Team:Stockholm/Project Idea/Proteins
From 2010.igem.org
(→Proteins) |
|||
| Line 41: | Line 41: | ||
---- | ---- | ||
| - | |||
=== Yeast copper chaperon (yCCS) === | === Yeast copper chaperon (yCCS) === | ||
| Line 80: | Line 79: | ||
---- | ---- | ||
| - | |||
=== Human basic fibroblast growth factor (bFGF) === | === Human basic fibroblast growth factor (bFGF) === | ||
| Line 116: | Line 114: | ||
---- | ---- | ||
| - | |||
=== Protein A, z domain === | === Protein A, z domain === | ||
| - | |||
| + | {|border="1" align="center" cellpadding="3" cellspacing="0" | ||
| + | !colspan="2"|Genepart | ||
| + | |rowspan="10" width="250"|[[Image:|250px]]<br />Primary citation ''et al''. 2010] | ||
| + | |- | ||
| + | |width="130"|'''Length''' | ||
| + | |width="130"|174 bp | ||
| + | |- | ||
| + | |'''Edited nucleotide(s)''' | ||
| + | | - | ||
| + | |- | ||
| + | |'''Removed restr. site(s)''' | ||
| + | | - | ||
| + | |- | ||
| + | |'''GenBank''' | ||
| + | | <br /> | ||
| + | |- | ||
| + | !colspan="2"|Protein | ||
| + | |- | ||
| + | |'''Length''' | ||
| + | |58 aa | ||
| + | |- | ||
| + | |'''Size''' | ||
| + | | | ||
| + | |- | ||
| + | |'''Fasta''' | ||
| + | |ProteinA<br /> | ||
| + | |- | ||
| + | |colspan="2"|First reported by <unknown><br /> | ||
| + | |} | ||
| - | |||
| - | |||
| + | ---- | ||
| + | === IgG protease (IdeS) === | ||
| - | {| | + | {|border="1" align="center" cellpadding="3" cellspacing="0" |
| - | | ''' | + | !colspan="2"|Gene (cDNA) |
| - | + | |rowspan="10" width="250"|[[Image:IdeS.jpg|250px]]<br />Primary citation [http://www.ncbi.nlm.nih.gov/pubmed/15574492 Wenig ''et al''. 2004] | |
|- | |- | ||
| - | | | + | |width="130"|'''Length''' |
| - | | | + | |width="130"|930 bp |
|- | |- | ||
| - | | | + | |'''Edited nucleotide(s)''' |
| - | | - | ||
|- | |- | ||
| - | | | + | |'''Removed restr. site(s)''' |
| - | | - | ||
|- | |- | ||
| - | | ''' | + | |'''GenBank''' |
| - | | | + | | <br /> |
|- | |- | ||
| - | | | + | !colspan="2"|Protein |
| - | + | ||
|- | |- | ||
| - | | | + | |'''Length''' |
| - | | | + | |339 aa |
|- | |- | ||
| - | | Fasta | + | |'''Size''' |
| - | | | + | |37,977 Da |
| + | |- | ||
| + | |'''Fasta''' | ||
| + | |[http://www.ncbi.nlm.nih.gov/protein/209559219?report=fasta IdeS]<br /> | ||
| + | |- | ||
| + | |colspan="2"|First reported by <unknown><br /> | ||
|} | |} | ||
| - | + | ---- | |
| + | ---- | ||
| - | + | == Cell penetrating peptides == | |
| + | This cell-penetrating peptides, (CPPs) may be used in N- and C-terminal fusions with full-length proteins to create transduction proteins with the ability to permeate the lipid bilayer of various cell types, making it a potential gene or protein delivery vector. | ||
| + | === TAT cell penetrating peptide (TAT) === | ||
| + | Purified full-length TAT fusion proteins expressed in ''Escherichia coli'' have been shown to successfully translocate into several human cell types, including all cells found in whole blood, as well as bone marrow stem cells and osteoblasts, while still retaining the fused protein's activity ([http://www.ncbi.nlm.nih.gov/pubmed/9846587 Nagahara ''et al.'' 1998]). The mechanism for transduction over the bilipid membrane is still a matter of debate, but has been suggested to occur through macropinocytosis, a specialized form of endocytosis ([http://www.ncbi.nlm.nih.gov/pubmed/17913584 Gump and Dowdy, 2007]). | ||
| + | TAT is an 11-amino acid derivative from the Human Immunodeficiency Virus 1 (HIV-1) ''trans''-activating transcriptional activator (Tat) ([http://www.ncbi.nlm.nih.gov/pubmed/2849509 Green and Loewenstein, 1988]; [http://www.ncbi.nlm.nih.gov/pubmed/9846587 Nagahara ''et al.'' 1998]). This part was back translated from the corresponding amino acid sequence and optimized for expression in ''Escherichia coli''. Codon usage has been varied for repetitive amino acids to enable DNA synthesis. | ||
| + | {|border="1" align="center" cellpadding="3" cellspacing="0" | ||
| + | !colspan="2"|Sequence | ||
| + | |rowspan="10" width="250"|[[Image:|250px]]<br />Primary citation [ ''et al''. 2001] | ||
| + | |- | ||
| + | |width="130"|'''Length''' | ||
| + | |width="130"|33 bp | ||
| + | |- | ||
| + | |'''GenBank''' | ||
| + | |<br /> | ||
| + | |- | ||
| + | !colspan="2"|Peptide | ||
| + | |- | ||
| + | |'''Length''' | ||
| + | |11 aa | ||
| + | |- | ||
| + | |'''Size''' | ||
| + | | | ||
| + | |- | ||
| + | |colspan="2"|YGRKKRRQRRR | ||
| + | |- | ||
| + | |colspan="2"|First reported by <unknown><br /> | ||
| + | |} | ||
---- | ---- | ||
| + | === Low molecular weight protamine (LMWP) === | ||
| + | Enzymatically prepared LMWP chemically conjugated to ovalbumin (OVA) and bovine serum albumin (BSA) have previously been shown to penetrate the lipid bilayer of human keratinocytes, as well as to successfully permeate mouse skin epidermis ([http://www.ncbi.nlm.nih.gov/pubmed/20232417 Huang ''et al.'', 2010]). Furthermore, LMWP/pDNA complexes can efficiently penetrate into human embryonic kidney cells ([http://www.ncbi.nlm.nih.gov/pubmed/12898639 Park ''et al.'', 2003]). As LMWP has been shown to be neither toxic nor immunogenic ([http://www.ncbi.nlm.nih.gov/pubmed/11741268 Chang ''et al.'' a, 2001]; [http://www.ncbi.nlm.nih.gov/pubmed/11741269 Chang ''et al.'' b, 2001]; [http://www.ncbi.nlm.nih.gov/pubmed/11741270 Lee ''et al.'', 2001]), it may be used as a potential vaccine, drug or gene delivery vector. | ||
| + | LMWP is a 14-amino acid derivative from Rainbow trout (''Oncorhynchus mykiss'') protamine, an arginine-rich protein that replaces histones in chromatin during spermatogenesis ([http://www.ncbi.nlm.nih.gov/pubmed/3755398 McKay ''et al.'', 1986]; [http://www.ncbi.nlm.nih.gov/pubmed/10213181 Byun ''et al.'', 1999]). This part was back translated from the corresponding amino acid sequence and optimized for expression in ''Escherichia coli''. Codon usage has been varied for repetitive amino acids to enable DNA synthesis. | ||
| - | === | + | {|border="1" align="center" cellpadding="3" cellspacing="0" |
| - | [ | + | !colspan="2"|Sequence |
| + | |rowspan="10" width="250"|[[Image:|250px]]<br />Primary citation [ ''et al''. 2001] | ||
| + | |- | ||
| + | |width="130"|'''Length''' | ||
| + | |width="130"|42 bp | ||
| + | |- | ||
| + | |'''GenBank''' | ||
| + | |<br /> | ||
| + | |- | ||
| + | !colspan="2"|Peptide | ||
| + | |- | ||
| + | |'''Length''' | ||
| + | |14 aa | ||
| + | |- | ||
| + | |'''Size''' | ||
| + | | | ||
| + | |- | ||
| + | |colspan="2"|VSRRRRRRGGRRRR | ||
| + | |- | ||
| + | |colspan="2"|First reported by <unknown><br /> | ||
| + | |} | ||
| - | + | ---- | |
| - | + | ||
| + | === Transportan 10 (Tp10) === | ||
| + | Chemically synthesized Tp10 peptides conjugated to different cargo, including pDNA and protein, have been shown to efficiently penetrate the lipid bilayer of both human and mouse cells ([http://www.ncbi.nlm.nih.gov/pubmed/15763630 Kilk ''et al.'', 2005]). Membrane permeation is both energy and temperature independent ([http://www.ncbi.nlm.nih.gov/pubmed/11718666 Hällbrink ''et al.'', 2001]). The exact mechanism for penetration is still unclear ([http://www.ncbi.nlm.nih.gov/pubmed/17218466 Yandek ''et al.'', 2007]). | ||
| + | Tp10 is a 21-amino acid derivative from the parent peptide transportan (originally known as galparan), which is a peptide chimera of the neuropeptide galanin and the wasp venom peptide mastoparan ([http://www.ncbi.nlm.nih.gov/pubmed/10930519 Soomets ''et al.'', 2000]; [http://www.ncbi.nlm.nih.gov/pubmed/8738882 Langel ''et al.'', 1996]). This part was back translated from the corresponding amino acid sequence and optimized for expression in ''Escherichia coli''. Codon usage has been varied for repetitive amino acids to enable DNA synthesis. | ||
| - | + | {|border="1" align="center" cellpadding="3" cellspacing="0" | |
| - | {| | + | !colspan="2"|Sequence |
| - | | ''' | + | |rowspan="10" width="250"|[[Image:|250px]]<br />Primary citation [ ''et al''. 2001] |
| - | + | ||
|- | |- | ||
| - | | | + | |width="130"|'''Length''' |
| - | | | + | |width="130"|63 bp |
|- | |- | ||
| - | | | + | |'''GenBank''' |
| - | | | + | |<br /> |
|- | |- | ||
| - | | | + | !colspan="2"|Peptide |
| - | + | ||
|- | |- | ||
| - | | ''' | + | |'''Length''' |
| - | | | + | |21 aa |
|- | |- | ||
| - | | | + | |'''Size''' |
| - | | | + | | |
|- | |- | ||
| - | | | + | |colspan="2"|AGYLLGKINLKALAALAKKIL |
| - | | | + | |
|- | |- | ||
| - | | | + | |colspan="2"|First reported by <unknown><br /> |
| - | | | + | |
|} | |} | ||
| - | + | ---- | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
{{Stockholm/Footer}} | {{Stockholm/Footer}} | ||
Revision as of 08:21, 27 October 2010
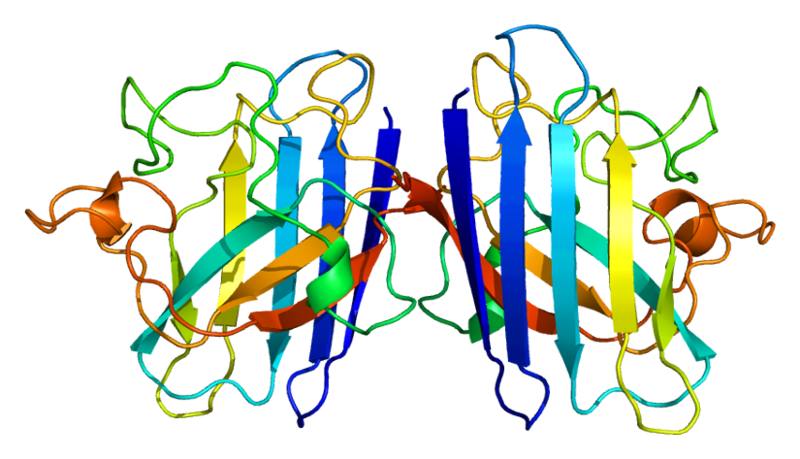
ProteinsSuperoxide dismutase 1 (SOD1)Human soluble Superoxide dismutase 1 (SOD1) is a soluble cytoplasmic protein functional as a homodimer that binds copper and zink ions. SOD1 catalyzes the reaction O-2 + O-2 + 2H+ → H2O2 + O2, protecting the cell from oxidative damage. SOD1 was first cloned and expressed in Escherichia coli by Hallewell et al., (1985).
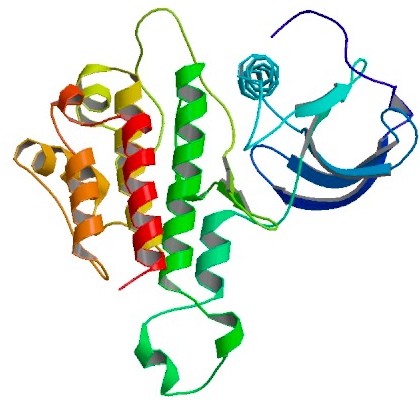
Yeast copper chaperon (yCCS)Yeast copper chaperon protein (yCCS) is a helper chaperon specific for copper/zinc superoxide dismutase located to the cytoplasm. yCCS generates fully metallized, active SOD1 proteins that in turn protects the cell from oxidative damage. yCCS has been shown to successfully mediate the delivery of copper ions to human SOD1 (Ahl et al. 2003). Co-expression of SOD1 and yCCS yields proteins with higher copper contents, leading to increased activity and more stable proteins.
Human basic fibroblast growth factor (bFGF)
Protein A, z domain
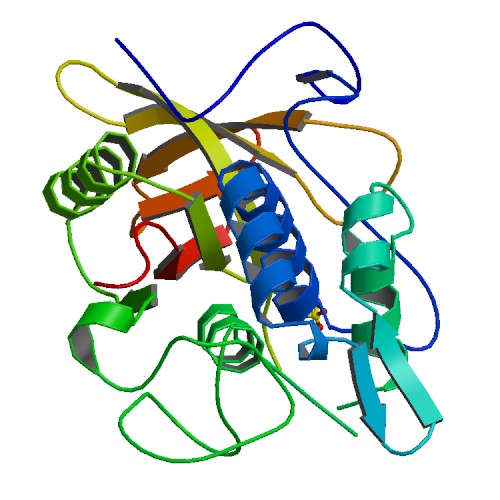
IgG protease (IdeS)
Cell penetrating peptidesThis cell-penetrating peptides, (CPPs) may be used in N- and C-terminal fusions with full-length proteins to create transduction proteins with the ability to permeate the lipid bilayer of various cell types, making it a potential gene or protein delivery vector.
TAT cell penetrating peptide (TAT)Purified full-length TAT fusion proteins expressed in Escherichia coli have been shown to successfully translocate into several human cell types, including all cells found in whole blood, as well as bone marrow stem cells and osteoblasts, while still retaining the fused protein's activity (Nagahara et al. 1998). The mechanism for transduction over the bilipid membrane is still a matter of debate, but has been suggested to occur through macropinocytosis, a specialized form of endocytosis (Gump and Dowdy, 2007). TAT is an 11-amino acid derivative from the Human Immunodeficiency Virus 1 (HIV-1) trans-activating transcriptional activator (Tat) (Green and Loewenstein, 1988; Nagahara et al. 1998). This part was back translated from the corresponding amino acid sequence and optimized for expression in Escherichia coli. Codon usage has been varied for repetitive amino acids to enable DNA synthesis.
Low molecular weight protamine (LMWP)Enzymatically prepared LMWP chemically conjugated to ovalbumin (OVA) and bovine serum albumin (BSA) have previously been shown to penetrate the lipid bilayer of human keratinocytes, as well as to successfully permeate mouse skin epidermis (Huang et al., 2010). Furthermore, LMWP/pDNA complexes can efficiently penetrate into human embryonic kidney cells (Park et al., 2003). As LMWP has been shown to be neither toxic nor immunogenic (Chang et al. a, 2001; Chang et al. b, 2001; Lee et al., 2001), it may be used as a potential vaccine, drug or gene delivery vector. LMWP is a 14-amino acid derivative from Rainbow trout (Oncorhynchus mykiss) protamine, an arginine-rich protein that replaces histones in chromatin during spermatogenesis (McKay et al., 1986; Byun et al., 1999). This part was back translated from the corresponding amino acid sequence and optimized for expression in Escherichia coli. Codon usage has been varied for repetitive amino acids to enable DNA synthesis.
Transportan 10 (Tp10)Chemically synthesized Tp10 peptides conjugated to different cargo, including pDNA and protein, have been shown to efficiently penetrate the lipid bilayer of both human and mouse cells (Kilk et al., 2005). Membrane permeation is both energy and temperature independent (Hällbrink et al., 2001). The exact mechanism for penetration is still unclear (Yandek et al., 2007). Tp10 is a 21-amino acid derivative from the parent peptide transportan (originally known as galparan), which is a peptide chimera of the neuropeptide galanin and the wasp venom peptide mastoparan (Soomets et al., 2000; Langel et al., 1996). This part was back translated from the corresponding amino acid sequence and optimized for expression in Escherichia coli. Codon usage has been varied for repetitive amino acids to enable DNA synthesis.
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"