Team:TU Delft/Project/rbs-characterization/parts
From 2010.igem.org

BioBricks, the making of the RBS Characterization
Experimental setup
The following RBS sequences from the Anderson RBS family were used:
For the purpose of comparison and standardization the following RBS was used as a reference for our characterization.
These RBS sequences were positioned upstream of the standard GFP coding sequence, so expression could be measured.
BBa_I13401 was PCR amplified using the universal primers G00100 and G00101. The purified PCR product was assembled into one of the Anderson RBS plasmids provided in the distribution plates by means of 2-way ligations.
Transformation into competent Top10 E.coli cells yielded positives, as determined by fluorescence analysis.
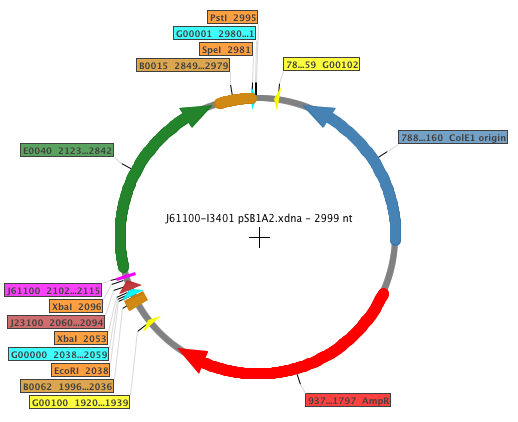
The Biobricks generated in order to perform the experiments were: K398500, K398501, K398502, K398503, K398504. The general map of the construction is shown below, where the RBS is displayed in fucsia.
| Feature | Function |
| AmpR | Ampicillin resistance |
| B0015 | Transcriptional (double) terminator |
| B0062 | Transcriptional terminator |
| E0040 | GFP |
| G00000 | Standard prefix |
| G00001 | Standard suffix |
| G00100 | VF2 primer binding site |
| G00101 | VR primer binding site |
| J61100 | RBS Anderson family |
| J23100 | Promoter |
Parts
Our constructs used for measurements: <partinfo>K398500 SpecifiedComponents</partinfo>
<partinfo>K398501 SpecifiedComponents</partinfo>
<partinfo>K398502 SpecifiedComponents</partinfo>
<partinfo>K398503 SpecifiedComponents</partinfo>
<partinfo>K398504 SpecifiedComponents</partinfo>
 "
"