Team:Slovenia/PROJECT/proof/domain
From 2010.igem.org
</strong></p>
DNA binding domains - which one to choose?
Arrangement of different functional proteins on DNA molecule is required for implementation of our ideas. Because enzymes and other functional polypeptides usually do not have specific affinity for DNA molecule, we decided to fuse them with specific DNA binding domains. These DNA binding domains should have strong affinity for DNA molecule, but interaction should also be sequence specific. Several classes of DNA binding proteins exist in nature, e.g. zinc finger family of proteins, TAL effectors, different transcription factors (TetR, CI, Gal4, ToxR, CinR, RhlR, LuxR) and endo/exonucleases with wrecked cutting activity. Unfortunately, most of them proved to be unusable due to multimerization nature, low affinity and low sequence specificity. Zinc fingers and TAL effectors were on the top of our list since the beginning, because they are modular in nature and can be programed into practically unlimited number of different sequence specific DNA binding domains.
</strong></p>
TAL effectors
TAL family of proteins consists of transcription activator like effectors from plant pathogenic bacteria of Xanthomonas spp. TAL effectors are proteins that are secreted into eukaryotic cells by type III secretion system. In host, they are translocated to the nucleus where they modulate transcription of different genes. TAL protein consists of signal N-terminal, C-terminal domain with transcription activator activity and central region of tandem repeats. It was shown that hipervariable pair of amino acids in every tandem repeat of TAL protein defines its binding properties (Boch et. al., 2009). As such TAL proteins can be programed to bind any desired nucleotide sequence. We designed TAL protein with predicted binding sequence that is the same as Tet operator sequence. With in vivo test of activity we confirmed specific binding of our novel TAL protein to Tet operator sequence. At the end we decided not to choose TAL effectors as DNA binding domains for our project, mainly because designing TAL proteins is still in its early stage.
</strong></p>
Zinc fingers in general
At the end we chose zinc finger proteins as DNA binding domains for our project. Zinc finger proteins form a small, independently folded zinc containing mini domain that recognizes specific nucleic acid sequence. They are present in eukaryotic cells as transcriptional factors and are involved in regulation of endogenous gene expression by specific DNA binding. However, some zinc finger proteins are able to bind to RNA or proteins as well. It is estimated that human genome encodes for more than 700 Cys2His2 zinc finger coding genes, thus being the most common DNA-binding proteins in eukaryotes. Nevertheless, recent biochemical and phylogenetical studies confirmed that they aren't restricted only to eukaryota but are also present in prokaryotes.
</strong></p>
Biochemical and binding properties of zinc fingers
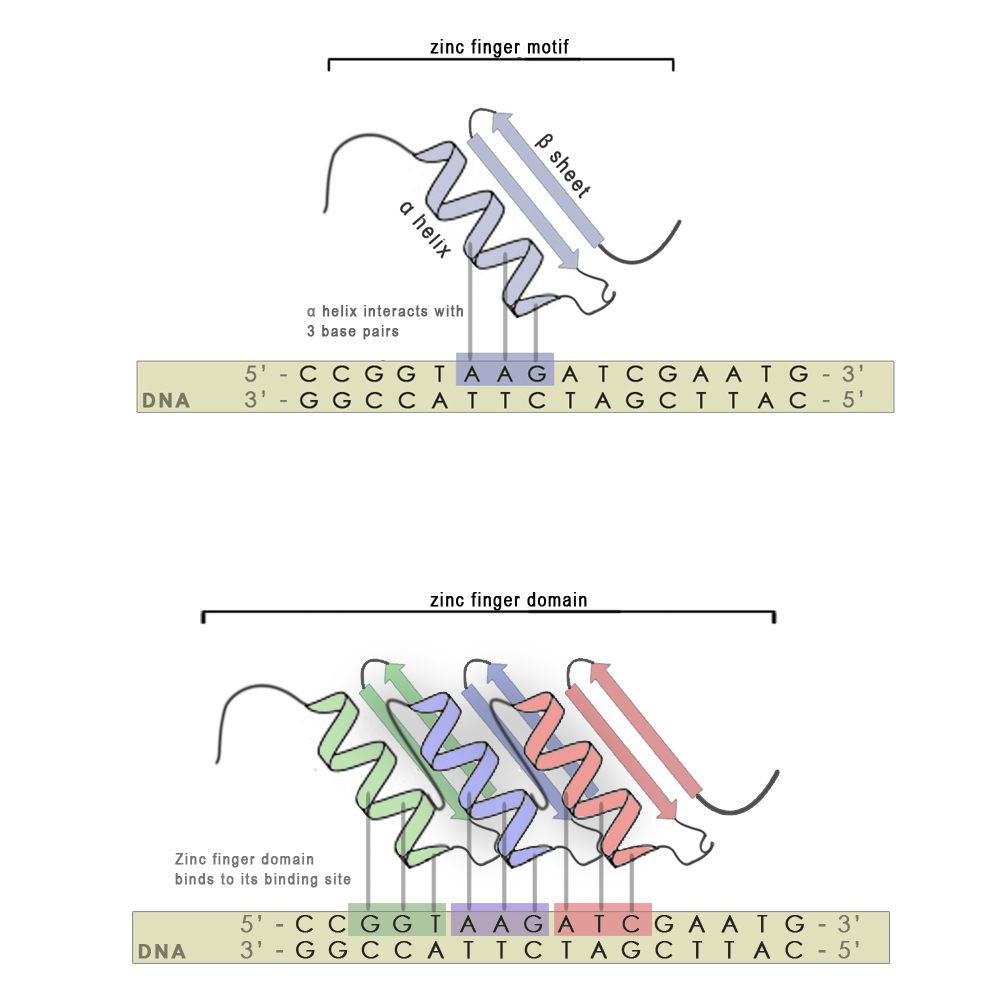
Zinc finger motifs are formed from a short stretch of amino acid residues folded around a zinc cation and have the shape of a finger. Each individual finger is highly conserved in its structure and consists of around 30 amino acid residues. There are two hisitdine residues in every finger on the α-helix and two cysteine residues on the β-sheet that interact with the zinc atom. A short stretch of amino acid residues in the α-helix is responsible for 3 - 4 bp binding specificity. Each finger recognizes and binds to 3 base-pair subsites (see scheme, upper panel), however using modular approach we can stitch several zinc finger mini domains together (scheme, bottom panel) and achieve chemical distinctiveness through variations in certain key amino acid residues. With every new attached zinc finger domain 3-4 base pairs are added to the recognized sequence, thus minimizing non-specific binding on nucleic acids present in host organisms and increasing the specificity of the zinc finger. They also have an advantage over most other DNA binding motifs from natural transcription factors, since the targeted DNA sequence does not have to be symmetric.
 "
"