Team:TU Delft/19 July 2010 content
From 2010.igem.org
(→Emulsifier) |
|||
| Line 1: | Line 1: | ||
| - | =Lab work= | + | ==Lab work== |
| - | + | ||
| + | <h4>Alkane degradation</h4> | ||
Biobricks in production: | Biobricks in production: | ||
| Line 13: | Line 14: | ||
|1 | |1 | ||
|2 μg J61100 + EcoRI + SpeI | |2 μg J61100 + EcoRI + SpeI | ||
| - | | | + | |Buffer 2 (BioLabs) |
|‘E – J61100 – S’ | |‘E – J61100 – S’ | ||
|- | |- | ||
|2 | |2 | ||
|2 μg J61100 + EcoRI + SpeI | |2 μg J61100 + EcoRI + SpeI | ||
| - | | | + | |Buffer 2 (BioLabs) |
|‘E – J61100 – S’ | |‘E – J61100 – S’ | ||
|- | |- | ||
|3 | |3 | ||
|1 μg rubR + EcoRI + SpeI | |1 μg rubR + EcoRI + SpeI | ||
| - | | | + | |Buffer 2 (BioLabs) |
|‘E – rubR– S’ | |‘E – rubR– S’ | ||
|- | |- | ||
|4 | |4 | ||
|1 μg J61101 + EcoRI + SpeI | |1 μg J61101 + EcoRI + SpeI | ||
| - | | | + | |Buffer 2 (BioLabs) |
|‘E – J61101– S’ | |‘E – J61101– S’ | ||
|- | |- | ||
|5 | |5 | ||
|1 μg J61107 + EcoRI + SpeI | |1 μg J61107 + EcoRI + SpeI | ||
| - | | | + | |Buffer 2 (BioLabs) |
| - | |‘E – | + | |‘E – J61107 – S’ |
|- | |- | ||
|6 | |6 | ||
| - | | | + | |1 μg alkB2 + Xbal + PstI |
| - | | | + | |Buffer 2 (BioLabs) |
|‘X – alkB2 – P’ | |‘X – alkB2 – P’ | ||
|- | |- | ||
|7 | |7 | ||
| - | | 1 μg rubA3 + Xbal + PstI | + | |1 μg rubA3 + Xbal + PstI |
| - | | | + | |Buffer 2 (BioLabs) |
|‘X – rubA3 – P’ | |‘X – rubA3 – P’ | ||
|- | |- | ||
|8 | |8 | ||
|1 μg rubA4 + Xbal + PstI | |1 μg rubA4 + Xbal + PstI | ||
| - | | | + | |Buffer 2 (BioLabs) |
|‘X – rubA4 – P’ | |‘X – rubA4 – P’ | ||
|- | |- | ||
|9 | |9 | ||
|1 μg B0015 + Xbal + PstI | |1 μg B0015 + Xbal + PstI | ||
| - | | | + | |Buffer 2 (BioLabs) |
|‘X – B0015 – P’ | |‘X – B0015 – P’ | ||
|- | |- | ||
|10 | |10 | ||
|1 μg ladA + Xbal + PstI | |1 μg ladA + Xbal + PstI | ||
| - | | | + | |Buffer 2 (BioLabs) |
|‘X – ladA – P’ | |‘X – ladA – P’ | ||
|- | |- | ||
|11 | |11 | ||
|1 μg ADH + Xbal + PstI | |1 μg ADH + Xbal + PstI | ||
| - | | | + | |Buffer 2 (BioLabs) |
|‘X – ADH – P’ | |‘X – ADH – P’ | ||
|- | |- | ||
|12 | |12 | ||
|1 μg ALDH + Xbal + PstI | |1 μg ALDH + Xbal + PstI | ||
| - | | | + | |Buffer 2 (BioLabs) |
|‘X – ALDH – P’ | |‘X – ALDH – P’ | ||
|- | |- | ||
|13 | |13 | ||
|3 μg pSB1T3 + EcoRI + PstI | |3 μg pSB1T3 + EcoRI + PstI | ||
| - | | | + | |Buffer 2 (BioLabs) |
| - | |‘E – | + | |‘E – ---- – P’ |
|} | |} | ||
| - | + | <h4>Emulsifier</h4> | |
| - | Since last weeks attempt to construct the inducible protomer R0011 with RBS B0032 failed, we decided to take a different approach. Instead of doing the standard 2 parts assembly, we amplify the promoter by PCR digest it and ligate in the plasmid that contains the RBS which has only be cut open at the left side. By doing so we do not risk losing the super small DNA fragments like the RBS. The problem is that we cannot use antibiotics selection. So we need to determine that by | + | Since last weeks attempt to construct the inducible protomer R0011 with RBS B0032 failed, we decided to take a different approach. Instead of doing the standard 2 parts assembly, we amplify the promoter by PCR digest it and ligate in the plasmid that contains the RBS which has only be cut open at the left side. By doing so we do not risk losing the super small DNA fragments like the RBS. The problem is that we cannot use antibiotics selection. So we need to determine that by colony PCR and sequencing. |
'''PCR Amplification''' | '''PCR Amplification''' | ||
| - | [ | + | R0011 was [[Team:TU_Delft/protocols/PCR|amplified]] with the universal primers G00100 and G00101. The product was put on [[Team:TU_Delft/protocols/agarose_gel|1% agarose gel]]: |
[[Image:TU_Delft_P1_2010-07-19_PCR_Product_promotor.png|thumb|right|1% agarose gel]] | [[Image:TU_Delft_P1_2010-07-19_PCR_Product_promotor.png|thumb|right|1% agarose gel]] | ||
| Line 91: | Line 92: | ||
|'''#''' | |'''#''' | ||
|'''Description''' | |'''Description''' | ||
| - | |||
|- | |- | ||
|1 | |1 | ||
| - | |BioRad EZ Ladder | + | |BioRad EZ Ladder (5 μL) |
| - | + | ||
|- | |- | ||
|2 | |2 | ||
| - | |PCR Product | + | |PCR Product of R0011 (10 μL + 2 μL Loading buffer) |
| - | + | ||
|} | |} | ||
| Line 106: | Line 104: | ||
'''Digestion''' | '''Digestion''' | ||
| - | The PCR product of promotor R0011 was cut with EcoRI and SpeI and the RBS containing plasmid has been cut open with EcoRI and Xbal at 37 | + | The PCR product of promotor R0011 was cut with EcoRI and SpeI and the RBS containing plasmid has been cut open with EcoRI and Xbal for 2.5 hours at 37 °C. This deviates from the standard biobrick assembly, thus is not completely as described in our [[Team:TU_Delft/protocols/restriction_enzyme_digestion|digestion protocol]]. |
{| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | {| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | ||
| Line 116: | Line 114: | ||
|1 | |1 | ||
|1 μg B0032 + EcoRI + Xbal: | |1 μg B0032 + EcoRI + Xbal: | ||
| - | |Buffer 2 | + | |Buffer 2 (BioLabs) |
|‘X – B0032 - Plasmid backbone – E’ | |‘X – B0032 - Plasmid backbone – E’ | ||
|- | |- | ||
|2 | |2 | ||
| - | |30 | + | |30 μL PCR product of R0011 + EcoRI + SpeI |
|Buffer 2 | |Buffer 2 | ||
|‘E - R0011 - S’ | |‘E - R0011 - S’ | ||
Revision as of 20:11, 19 July 2010
Lab work
Alkane degradation
Biobricks in production:
| # | Digestion reaction | Used Buffer | Needed fragment |
| 1 | 2 μg J61100 + EcoRI + SpeI | Buffer 2 (BioLabs) | ‘E – J61100 – S’ |
| 2 | 2 μg J61100 + EcoRI + SpeI | Buffer 2 (BioLabs) | ‘E – J61100 – S’ |
| 3 | 1 μg rubR + EcoRI + SpeI | Buffer 2 (BioLabs) | ‘E – rubR– S’ |
| 4 | 1 μg J61101 + EcoRI + SpeI | Buffer 2 (BioLabs) | ‘E – J61101– S’ |
| 5 | 1 μg J61107 + EcoRI + SpeI | Buffer 2 (BioLabs) | ‘E – J61107 – S’ |
| 6 | 1 μg alkB2 + Xbal + PstI | Buffer 2 (BioLabs) | ‘X – alkB2 – P’ |
| 7 | 1 μg rubA3 + Xbal + PstI | Buffer 2 (BioLabs) | ‘X – rubA3 – P’ |
| 8 | 1 μg rubA4 + Xbal + PstI | Buffer 2 (BioLabs) | ‘X – rubA4 – P’ |
| 9 | 1 μg B0015 + Xbal + PstI | Buffer 2 (BioLabs) | ‘X – B0015 – P’ |
| 10 | 1 μg ladA + Xbal + PstI | Buffer 2 (BioLabs) | ‘X – ladA – P’ |
| 11 | 1 μg ADH + Xbal + PstI | Buffer 2 (BioLabs) | ‘X – ADH – P’ |
| 12 | 1 μg ALDH + Xbal + PstI | Buffer 2 (BioLabs) | ‘X – ALDH – P’ |
| 13 | 3 μg pSB1T3 + EcoRI + PstI | Buffer 2 (BioLabs) | ‘E – ---- – P’ |
Emulsifier
Since last weeks attempt to construct the inducible protomer R0011 with RBS B0032 failed, we decided to take a different approach. Instead of doing the standard 2 parts assembly, we amplify the promoter by PCR digest it and ligate in the plasmid that contains the RBS which has only be cut open at the left side. By doing so we do not risk losing the super small DNA fragments like the RBS. The problem is that we cannot use antibiotics selection. So we need to determine that by colony PCR and sequencing.
PCR Amplification
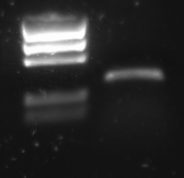
R0011 was amplified with the universal primers G00100 and G00101. The product was put on 1% agarose gel:
Lane description:
| # | Description |
| 1 | BioRad EZ Ladder (5 μL) |
| 2 | PCR Product of R0011 (10 μL + 2 μL Loading buffer) |
The PCR band is about 300 bps long. R0011 itself is just 55 bp, but the flanking primer regions are about 100 bp each.
Digestion
The PCR product of promotor R0011 was cut with EcoRI and SpeI and the RBS containing plasmid has been cut open with EcoRI and Xbal for 2.5 hours at 37 °C. This deviates from the standard biobrick assembly, thus is not completely as described in our digestion protocol.
| # | Digestion reaction | Used Buffer | Needed fragment |
| 1 | 1 μg B0032 + EcoRI + Xbal: | Buffer 2 (BioLabs) | ‘X – B0032 - Plasmid backbone – E’ |
| 2 | 30 μL PCR product of R0011 + EcoRI + SpeI | Buffer 2 | ‘E - R0011 - S’ |
Ligation
The digestion products were ligated over night:
| # | Ligation reaction |
| 1 | 5 μL Cut plasmid with B0032 + 10 μL PCR Product of R0011 |
Hopefully this will lead to E - X - R0011 - B0032 - S - P in the B0032 plasmid backbone with Amp resistance marker.
 "
"