Team:SDU-Denmark/labnotes
From 2010.igem.org
(Difference between revisions)
(→Exp 1.) |
(→Lab notes (7/12 - 7/18)) |
||
| Line 9: | Line 9: | ||
== Group: Flagella == | == Group: Flagella == | ||
| - | |||
=== Extraction of psb3k3 plasmids with incerted RFP from E. coli MG1655 === | === Extraction of psb3k3 plasmids with incerted RFP from E. coli MG1655 === | ||
| - | |||
==== Exp. 1 ==== | ==== Exp. 1 ==== | ||
| - | + | <br> | |
| - | ''Methods:'' MiniPrep, NanoDrop and gel electrophoresis | + | ''Methods:'' [[https://2010.igem.org/Team:SDU-Denmark/protocols#Plasmid_miniprep_kit_.28Fermentas.29 MiniPrep]], NanoDrop and gel electrophoresis <br><br> |
| - | + | ''Notes:'' We used 9 ml ON culture (Lag phase cells), loaded 2ul sample and 8ul agarose loading dye on a 1,5% gel,used a DNA ladder mix (100-10000 nucleotides) as marker <br><br> | |
| - | ''Notes:'' We used 9 ml ON culture (Lag phase cells), loaded 2ul sample and 8ul agarose loading dye on a 1,5% gel,used a DNA ladder mix (100-10000 nucleotides) as marker | + | ''Results:'' Nanodrop: Tube 1: 30.7 mg/ul Tube 2: 27.4 mg/ul <br><br> |
| - | + | Gel electrophoresis: No result <br><br> | |
| - | ''Results:'' Nanodrop: Tube 1: 30.7 mg/ul Tube 2: 27.4 mg/ul | + | |
| - | + | ||
| - | Gel electrophoresis: No result | + | |
| - | + | ||
--[[User:Louch07|Louch07]] 15:00, 9 July 2010 (UTC) | --[[User:Louch07|Louch07]] 15:00, 9 July 2010 (UTC) | ||
<br><br> | <br><br> | ||
==== Exp. 2 ==== | ==== Exp. 2 ==== | ||
| - | + | <br> | |
| - | ''Methods:'' gel electrophoresis | + | ''Methods:'' gel electrophoresis <br><br> |
| - | + | ''Notes:'' We ran another gel electrophoresis on the miniPrep sample form above. But now we loaded 4ul sample and 4ul agarose loading dye on a 1,5% gel,used a DNA ladder mix (100-10000 nucleotides) as marker. <br><br> | |
| - | ''Notes:'' We ran another gel electrophoresis on the miniPrep sample form above. But now we loaded 4ul sample and 4ul agarose loading dye on a 1,5% gel,used a DNA ladder mix (100-10000 nucleotides) as marker. | + | |
| - | + | ||
''Results:'' Gel electrophoresis: Bands were detected. The psb3k3 plasmid is 2750bp long and the RFP with generator is 1096 bp, which gives a band at about 4000bp compaired with the marker. | ''Results:'' Gel electrophoresis: Bands were detected. The psb3k3 plasmid is 2750bp long and the RFP with generator is 1096 bp, which gives a band at about 4000bp compaired with the marker. | ||
| + | <br><br> | ||
==== Exp. 3 ==== | ==== Exp. 3 ==== | ||
| - | ''Methods:'' MiniPrep, NanoDrop and gel electrophoresis | + | ''Methods:'' [[https://2010.igem.org/Team:SDU-Denmark/protocols#Plasmid_miniprep_kit_.28Fermentas.29 MiniPrep]], NanoDrop and gel electrophoresis. <br><br> |
| - | + | ''Notes:'' We used 10 ml af a culture in Log phase (1ml cells from an ON culture + 9ml LB medium, incubated at 37 degrees celcius for 4 hours), loaded 4ul sample and 4ul agarose loading dye on a 1,5% gel,used a DNA ladder mix (100-10000 nucleotides) as marker. <br><br> | |
| - | ''Notes:'' We used 10 ml af a culture in Log phase (1ml cells from an ON culture + 9ml LB medium, incubated at 37 degrees celcius for 4 hours), loaded 4ul sample and 4ul agarose loading dye on a 1,5% gel,used a DNA ladder mix (100-10000 nucleotides) as marker | + | ''Results:'' Nanodrop: Tube 1: 38.7 mg/ul Tube 2: 32.4 mg/ul. <br><br> |
| - | + | Gel electrophoresis: Bands were detected.<br><br> | |
| - | ''Results:'' Nanodrop: Tube 1: 38.7 mg/ul Tube 2: 32.4 mg/ul | + | |
| - | + | ||
| - | Gel electrophoresis: Bands were detected. | + | |
| - | + | ||
--[[User:Louch07|Louch07]] 17:21, 12 July 2010 (UTC) | --[[User:Louch07|Louch07]] 17:21, 12 July 2010 (UTC) | ||
| - | + | <br><br> | |
=== Polyferation of FlhDC, FlhD and FlhC genes === | === Polyferation of FlhDC, FlhD and FlhC genes === | ||
| - | + | <br><br> | |
| - | ''Methods:'' PCR and Gel electrophoresis | + | ''Methods:'' PCR and Gel electrophoresis.<br><br> |
| - | + | ''Notes:''Since our FldhDC primers have yet to work, we have decided to test them on previously purified cromosomal DNA. Examination of the primers showed that the FlhC reverse primer had a melting temperature of only 45˚C. Therefore we decided to run the samples on a gradient PCR. Simultaneously, we prepared 2 extra samples, isolating FlhD and FlhC, respectively. We did this because we wanted to see if our problems were caused because the combined gene-sequence was to long (932bp).<br> | |
| - | ''Notes:''Since our FldhDC primers have yet to work, we have decided to test them on previously purified cromosomal DNA. Examination of the primers showed that the FlhC reverse primer had a melting temperature of only 45˚C. Therefore we decided to run the samples on a gradient PCR. Simultaneously, we prepared 2 extra samples, isolating FlhD and FlhC, respectively. We did this because we wanted to see if our problems were caused because the combined gene-sequence was to long (932bp). | + | |
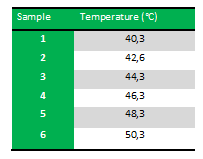
Because we just wanted to test our primers in this PCR, we used Taq polymerase, because although it doesn’t proofread, it is remarkably cheaper than Pfu. On the [http://www.fermentas.com/en/products/all/pcr-qpcr-rt-pcr/standard-pcr/ep028-taq-dna-native Fermentas homepage] we found that the annealing temperature for Taq is Tm-5 , which in this case means 40˚C. However, Taq polymerase is not very effective at temperatures under 50˚C so we designed the gradient to lies between 40 and 55˚C. More specifically we chose the following temperatures: | Because we just wanted to test our primers in this PCR, we used Taq polymerase, because although it doesn’t proofread, it is remarkably cheaper than Pfu. On the [http://www.fermentas.com/en/products/all/pcr-qpcr-rt-pcr/standard-pcr/ep028-taq-dna-native Fermentas homepage] we found that the annealing temperature for Taq is Tm-5 , which in this case means 40˚C. However, Taq polymerase is not very effective at temperatures under 50˚C so we designed the gradient to lies between 40 and 55˚C. More specifically we chose the following temperatures: | ||
| - | + | <br><br> | |
| - | + | ||
[[Image:Team-SDU-Denmark-PCR_temp_for_FlhDC.png]] | [[Image:Team-SDU-Denmark-PCR_temp_for_FlhDC.png]] | ||
| - | + | <br><br> | |
| - | ''Results:'' The experiment was succesfull! We could detect FlhDC DNA at temperatures between 42.6˚C and 48.3˚C. FlhD DNA at temperatures between 40.3˚C and 44.3˚C and also between 48.3˚C and 50.3˚C. FlhC DNA at temperatures run between 40.3˚C and 50.3˚C. | + | ''Results:'' The experiment was succesfull! We could detect FlhDC DNA at temperatures between 42.6˚C and 48.3˚C. FlhD DNA at temperatures between 40.3˚C and 44.3˚C and also between 48.3˚C and 50.3˚C. FlhC DNA at temperatures run between 40.3˚C and 50.3˚C.<br><br> |
| - | + | ||
--[[User:Louch07|Louch07]] 10:13, 12 July 2010 (UTC) | --[[User:Louch07|Louch07]] 10:13, 12 July 2010 (UTC) | ||
| + | <br><br> | ||
== Group: Photosensor == | == Group: Photosensor == | ||
| Line 90: | Line 77: | ||
=== Mini-prep of pSB1A2 w. B0034, pSB1AK3 w. B0015 and pSB3K3 w. J04450(transformation from 08/07) === | === Mini-prep of pSB1A2 w. B0034, pSB1AK3 w. B0015 and pSB3K3 w. J04450(transformation from 08/07) === | ||
Start date: 12/07 End date: 12/07<br> | Start date: 12/07 End date: 12/07<br> | ||
| - | ''Methods:'' | + | ''Methods:'' [[https://2010.igem.org/Team:SDU-Denmark/protocols#Plasmid_miniprep_kit_.28Fermentas.29 MiniPrep]] <br><br> |
''Protocol'': Fermentas protocol <br><br> | ''Protocol'': Fermentas protocol <br><br> | ||
''Notes:'' pellet from 10 mL ON-culture was resuspended in 500uL resuspension buffer, and transferred into two eppendorf tubes, which were run in parellel <br><br> | ''Notes:'' pellet from 10 mL ON-culture was resuspended in 500uL resuspension buffer, and transferred into two eppendorf tubes, which were run in parellel <br><br> | ||
Revision as of 15:34, 12 July 2010
 "
"