Team:TU Delft/8 July 2010 content
From 2010.igem.org
(Difference between revisions)
(→Ordered DNA stocks) |
|||
| Line 4: | Line 4: | ||
We harvested the 200 mL bacterial cells of the [https://2010.igem.org/Team:TU_Delft#page=pages/blog&blog=6_July_2010 16 DNA parts]. We used 3 mL of the bacterial cells to make [[Team:TU_Delft/protocols/freezing_bacterial_stocks|-80 °C stocks]]. With the rest we centrifuged at 4,000 rmp for 15 minutes and stored the pellets in the -20 °C. | We harvested the 200 mL bacterial cells of the [https://2010.igem.org/Team:TU_Delft#page=pages/blog&blog=6_July_2010 16 DNA parts]. We used 3 mL of the bacterial cells to make [[Team:TU_Delft/protocols/freezing_bacterial_stocks|-80 °C stocks]]. With the rest we centrifuged at 4,000 rmp for 15 minutes and stored the pellets in the -20 °C. | ||
| - | + | ==Characterization of Anderson RBS sequences== | |
[https://2010.igem.org/Team:TU_Delft#page=pages/blog&blog=6_July_2010 Tuesday's] ligation products of BioBricks K398500-K398504 yielded successful transformants. [[Team:TU_Delft/protocols/colony_PCR|Single colony PCRs]] were performed and loaded onto a gel. The bands on the [[Team:TU_Delft/protocols/agarose_gel |1% agarose gel]] indicated the presence of inserts with the proper lengths: | [https://2010.igem.org/Team:TU_Delft#page=pages/blog&blog=6_July_2010 Tuesday's] ligation products of BioBricks K398500-K398504 yielded successful transformants. [[Team:TU_Delft/protocols/colony_PCR|Single colony PCRs]] were performed and loaded onto a gel. The bands on the [[Team:TU_Delft/protocols/agarose_gel |1% agarose gel]] indicated the presence of inserts with the proper lengths: | ||
Latest revision as of 19:40, 5 August 2010
Contents |
Lab work
Ordered DNA stocks
We harvested the 200 mL bacterial cells of the 16 DNA parts. We used 3 mL of the bacterial cells to make -80 °C stocks. With the rest we centrifuged at 4,000 rmp for 15 minutes and stored the pellets in the -20 °C.
Characterization of Anderson RBS sequences
Tuesday's ligation products of BioBricks K398500-K398504 yielded successful transformants. Single colony PCRs were performed and loaded onto a gel. The bands on the 1% agarose gel indicated the presence of inserts with the proper lengths:
Lane descriptions:
| # | Description | Expected Length (bp) | Primers | Status | Remarks |
| 1 | transformant #1 of ligation mix K398500 | 1158 | G00100 + G00101 | ✓ | Unexpected band at 2700 |
| 2 | transformant #2 of ligation mix K398500 | 1158 | G00100 + G00101 | ✓ | |
| 3 | transformant #1 of ligation mix K398501 | 1158 | G00100 + G00101 | ✓ | |
| 4 | transformant #2 of ligation mix K398501 | 1158 | G00100 + G00101 | ✓ | |
| 5 | transformant #1 of ligation mix K398502 | 1158 | G00100 + G00101 | ✓ | |
| 6 | transformant #2 of ligation mix K398502 | 1158 | G00100 + G00101 | ✓ | |
| 7 | transformant #1 of ligation mix K398503 | 1158 | G00100 + G00101 | ✓ | |
| 8 | transformant #2 of ligation mix K398503 | 1158 | G00100 + G00101 | ✓ | probably run of the gel |
| 9 | transformant #1 of ligation mix K398504 | 1158 | G00100 + G00101 | ✓ | |
| 10 | transformant #2 of ligation mix K398504 | 1158 | G00100 + G00101 | ✓ | Sample not fully loaded on gel |
| 11 | transformant #1 of ligation control | G00100 + G00101 | probably run of the gel | ||
| 12 | transformant #2 of ligation control | G00100 + G00101 | |||
| 13 | transformant #1 of digestion control | G00100 + G00101 | probably run of the gel | ||
| 14 | transformant #2 of digestion control | G00100 + G00101 | |||
| M1 | SmartLadder Marker | n/a | n/a | n/a |
The colonies belonging to lanes 2, 4, 6, 8 and 10 were plated out on AMP plates to yield reincultures for subsequent characterization experiments.
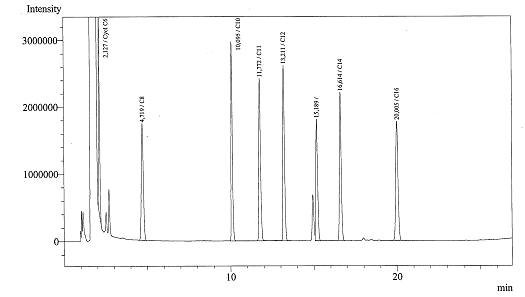
Alkane degradation
Today we got the Gas Chromatograph working, we could identify several peaks. Now we are ready for the real experiments!
 "
"