Team:Newcastle/Filamentous Cells
From 2010.igem.org
(Difference between revisions)
RachelBoyd (Talk | contribs) |
RachelBoyd (Talk | contribs) (→Filamentous cells genes list) |
||
| Line 2: | Line 2: | ||
=Filamentous Cells= | =Filamentous Cells= | ||
==Filamentous cells genes list== | ==Filamentous cells genes list== | ||
| - | + | ||
| - | + | *''yneA'' (Transcribed with ''yneB'', ''ynzC'', is an analogue of ''sulA'' in ''E.coli'') * our biobrick is designed to over express this gene r educing cell division possibly by inhibiting FtsZ ring formation or constriction. | |
| - | + | *''dinR'' (Homologue of ''lexA'' in ''E.coli'' transcribed in the opposite direction) | |
| - | + | *''ftsZ'' (Involved in the recruitment of other proteins to the divisisome for cytokinesis, strangely over expression results in disruption of Zring formation as well as reduced expression) | |
| - | + | *''secA'' (Involved in the secretion of extracellular proteins and the insertion of transmembrane proteins) | |
| - | + | * ''recA'' (Involved in SOS response removing the repressor DinR (LexA)) | |
| - | + | *''wpr'' and ''epr'' produce extracellular proteases that cleave the signal peptide/transmembrane domain of YneA | |
| - | + | *''ezr'' produces a protein which sequesters FtsZ monomer by binding its C terminal domain and also inhibits GTP binding; however overexpression does not result in filamentation. | |
| - | + | * ''min C,D ,J and divIVA'' prevent polar cell division . | |
| - | + | *Positive regulators of FtsZ: ''ftsA, zapA, zipA, ftsL and divIC'' | |
| - | + | *Inhibitors of Daughter cell separation: ''lytC,D,E,F and cwlS'' *Chains rather than filaments, ''yneA'' is also reported to increase the time spent in chains well into the stationary phase of bacterial growth. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Revision as of 12:05, 24 June 2010

| |||||||||||||
| |||||||||||||
Contents |
Filamentous Cells
Filamentous cells genes list
- yneA (Transcribed with yneB, ynzC, is an analogue of sulA in E.coli) * our biobrick is designed to over express this gene r educing cell division possibly by inhibiting FtsZ ring formation or constriction.
- dinR (Homologue of lexA in E.coli transcribed in the opposite direction)
- ftsZ (Involved in the recruitment of other proteins to the divisisome for cytokinesis, strangely over expression results in disruption of Zring formation as well as reduced expression)
- secA (Involved in the secretion of extracellular proteins and the insertion of transmembrane proteins)
- recA (Involved in SOS response removing the repressor DinR (LexA))
- wpr and epr produce extracellular proteases that cleave the signal peptide/transmembrane domain of YneA
- ezr produces a protein which sequesters FtsZ monomer by binding its C terminal domain and also inhibits GTP binding; however overexpression does not result in filamentation.
- min C,D ,J and divIVA prevent polar cell division .
- Positive regulators of FtsZ: ftsA, zapA, zipA, ftsL and divIC
- Inhibitors of Daughter cell separation: lytC,D,E,F and cwlS *Chains rather than filaments, yneA is also reported to increase the time spent in chains well into the stationary phase of bacterial growth.
YneA
| SOS response is believe to be a universal bacteria phenomenon first studied in E.coli -LexA, recA |
| In Bacillus subtillis (gram positive) dinR protein is homologous to lexA (Repressor of din-damage inducible genes). din genes include uvrA, uvrB, dinB, dinC dinR and recA. DNA damage inhibits cell division. |
| Wild type Bacillus subtillis |
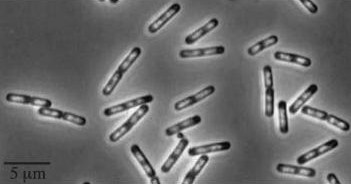
|
| dinR KO |
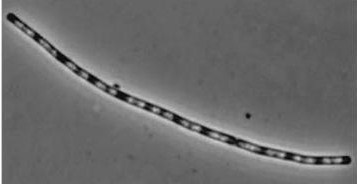
|
| dinR KO mutant over expressed the divergent (opposite direction) transcript for YneA, YneB and YnzC. These genes form the SOS regulon (recA independent SOS response) |
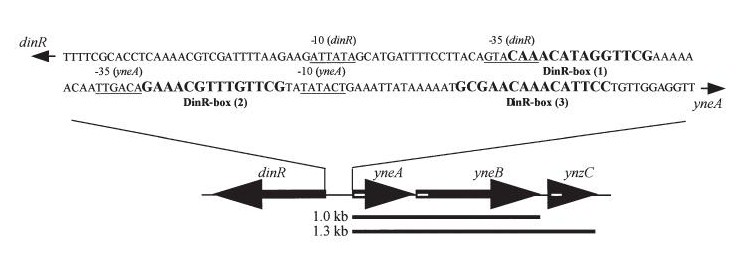
|
| YneA suppressed in wt without SOS induction # |
| Expression of YneA from IPTG controlled promoter in wt leads to elongation. |
| Disruption of YneA in SOS response leads to reduced elongation. Altering YneB and YnzC expression does not affect cell morphology. |
| Double mutant (dinR YneA) |
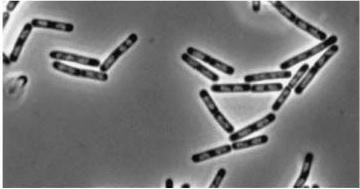
|
| YneA protein required to suppress cell division. Not chromosome replication or segregation. |
| FtsZ is important for bacterial cell division forming a ring structure at the division site by polymerising
assembling other proteins necessary for division at the site. |
| FtsZ localises to the cell division cycle unless dinR is disrupted or YneA is being induced. |
| YneA suppresses FtsZ ring formation- no proven direct interaction by two-hybrid. |
| Filamentous cells less colony formation. |
| YneA expression via the inactivation of dinR by Rec A is important. |
| Sequence of YneA: http://www.ncbi.nlm.nih.gov/nuccore/NC_000964.3?from=1918391&to=1918738&report=graph&content=5 |
Kawai, Y., Moriya, S., & Ogasawara, N. (2003). Identification of a protein, YneA, responsible for cell division suppression during the SOS response in Bacillus subtilis. Molecular microbiology, 47(4), 1113-22. Retrieved from http://www.ncbi.nlm.nih.gov/pubmed/12581363.
MinCDJ
| In B.subtilis cell division occurs precisely mid cell via the formation of the FtsZ ring (tubulin homologue), through Noc (Nucleoid occlusion: prevents division over nucleoids) and the min system (well described in E.coli), which prevents division taking place at the ends of the cell (poles). |
| Main function of Min system to prevent mini cell formation and ensuring only one cell division occurs per cell cycle. |
| Min’s role is not just in the inhibition of FtsZ. FtsZ recruits other components causing synthesis of the new wall and cell invagination. |
|
| Cell division is regulated spatially and temporally. |
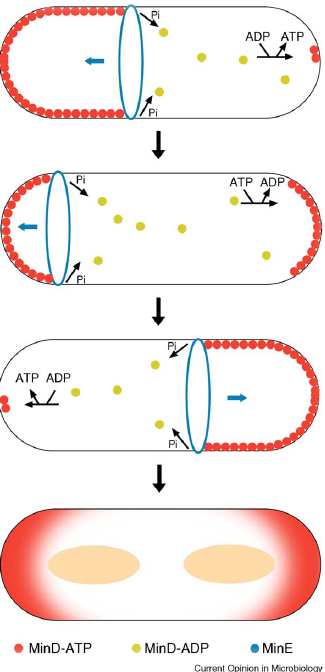
| Min C inhibits FtsZ ring formation; Min C interacts with Min D via its C-terminal domain. Min C inhibits lateral interaction between the filaments. |
| Min D is a membrane associated ATPase. (Min E ensures high concentrations of MinCD at the poles in E.coli, no Min E homologue in B.subtilis! Instead DivIVA acts as a topological factor). |
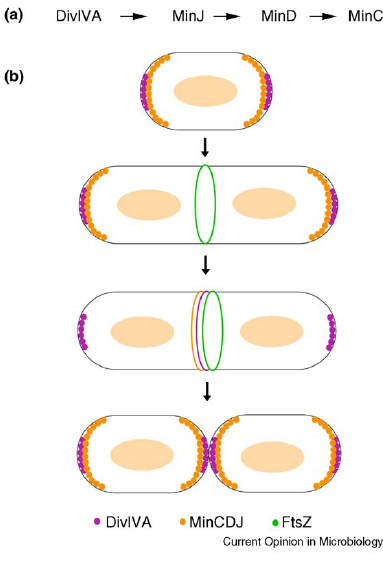
| Min C in B.subtilis is responsible for localisation (shown using GFP) showed a primary site of localisation at the site of active division challenging the original model for the role of Min. |
| Min J was discovered linking Min D to DivIVA and therefore necessary for localisation. |
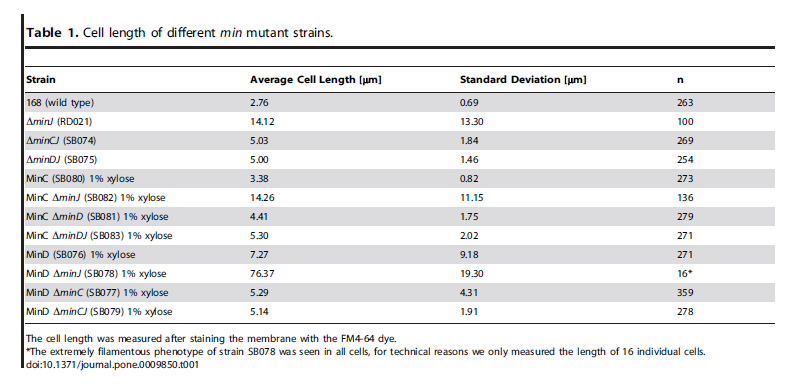
| Over expression of Min D in the absence of Min J causes lethal filamentation. |
|
| Min C sequence |
| http://www.ncbi.nlm.nih.gov/nuccore/NC_000964.3?from=2858550&to=2859298&strand=true&report=graph&content=5 |
| http://www.ncbi.nlm.nih.gov/gene/937500 |
| MinD sequence |
| http://www.ncbi.nlm.nih.gov/nuccore/NC_000964.3?from=2857736&to=2858622&strand=true&report=graph&content=5 |
| http://www.ncbi.nlm.nih.gov/gene/937499 |
| MinJ sequence |
| http://www.ncbi.nlm.nih.gov/nuccore/NC_000964.3?from=3620287&to=3621598&strand=true&report=graph&content=5 |
| http://www.ncbi.nlm.nih.gov/gene/936668 |
| van Baarle, S., & Bramkamp, M. (2010). The MinCDJ system in Bacillus subtilis prevents minicell formation by promoting divisome disassembly. PloS one, 5(3), e9850. doi: 10.1371/journal.pone.0009850. |
| Bramkamp, M. & van Baarle, S., 2009. Division site selection in rod-shaped bacteria. Current opinion in microbiology, 12(6), 683-8. Available at: http://www.ncbi.nlm.nih.gov/pubmed/19884039. |
 "
"