Team:Newcastle/23 August 2010
From 2010.igem.org

| |||||||||||||
| |||||||||||||
Contents |
rocF BioBrick gel electrophoresis
Aims
The aim of this experiment is to run gel electrophoresis for the extracted plasmid pSB1C3 containing rocF BioBrick.
Materials and Protocol
Please refer to: Gel electrophoresis for gel electrophoresis protocol.
Result
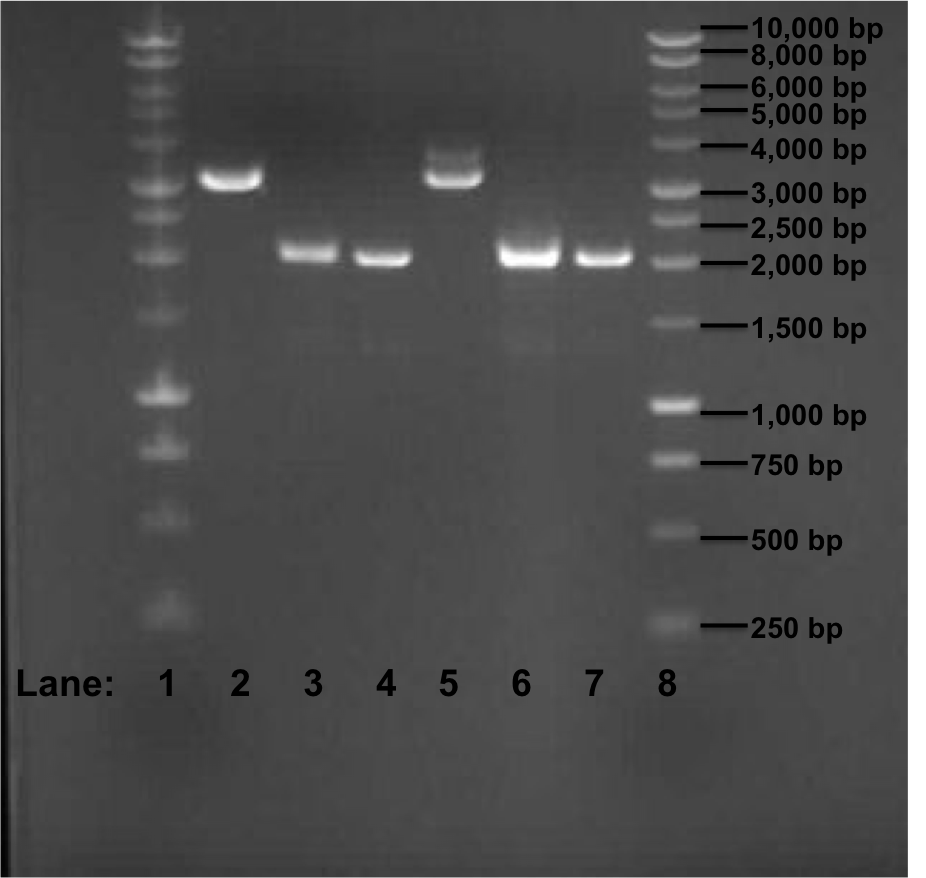
Figure 1: Gel electrophoresis of the miniprep products required for the confirmation of Gibson procedure.
- Lane 1: 1 kb DNA ladder
- Lane 2: pSB1C3 with rocF BioBrick (no. 1)
- Lane 3: pSB1C3 with rocF BioBrick (no. 2)
- Lane 4: pSB1C3 with rocF BioBrick (no. 3)
- Lane 5: pSB1C3 with rocF BioBrick (no. 4)
- Lane 6: pSB1C3 with rocF BioBrick (no. 5)
- Lane 7: pSB1C3 with rocF BioBrick (no. 6)
- Lane 8: 1 kb DNA ladder
Discussion
The expect band size is supposed to be 3195 bp after Gibson procedure ligation. In Figure 1, we can see that in Lane 2 and 5 we got a band of approximately 3200 bp but we found that it is because of the initial insert (rfp gene) which was present in the plasmid pSB1C3 i.e. these 2 sequeich was present in the plasmid pSB1C3 i.e. these 2 sequences are of the template DNA which were used during PCR reaction few days ago. The other 4 bands are of approximately 2100 bp in size.
Conclusion
Overall the Gibson protocol has failed. We did not receive bands of correct size thus showing that the rfp fragments and the plasmid pSB1C3 did not ligate at all. The 2 bands of approximately 3200 bp size are of the plasmid pSB1C3 containing rfp gene insert and thus these fragments are of template DNA for the PCR reaction which had taken place few days ago. The explanation for the failure of the Gibson protocol is as following:
- Because of the presence of the BioBrick prefix and suffix sequences at the ends of the fragments, the fragments circularized and ligated with themselves. This is because both prefix and suffix contains Not1 restriction site and Xba1 and Spe1 restriction site which have similar sequences and thus during ligation step of the Gibson reaction, the fragments containing prefix and suffix religate with themselves and thus the fragments have not been able to ligate with each other as expected earlier.
 
|
 "
"