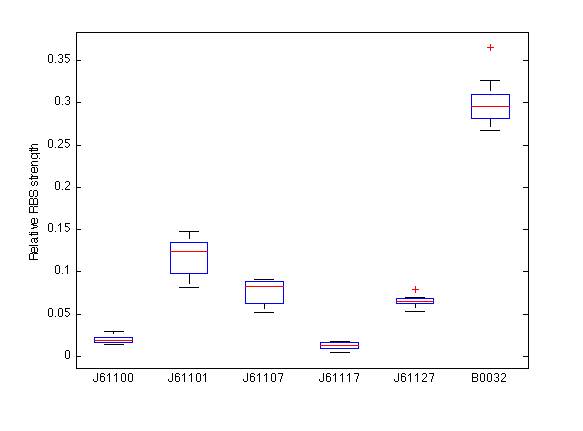
| - | In the boxplot below the RBS strengths for the five tested members of the Anderson family are given along with the deviations from the mean. The strengths are given relative to the community RBS BBa_B0034, which has a strength of unity (1). This relationship could be built by our characterizing the community RBS [http://partsregistry.org/Part:BBa_B0032 BBa_B0032] simultaneously, which is known to have a strength of 0.3 relative to [http://partsregistry.org/Part:BBa_B0032 BBa_B0034]. Therefore in the plot below, the mean value of BBa_B0032 is fixed at 0.3. | + | In the boxplot below the RBS strengths for the five tested members of the Anderson family are given along with the deviations from the mean. The strengths are given relative to the community RBS BBa_B0034, which has a strength of unity (1). This relationship could be built by our characterizing the community RBS [http://partsregistry.org/Part:BBa_B0032 BBa_B0032] simultaneously, which is known to have a strength (or efficiency) of 0.3 relative to [http://partsregistry.org/Part:BBa_B0032 BBa_B0034]. Therefore in the plot below, the mean value of BBa_B0032 is fixed at 0.3. |
[[Image:TUDelft2010_Rbs_strength_boxplot.png]] | [[Image:TUDelft2010_Rbs_strength_boxplot.png]] | ||
| Line 34: | Line 34: | ||
<html><div id="rbs-results"></html> | <html><div id="rbs-results"></html> | ||
| - | + | ==Conclusions== | |
| + | We've succeeded in characterizing five memebers of the Anderson ribosome binding site family by simple culturing on a 96 well plate while measuring GFP fluorescence and biomass absorbance. Fitting the obtained data to our improved [https://2010.igem.org/Team:TU_Delft#page=Project/rbs-characterization/characterization protein production model] yielded RBS strengths values with negligible standard deviations. | ||
| + | |||
{| | {| | ||
|RBS | |RBS | ||
Revision as of 10:03, 24 October 2010

The Results
In the boxplot below the RBS strengths for the five tested members of the Anderson family are given along with the deviations from the mean. The strengths are given relative to the community RBS BBa_B0034, which has a strength of unity (1). This relationship could be built by our characterizing the community RBS BBa_B0032 simultaneously, which is known to have a strength (or efficiency) of 0.3 relative to BBa_B0034. Therefore in the plot below, the mean value of BBa_B0032 is fixed at 0.3.
Conclusions
We've succeeded in characterizing five memebers of the Anderson ribosome binding site family by simple culturing on a 96 well plate while measuring GFP fluorescence and biomass absorbance. Fitting the obtained data to our improved protein production model yielded RBS strengths values with negligible standard deviations.
| RBS | Mean strength | Standard deviation |
| J61100 | 0.026560 | 0.005126 |
| J61117 | 0.123819 | 0.024725 |
| J61101 | 0.081210 | 0.013402 |
| J61107 | 0.019763 | 0.003963 |
| J61127 | 0.069657 | 0.007422 |
| B0032 | 0.300000 | 0.028316 |

 "
"