Team:Panama/Project
From 2010.igem.org
Nikkibiotech (Talk | contribs) (→The Experiments) |
Ernestopro (Talk | contribs) (→The Experiment) |
||
| (30 intermediate revisions not shown) | |||
| Line 6: | Line 6: | ||
| - | There is considerable interest among bio-industries in bioremediation products such as Rhamnolipids. Rhamnolipids as biosurfactants are important in the remediation of oil spill areas. The cleanup of the Exxon Valdez oil spill with rhamnolipids as biosurfactants was too expensive and complicated, therefore impractical for large-scale bioremediation. However, advances in genetic engineering and synthetic biology offer a viable solution to oil spill pollution | + | There is considerable interest among bio-industries in bioremediation products such as Rhamnolipids. Rhamnolipids as biosurfactants are important in the remediation of oil spill areas. The cleanup of the Exxon Valdez oil spill with rhamnolipids as biosurfactants was too expensive and complicated, therefore impractical for large-scale bioremediation. However, advances in genetic engineering and synthetic biology offer a viable solution to oil spill pollution cleanup. In this project we modify a naturally occurring gene which synthesizes rhamnolipids to make it compatible with Assembly Standard 10 for rhamnosiltransferase 1 complex (rhlAB) gene expression in ''Escherichia coli'' for standardized rhamnolipid production. In this future, we will integrate this and other genetic parts to create an ''E. coli'' based factory for rhamnolipids. Some version of these devices may even include hydrocarbon sensors for Just In Time production of rhamnolipids. |
== Project Details== | == Project Details== | ||
| - | ===The | + | ===The Experiment === |
| - | The purpose behind our project is to take a gene that produces the Rhamnosyltransferase 1 enzyme in a pathogenic | + | The purpose behind our project is to take a gene that produces the Rhamnosyltransferase 1 enzyme in a pathogenic bacteria known as ''Pseudomonas aeruginosa'' and insert it into an efficient and not so pathogenic bacteria that is ''Escherichia coli''. This will help biosynthezise a rhamnolipid that can aid in the more efficent degradation of hydrocarbons. |
| - | + | Upon obtaining and evaluating the sequence of natural Rh1A and Rh1B from GenBank (ID 878955 and 878954) respectively, we realized it contains illegal restriction sites. We therefore needed to fix this gene to be compatible with Assembly Standard 10. We did this by identifying that the PstI restriction enzyme cut our gene sequence in three parts. In the natural rhlA is found an illegal site located at the 821 to 823 base pair corresponding to the codon TGC. In the natural rhlB is found two illegal sites, the first located at the 1843 to 1845 base pair corresponding to the codon CTG and the second illegal site located at the 1957 to 1959 base pair corresponding to the codon CTG. We design an alternative gene sequence using a multiple site mutagenesis and we also designed three different set of primers (we use the Quick Change Primer design tool from Agilent Technologies), because the base pairs that surround the PstI unwanted sequence were different. Nevertheless, before we design the primers we analyzed the RhlAB codon sequence; because we want to perform silent mutations (change the codon but not the amino acid). We follow the manufacturer guidelines for the primer design. | |
| - | In the meantime, we have to obtain our gene from the ''P. aeruginosa''. This bacteria | + | In the meantime, we have to obtain our gene from the ''P. aeruginosa''. This bacteria was a kind donation from Marcelino Gutierrez, Ph.D, who is a researcher from the Drug Discovery department at INDICASAT-AIP. The ''P. aeruginosa'' culture was given to us in liquid media ready for extraction of genomic DNA. Our DNA of interest will be yielded by performing a PCR. |
| - | Once we have this, we have to mutate the Rhamnosyltransferase 1 gene to eliminate the restriction site of | + | Once we have this, we have to mutate the Rhamnosyltransferase 1 gene to eliminate the restriction site of PstI that clashes with the iGEM disposition for design and BioBrick construction that indicates that our gene cannot be cut in any way by the restriction enzymes of the Assembly Standard Protocol 10. This mutation will allow us to have the same gene without the restriction sites within our construct permitted. To resolve this issue we propose to do a multiple site mutagenesis. |
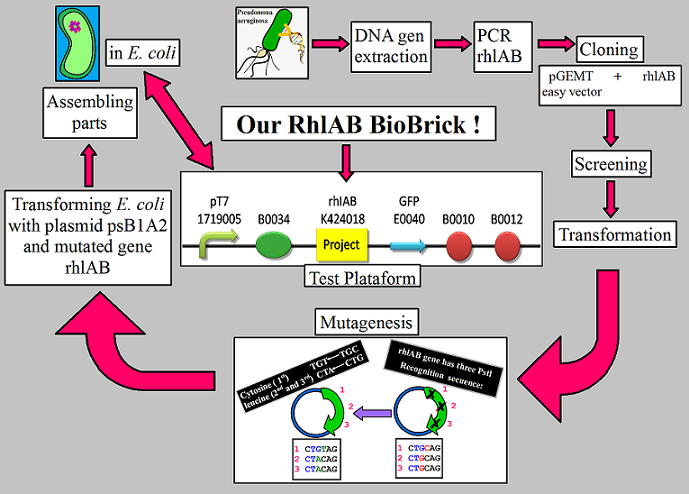
| - | In the end, we'll be able to ligate all the parts together to have one general section that contains the Promoter+RBS+ OUR GENE +Reporter+Terminator | + | In the end, we'll be able to ligate all the parts together to have one general section that contains the Promoter+RBS+ OUR GENE +Reporter+Terminator as a test plataform of our rhamnosyltransferase 1 BioBrick (Rh1AB_BB)expression in ''E. coli'' and EUREKA! we bring a new BioBrick ready to be characterized (see figure 1). |
| + | |||
| + | [[Image:E.Dwiki3.png|center]] | ||
| + | Figure 1: All the steps that we make to get our rhamnosyltransferase BioBrick (Rh1AB_BB). | ||
| + | |||
| + | In the future experiments we will do different tests to ensure that the expected rhamnolipid is being produce. We will create a control platform with: P+RBS+GFP+T and an experimental platform with P+RBS+rh1AB-BB+GFP+T. The control platform will produce no rhamnolipids, and the test platform will produce these rhamnolipids. Also in the future we want to do biochemical and biological characterization of our experimental platform for rhamnolipid production. To measure the quantity of rhamnolipid produced we will do it by performing a biomolecule isolation and purification by chromatographic techniques as a biochemistry characterization. The biological characterization will be performed by a comparative bioassay experiment between ''Pseudomonas aeruginosa'' and our new ''E. coli Rh1AB_BB'' strain. In these bioassays we will compare the growth time and efficiency of rhamnolipid quantity production. We expect that our new ''E. coli Rh1AB_BB'' strain produce more rhamnolipid quantity in less time and in a safer way than the ''P.aeruginosa'' bacteria, our control for these bioassay will be the ''E. coli k12'' strain given by iGEM that is the same strain that we used to test our BioBrick platform but without the RhlAB gene sequence. | ||
=== Part 3: Results === | === Part 3: Results === | ||
| + | |||
| + | We were able to standardize the BioBrick extraction procedures and assembly of the parts once they were ready. | ||
| + | Also we isolate the rhlAB gene from the ''Pseudomonas aeruginosa'' genomic DNA, cloned this gene in a plasmid vector pGEMT Easy Vector for its further used in the mutagenesis protocol. | ||
| + | |||
| + | Once each of the parts were obtained, we were able to ligate them together (promoter + RBS and reporter + terminator) and this was confirmed by performing several gel electrophoresis. We accomplish the construction of test plataform for our coding sequence project, but we have not test the plataform yet for the expression of our rhamnosyltransferase BioBrick (Rh1AB_BB) in ''E. coli.'' | ||
Latest revision as of 04:00, 28 October 2010
Republic of Panama
Situated on the isthmus connecting North and South America, it
is bordered by Costa Rica to the northwest, Colombia to the southeast,
the Caribbean Sea to the north and the Pacific Ocean to the south.
Republic of Panama
Situated on the isthmus connecting North and South America, it is bordered by Costa Rica to the northwest, Colombia to the southeast, the Caribbean Sea to the north and the Pacific Ocean to the south.

iGEM PANAMA
In this picture: Carolina, Yisett, Claudio, Nicole, Lorena,
Grimaldo, Natasha, Laura, Ernesto, Dra. Carmenza, Dr. Rao, Ezequiel
iGEM PANAMA
In this picture: Carolina, Yisett, Claudio, Nicole, Lorena, Grimaldo, Natasha, Laura, Ernesto, Dra. Carmenza, Dr. Rao, Ezequiel

iGEM PANAMA
In this picture: Yisett, Silke, Yaraví, Carlos, Dr. Patrick
Nee.
iGEM PANAMA
In this picture: Yisett, Silke, Yaraví, Carlos, Dr. Patrick Nee.

iGEM PANAMA
In this picture: Leyda, Zeuz, Laura
iGEM PANAMA
In this picture: Leyda, Zeuz, Laura

iGEM PANAMA
Labs.
iGEM PANAMA
Labs.

Contents |
Overall project
Standardization of the Rhamnosiltransferase 1 gene complex (rhlAB) into a Biobrick-friendly for rhamnolipid production in E. coli.
There is considerable interest among bio-industries in bioremediation products such as Rhamnolipids. Rhamnolipids as biosurfactants are important in the remediation of oil spill areas. The cleanup of the Exxon Valdez oil spill with rhamnolipids as biosurfactants was too expensive and complicated, therefore impractical for large-scale bioremediation. However, advances in genetic engineering and synthetic biology offer a viable solution to oil spill pollution cleanup. In this project we modify a naturally occurring gene which synthesizes rhamnolipids to make it compatible with Assembly Standard 10 for rhamnosiltransferase 1 complex (rhlAB) gene expression in Escherichia coli for standardized rhamnolipid production. In this future, we will integrate this and other genetic parts to create an E. coli based factory for rhamnolipids. Some version of these devices may even include hydrocarbon sensors for Just In Time production of rhamnolipids.
Project Details
The Experiment
The purpose behind our project is to take a gene that produces the Rhamnosyltransferase 1 enzyme in a pathogenic bacteria known as Pseudomonas aeruginosa and insert it into an efficient and not so pathogenic bacteria that is Escherichia coli. This will help biosynthezise a rhamnolipid that can aid in the more efficent degradation of hydrocarbons.
Upon obtaining and evaluating the sequence of natural Rh1A and Rh1B from GenBank (ID 878955 and 878954) respectively, we realized it contains illegal restriction sites. We therefore needed to fix this gene to be compatible with Assembly Standard 10. We did this by identifying that the PstI restriction enzyme cut our gene sequence in three parts. In the natural rhlA is found an illegal site located at the 821 to 823 base pair corresponding to the codon TGC. In the natural rhlB is found two illegal sites, the first located at the 1843 to 1845 base pair corresponding to the codon CTG and the second illegal site located at the 1957 to 1959 base pair corresponding to the codon CTG. We design an alternative gene sequence using a multiple site mutagenesis and we also designed three different set of primers (we use the Quick Change Primer design tool from Agilent Technologies), because the base pairs that surround the PstI unwanted sequence were different. Nevertheless, before we design the primers we analyzed the RhlAB codon sequence; because we want to perform silent mutations (change the codon but not the amino acid). We follow the manufacturer guidelines for the primer design.
In the meantime, we have to obtain our gene from the P. aeruginosa. This bacteria was a kind donation from Marcelino Gutierrez, Ph.D, who is a researcher from the Drug Discovery department at INDICASAT-AIP. The P. aeruginosa culture was given to us in liquid media ready for extraction of genomic DNA. Our DNA of interest will be yielded by performing a PCR.
Once we have this, we have to mutate the Rhamnosyltransferase 1 gene to eliminate the restriction site of PstI that clashes with the iGEM disposition for design and BioBrick construction that indicates that our gene cannot be cut in any way by the restriction enzymes of the Assembly Standard Protocol 10. This mutation will allow us to have the same gene without the restriction sites within our construct permitted. To resolve this issue we propose to do a multiple site mutagenesis.
In the end, we'll be able to ligate all the parts together to have one general section that contains the Promoter+RBS+ OUR GENE +Reporter+Terminator as a test plataform of our rhamnosyltransferase 1 BioBrick (Rh1AB_BB)expression in E. coli and EUREKA! we bring a new BioBrick ready to be characterized (see figure 1).
Figure 1: All the steps that we make to get our rhamnosyltransferase BioBrick (Rh1AB_BB).
In the future experiments we will do different tests to ensure that the expected rhamnolipid is being produce. We will create a control platform with: P+RBS+GFP+T and an experimental platform with P+RBS+rh1AB-BB+GFP+T. The control platform will produce no rhamnolipids, and the test platform will produce these rhamnolipids. Also in the future we want to do biochemical and biological characterization of our experimental platform for rhamnolipid production. To measure the quantity of rhamnolipid produced we will do it by performing a biomolecule isolation and purification by chromatographic techniques as a biochemistry characterization. The biological characterization will be performed by a comparative bioassay experiment between Pseudomonas aeruginosa and our new E. coli Rh1AB_BB strain. In these bioassays we will compare the growth time and efficiency of rhamnolipid quantity production. We expect that our new E. coli Rh1AB_BB strain produce more rhamnolipid quantity in less time and in a safer way than the P.aeruginosa bacteria, our control for these bioassay will be the E. coli k12 strain given by iGEM that is the same strain that we used to test our BioBrick platform but without the RhlAB gene sequence.
Part 3: Results
We were able to standardize the BioBrick extraction procedures and assembly of the parts once they were ready. Also we isolate the rhlAB gene from the Pseudomonas aeruginosa genomic DNA, cloned this gene in a plasmid vector pGEMT Easy Vector for its further used in the mutagenesis protocol.
Once each of the parts were obtained, we were able to ligate them together (promoter + RBS and reporter + terminator) and this was confirmed by performing several gel electrophoresis. We accomplish the construction of test plataform for our coding sequence project, but we have not test the plataform yet for the expression of our rhamnosyltransferase BioBrick (Rh1AB_BB) in E. coli.
 "
"