Team:TU Delft/16 July 2010 content
From 2010.igem.org
(Difference between revisions)
(→Characterization of Anderson RBS sequences) |
|||
| (11 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
=Lab work= | =Lab work= | ||
| - | + | ==Ordered DNA + Solvent Tolerance and Hydrocarbon Sensing== | |
| - | The colony PCR of [https://2010.igem.org/Team:TU_Delft#/blog | + | The colony PCR of [https://2010.igem.org/Team:TU_Delft#page=pages/blog&blog=15_July_2010 yesterday] was put on[[Team:TU_Delft/protocols/agarose_gel|1% agarose gel]] |
| + | |||
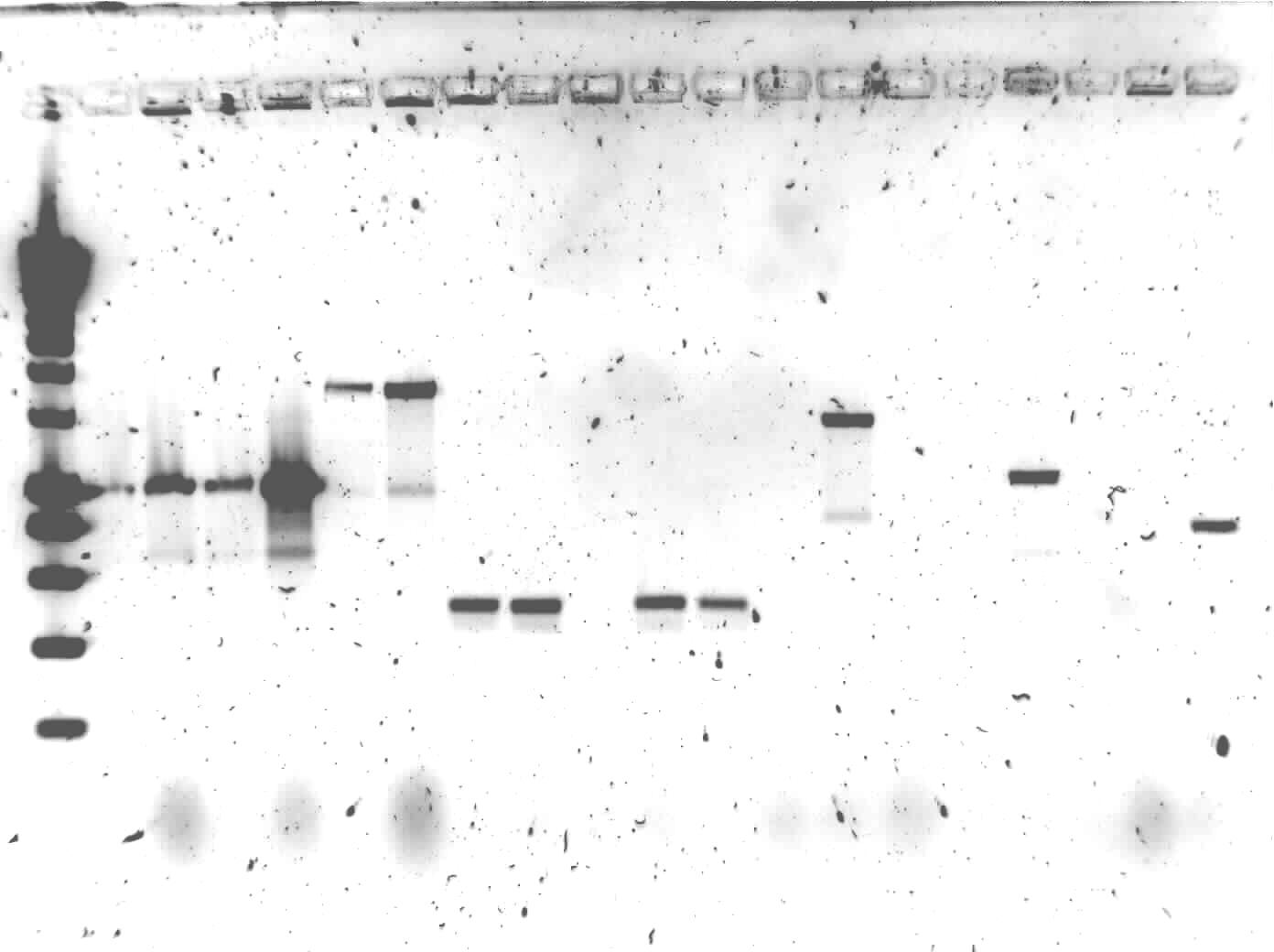
| + | [[Image:TU Delft E 20100716 PCR.jpg|450px|1% agarose of colony PCR. Gel runned at 100V for 1 hour. Of all samples 10 μL + 2 μL loadingbuffer was loaded. 5 μL was loaded of marker]] | ||
| + | |||
| + | Lane description: | ||
| + | {| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | ||
| + | |'''#''' | ||
| + | |'''Description''' | ||
| + | |'''Expected lenght (bp)''' | ||
| + | |'''Primers''' | ||
| + | |'''Status''' | ||
| + | |- | ||
| + | |0 | ||
| + | |SmartLadder | ||
| + | |n/a | ||
| + | |n/a | ||
| + | |n/a | ||
| + | |- | ||
| + | |1 | ||
| + | |transformant #1 of ligation mix AlkS + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |2 | ||
| + | |transformant #2 of ligation mix AlkS + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |3 | ||
| + | |transformant #3 of ligation mix AlkS + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |4 | ||
| + | |transformant #4 of ligation mix AlkS + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |5 | ||
| + | |transformant #1 of ligation mix ladA + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |6 | ||
| + | |transformant #2 of ligation mix ladA + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |7 | ||
| + | |transformant #1 of ligation mix RubA3 + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |8 | ||
| + | |transformant #2 of ligation mix RubA3 + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |9 | ||
| + | |transformant #3 of ligation mix RubA3 + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |10 | ||
| + | |transformant #4 of ligation mix RubA3 + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |11 | ||
| + | |transformant #5 of ligation mix RubA3 + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |12 | ||
| + | |transformant #1 of ligation mix PhPFDbeta + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |13 | ||
| + | |transformant #1 of ligation mix AlnA + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |14 | ||
| + | |transformant #2 of ligation mix AlnA + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |15 | ||
| + | |transformant #1 of ligation mix OprG + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |16 | ||
| + | |transformant #2 of ligation mix OprG + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |17 | ||
| + | |transformant #3 of ligation mix OprG + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |18 | ||
| + | |transformant #1 of ligation mix PhPFDalpha + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |19 | ||
| + | |transformant #2 of ligation mix PhPFDalpha + pSB1C3 | ||
| + | | | ||
| + | |G00100 + G00101 | ||
| + | |<font color=red>✗</font> | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | |||
We harvested the 1 mL bacterial cells of possible correct transformations. The pellets were stored at -20 °C. | We harvested the 1 mL bacterial cells of possible correct transformations. The pellets were stored at -20 °C. | ||
| Line 8: | Line 142: | ||
==Characterization of Anderson RBS sequences== | ==Characterization of Anderson RBS sequences== | ||
| - | + | ====Fluorescence measurements Attempt #2==== | |
| + | |||
Data analysis to follow shortly (good stuff!) | Data analysis to follow shortly (good stuff!) | ||
| - | + | ====Assembly of reference construct & positive control==== | |
| - | The [[Team:TU_Delft/protocols/birnboim_plasmid_isolation Birnboim method ]] was used to isolate J23100 and I13522 (both in pSB1A2) from the o/n 5 mL LB cultures. | + | |
| + | [https://2010.igem.org/Team:TU_Delft#page=pages/blog&blog=15_July_2010 Yesterday's] digestion products of K081005 and I13401 were set for ligation over the weekend at 4 °C: | ||
| + | |||
| + | {| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | ||
| + | |'''#''' | ||
| + | |'''BioBrick''' | ||
| + | |'''Fragment''' | ||
| + | |'''Recipient plasmid''' | ||
| + | |'''Final volume''' | ||
| + | |- | ||
| + | |1 | ||
| + | |[http://partsregistry.org/wiki/index.php?title=Part:BBa_K136011 K136011] | ||
| + | |17.5 μL ‘E-K081005-S’ | ||
| + | |4 μL ‘E-pSB1A2 I13401-X’ | ||
| + | |25 μL | ||
| + | |- | ||
| + | |2 | ||
| + | |[http://partsregistry.org/wiki/index.php?title=Part:BBa_K136011 K136011] | ||
| + | |17.5 μL ‘E-K081005-S’ | ||
| + | |4 μL ‘E-pSB1A2 I13401-X | ||
| + | |25 μL | ||
| + | |- | ||
| + | |3 | ||
| + | |Ligation control | ||
| + | |None | ||
| + | |4 μL ‘E-pSB1A2 I13401-X | ||
| + | |25 μL | ||
| + | |} | ||
| + | |||
| + | To all samples with an end volume of 25 μL, 2.5 μL Ligase buffer was added. | ||
| + | |||
| + | The [[Team:TU_Delft/protocols/birnboim_plasmid_isolation|Birnboim method]] was used to isolate J23100 and I13522 (both in pSB1A2) from the o/n 5 mL LB cultures. We used 3 mL of the bacterial cells to make [[Team:TU_Delft/protocols/freezing_bacterial_stocks|-80 °C stocks]] . The following concentrations of plasmid were obtained: | ||
| + | {| style="color:black; background-color:white;" cellpadding="5" cellspacing="0" border="1" | ||
| + | |'''BioBrick''' | ||
| + | |'''Concentration (ng/μL)''' | ||
| + | |- | ||
| + | |[http://partsregistry.org/wiki/index.php?title=Part:BBa_J23100 J23100] | ||
| + | |130.7 | ||
| + | |- | ||
| + | |[http://partsregistry.org/wiki/index.php?title=Part:BBa_I13522 I13522] | ||
| + | |106.7 | ||
| + | |} | ||
Latest revision as of 20:22, 3 August 2010
Contents |
Lab work
Ordered DNA + Solvent Tolerance and Hydrocarbon Sensing
The colony PCR of yesterday was put on1% agarose gel
Lane description:
| # | Description | Expected lenght (bp) | Primers | Status |
| 0 | SmartLadder | n/a | n/a | n/a |
| 1 | transformant #1 of ligation mix AlkS + pSB1C3 | G00100 + G00101 | ✗ | |
| 2 | transformant #2 of ligation mix AlkS + pSB1C3 | G00100 + G00101 | ✗ | |
| 3 | transformant #3 of ligation mix AlkS + pSB1C3 | G00100 + G00101 | ✗ | |
| 4 | transformant #4 of ligation mix AlkS + pSB1C3 | G00100 + G00101 | ✗ | |
| 5 | transformant #1 of ligation mix ladA + pSB1C3 | G00100 + G00101 | ✗ | |
| 6 | transformant #2 of ligation mix ladA + pSB1C3 | G00100 + G00101 | ✗ | |
| 7 | transformant #1 of ligation mix RubA3 + pSB1C3 | G00100 + G00101 | ✗ | |
| 8 | transformant #2 of ligation mix RubA3 + pSB1C3 | G00100 + G00101 | ✗ | |
| 9 | transformant #3 of ligation mix RubA3 + pSB1C3 | G00100 + G00101 | ✗ | |
| 10 | transformant #4 of ligation mix RubA3 + pSB1C3 | G00100 + G00101 | ✗ | |
| 11 | transformant #5 of ligation mix RubA3 + pSB1C3 | G00100 + G00101 | ✗ | |
| 12 | transformant #1 of ligation mix PhPFDbeta + pSB1C3 | G00100 + G00101 | ✗ | |
| 13 | transformant #1 of ligation mix AlnA + pSB1C3 | G00100 + G00101 | ✗ | |
| 14 | transformant #2 of ligation mix AlnA + pSB1C3 | G00100 + G00101 | ✗ | |
| 15 | transformant #1 of ligation mix OprG + pSB1C3 | G00100 + G00101 | ✗ | |
| 16 | transformant #2 of ligation mix OprG + pSB1C3 | G00100 + G00101 | ✗ | |
| 17 | transformant #3 of ligation mix OprG + pSB1C3 | G00100 + G00101 | ✗ | |
| 18 | transformant #1 of ligation mix PhPFDalpha + pSB1C3 | G00100 + G00101 | ✗ | |
| 19 | transformant #2 of ligation mix PhPFDalpha + pSB1C3 | G00100 + G00101 | ✗ |
We harvested the 1 mL bacterial cells of possible correct transformations. The pellets were stored at -20 °C. We used 3 mL of the bacterial cells to make -80 °C stocks.
Characterization of Anderson RBS sequences
Fluorescence measurements Attempt #2
Data analysis to follow shortly (good stuff!)
Assembly of reference construct & positive control
Yesterday's digestion products of K081005 and I13401 were set for ligation over the weekend at 4 °C:
| # | BioBrick | Fragment | Recipient plasmid | Final volume |
| 1 | K136011 | 17.5 μL ‘E-K081005-S’ | 4 μL ‘E-pSB1A2 I13401-X’ | 25 μL |
| 2 | K136011 | 17.5 μL ‘E-K081005-S’ | 4 μL ‘E-pSB1A2 I13401-X | 25 μL |
| 3 | Ligation control | None | 4 μL ‘E-pSB1A2 I13401-X | 25 μL |
To all samples with an end volume of 25 μL, 2.5 μL Ligase buffer was added.
The Birnboim method was used to isolate J23100 and I13522 (both in pSB1A2) from the o/n 5 mL LB cultures. We used 3 mL of the bacterial cells to make -80 °C stocks . The following concentrations of plasmid were obtained:
| BioBrick | Concentration (ng/μL) |
| J23100 | 130.7 |
| I13522 | 106.7 |
 "
"