Team:TU Delft/Project/rbs-characterization/parts
From 2010.igem.org
(→Experimental setup) |
(→The Reference Construct) |
||
| Line 25: | Line 25: | ||
====The Reference Construct==== | ====The Reference Construct==== | ||
| + | [http://partsregistry.org/Part:BBa_J23100 BBa_J23100], which is contained in the pSB1A2 plasmid was digested using SpeI and PstI. [http://partsregistry.org/Part:BBa_E0240 BBa_E0240] was digested using XbaI and PstI and ligated with the digest product of J23100. Transformation into <i>E.coli</i> TOP10 yielded positives as analyzed by UV-light. | ||
| - | + | ====The BioBricks==== | |
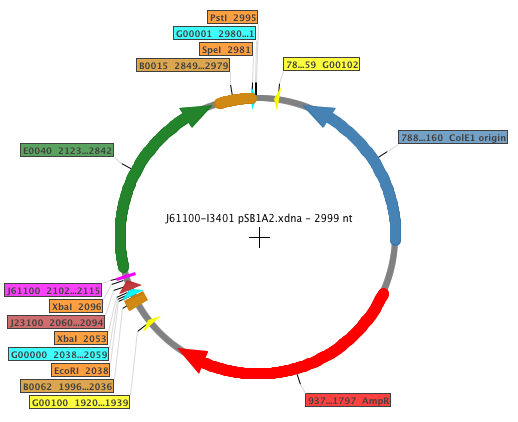
| - | The Biobricks generated | + | The Biobricks generated for the measurements were: [http://partsregistry.org/Part:BBa_K398500 K398500], [http://partsregistry.org/Part:BBa_K398501 K398501], [http://partsregistry.org/Part:BBa_K398502 K398502], [http://partsregistry.org/Part:BBa_K398503 K398503], [http://partsregistry.org/Part:BBa_K398504 K398504] and [http://partsregistry.org/Part:BBa_K173000 K173000]. A general example map of the construction is shown below, where the RBS is displayed in fucsia. |
[[Image:RBS1.jpg|left|650px]] | [[Image:RBS1.jpg|left|650px]] | ||
Revision as of 10:41, 24 October 2010

RBS measurement parts
Experimental setup
The following RBS sequences from the Anderson RBS family were used:
For the purpose of comparison and standardization the following RBS was used as a reference for our characterization.
These RBS sequences were positioned upstream of the standard GFP coding sequence, in order to be able to measure expression.
Cloning method
The Anderson Constructs
BBa_I13401 was PCR amplified by using the universal primers G00100 and G00101. The purified PCR product was back-digested using XbaI and PstI. The Anderson RBSs are provided in pSB1A2 plasmids. These were digested using SpeI and PstI and ligated with the digested I13401 insert. Transformation into competent Top10 E.coli cells yielded positives, as determined by UV-light plate analysis.
The Reference Construct
BBa_J23100, which is contained in the pSB1A2 plasmid was digested using SpeI and PstI. BBa_E0240 was digested using XbaI and PstI and ligated with the digest product of J23100. Transformation into E.coli TOP10 yielded positives as analyzed by UV-light.
The BioBricks
The Biobricks generated for the measurements were: K398500, K398501, K398502, K398503, K398504 and K173000. A general example map of the construction is shown below, where the RBS is displayed in fucsia.
| Feature | Function |
| AmpR | Ampicillin resistance |
| B0015 | Transcriptional (double) terminator |
| B0062 | Transcriptional terminator |
| E0040 | GFP |
| G00000 | Standard prefix |
| G00001 | Standard suffix |
| G00100 | VF2 primer binding site |
| G00101 | VR primer binding site |
| J61100 | RBS Anderson family |
| J23100 | Promoter |
Parts
Our constructs used for measurements: <partinfo>K398500 SpecifiedComponents</partinfo>
<partinfo>K398501 SpecifiedComponents</partinfo>
<partinfo>K398502 SpecifiedComponents</partinfo>
<partinfo>K398503 SpecifiedComponents</partinfo>
<partinfo>K398504 SpecifiedComponents</partinfo>
 "
"