Team:TU Delft/Modeling/MFA/explanation
From 2010.igem.org
(→Metabolic Flux Analysis, how does it work) |
|||
| Line 1: | Line 1: | ||
| + | __NOTOC__ | ||
| + | {{Team:TU_Delft/frame_check}} | ||
==Metabolic Flux Analysis, how does it work== | ==Metabolic Flux Analysis, how does it work== | ||
| - | |||
Metabolic flux analysis (MFA) is a tool to study networks quantitatively. It can be used to calculate theoretical maximal yields. For our MFA we useed CellNetAnalyzer, a tool for Matlab. CellNetAnalyzer already has a simplified, yet extensive metabolic network of ''E. coli''. To this network the hydrocarbon degradation pathway was added. After this, further pathways were added to the network see how efficient certain products could be produced and alternative electron acceptor was investigated. | Metabolic flux analysis (MFA) is a tool to study networks quantitatively. It can be used to calculate theoretical maximal yields. For our MFA we useed CellNetAnalyzer, a tool for Matlab. CellNetAnalyzer already has a simplified, yet extensive metabolic network of ''E. coli''. To this network the hydrocarbon degradation pathway was added. After this, further pathways were added to the network see how efficient certain products could be produced and alternative electron acceptor was investigated. | ||
| Line 15: | Line 16: | ||
The change in the concentration of the substrate and products outside the cell can easily be calculated from measurements. This results in uptake and excretion speeds. The intracellelular rates (inside the cell) can not be figured out as easily, but if enough information is known from the uptake en excretion rates, the intracellular rates can often be calculated. | The change in the concentration of the substrate and products outside the cell can easily be calculated from measurements. This results in uptake and excretion speeds. The intracellelular rates (inside the cell) can not be figured out as easily, but if enough information is known from the uptake en excretion rates, the intracellular rates can often be calculated. | ||
| - | |||
===Units in MFA=== | ===Units in MFA=== | ||
| - | |||
The term 'rate' has occured in the previous alinea and needs to be specified further. Rates indicate how fast a reaction goes. Rates are usually defined in moles per amount of biomass per time. In CellNetAnalyzer the units 'mmol per gram of dry weight biomass per hour' were used. The reaction that genereates biomass, called μ, is an exception and has the unit per hour, which is derived from biomass produced per biomass per hour. | The term 'rate' has occured in the previous alinea and needs to be specified further. Rates indicate how fast a reaction goes. Rates are usually defined in moles per amount of biomass per time. In CellNetAnalyzer the units 'mmol per gram of dry weight biomass per hour' were used. The reaction that genereates biomass, called μ, is an exception and has the unit per hour, which is derived from biomass produced per biomass per hour. | ||
| - | |||
===Steady-state assumption=== | ===Steady-state assumption=== | ||
| - | |||
In order to determine the intracellular rates, it is assumed that all the intermediates inside the cell are in a steady-state. This means that the amount of the substance is constant, meaning that the total production and the total consumption of a substance are equal to each other. This holds also for co-factors such as ATP and NADH which are used by many reactions. This means that all the rates need to have a ratio that balances out the co-factor production and consumption. | In order to determine the intracellular rates, it is assumed that all the intermediates inside the cell are in a steady-state. This means that the amount of the substance is constant, meaning that the total production and the total consumption of a substance are equal to each other. This holds also for co-factors such as ATP and NADH which are used by many reactions. This means that all the rates need to have a ratio that balances out the co-factor production and consumption. | ||
| - | |||
===Full flux determination=== | ===Full flux determination=== | ||
| - | + | With this analysis all the intracellular fluxes are calculated to fit with a set of measured transport fluxes. For figure 1 this would mean that, if it is known that 1 mmol gx<sup>-1</sup> h<sup>-1</sup> of glucose is going in and 1.2 mmol gx<sup>-1</sup> h<sup>-1</sup> of CO<sub>2</sub> is coming out, then the other rates can be calculated. The calculation results in 1.32 mmol gx<sup>-1</sup> h<sup>-1</sup> of ethanol coming out and 0.36 mmol gx<sup>-1</sup> h<sup>-1</sup> goes through intermediate 2 to the biomass. This is essentially the metabolic flux analysis. For larger networks, however it is required to used lineair algebra or tools such as Cell Net Analyzer. | |
| - | With this analysis all the intracellular fluxes are calculated to fit with a set of measured transport fluxes. For figure 1 this would mean that, if it is known that 1 mmol gx | + | |
| - | + | ||
===Flux optimization=== | ===Flux optimization=== | ||
| + | This is the analysis that is used in our iGEM project. This analysis does not try to calculated all the intracellular fluxes with a set of measured transport fluxes, instead it tries to optimize a (set of) fluxes to a set of transport fluxes. To calculate a maximal theoretical yield of a product on a substrate, the production rate needs to be optimized toward a certain substrate uptake rate. | ||
| - | + | Production by a metabolic network can be optimized with a built in function of CellNetAnalyzer. This function will calculate with a given set of substrate inflows the maximal rate for a product. An advantage of this is that not all the transport rates need to be known, the function simply finds an optimal solution. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | For example, in figure 1 there is a inflow of 1 mmol gx<sup>-1</sup> h<sup>-1</sup> substrate and the rate for product 2 needs to be maximized. The function will then assign values to all the reaction rates to make the outflow of product 2 as large as possible, while still fulfilling the pseudo-steady-state condition. For this example the outflow of product 2 will be 0.6 mmol gx<sup>-1</sup> h<sup>-1</sup>. The rest goes into biomass. The maximal yield is then 0.6 mol product 2 per mol substrate. | |
| + | It is also possible to optimize the growth by maximizing μ. If there is no feasible solution for this, then the cell is not viable because it can not use the substrate for growth. This may occur for example when a lot of NADH is produced, but cannot be consumed sufficiently somewhere else in the network. Because of this the network can not maintain the pseudo-steady-state condition. This situation is called redox stress. | ||
| - | [[Team:TU_Delft/Modeling/MFA|Go back now you are enlightened about how | + | [[Team:TU_Delft/Modeling/MFA|Go back to the MFA overview, now you are enlightened about how it works]] |
Revision as of 08:24, 20 October 2010
Metabolic Flux Analysis, how does it work
Metabolic flux analysis (MFA) is a tool to study networks quantitatively. It can be used to calculate theoretical maximal yields. For our MFA we useed CellNetAnalyzer, a tool for Matlab. CellNetAnalyzer already has a simplified, yet extensive metabolic network of E. coli. To this network the hydrocarbon degradation pathway was added. After this, further pathways were added to the network see how efficient certain products could be produced and alternative electron acceptor was investigated.
Metabolic networks - definition
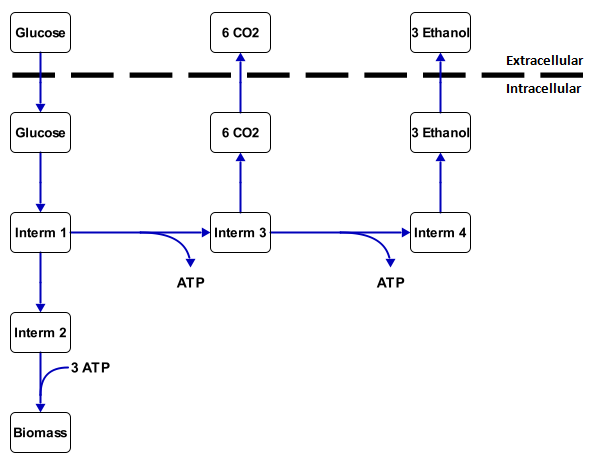
A metabolic network is the collection of all the reactions that occur inside a cell. All these reactions are coupled and form a network together. This network has a couple of transport reactions with substrate (food, such as sugar and alkanes) as an input and products (such as biomass, CO2, ethanol) as output. Other products of the metabolic network are energy, which is stored in molecules such as ATP and NADH, and building blocks for the cell (biomass). The substances between the substrates and the products are called intermediates. An example of this is in figure 1.
Figure 1: Small made-up metabolic network
The change in the concentration of the substrate and products outside the cell can easily be calculated from measurements. This results in uptake and excretion speeds. The intracellelular rates (inside the cell) can not be figured out as easily, but if enough information is known from the uptake en excretion rates, the intracellular rates can often be calculated.
Units in MFA
The term 'rate' has occured in the previous alinea and needs to be specified further. Rates indicate how fast a reaction goes. Rates are usually defined in moles per amount of biomass per time. In CellNetAnalyzer the units 'mmol per gram of dry weight biomass per hour' were used. The reaction that genereates biomass, called μ, is an exception and has the unit per hour, which is derived from biomass produced per biomass per hour.
Steady-state assumption
In order to determine the intracellular rates, it is assumed that all the intermediates inside the cell are in a steady-state. This means that the amount of the substance is constant, meaning that the total production and the total consumption of a substance are equal to each other. This holds also for co-factors such as ATP and NADH which are used by many reactions. This means that all the rates need to have a ratio that balances out the co-factor production and consumption.
Full flux determination
With this analysis all the intracellular fluxes are calculated to fit with a set of measured transport fluxes. For figure 1 this would mean that, if it is known that 1 mmol gx-1 h-1 of glucose is going in and 1.2 mmol gx-1 h-1 of CO2 is coming out, then the other rates can be calculated. The calculation results in 1.32 mmol gx-1 h-1 of ethanol coming out and 0.36 mmol gx-1 h-1 goes through intermediate 2 to the biomass. This is essentially the metabolic flux analysis. For larger networks, however it is required to used lineair algebra or tools such as Cell Net Analyzer.
Flux optimization
This is the analysis that is used in our iGEM project. This analysis does not try to calculated all the intracellular fluxes with a set of measured transport fluxes, instead it tries to optimize a (set of) fluxes to a set of transport fluxes. To calculate a maximal theoretical yield of a product on a substrate, the production rate needs to be optimized toward a certain substrate uptake rate.
Production by a metabolic network can be optimized with a built in function of CellNetAnalyzer. This function will calculate with a given set of substrate inflows the maximal rate for a product. An advantage of this is that not all the transport rates need to be known, the function simply finds an optimal solution.
For example, in figure 1 there is a inflow of 1 mmol gx-1 h-1 substrate and the rate for product 2 needs to be maximized. The function will then assign values to all the reaction rates to make the outflow of product 2 as large as possible, while still fulfilling the pseudo-steady-state condition. For this example the outflow of product 2 will be 0.6 mmol gx-1 h-1. The rest goes into biomass. The maximal yield is then 0.6 mol product 2 per mol substrate.
It is also possible to optimize the growth by maximizing μ. If there is no feasible solution for this, then the cell is not viable because it can not use the substrate for growth. This may occur for example when a lot of NADH is produced, but cannot be consumed sufficiently somewhere else in the network. Because of this the network can not maintain the pseudo-steady-state condition. This situation is called redox stress.
Go back to the MFA overview, now you are enlightened about how it works
 "
"