Team:Newcastle/13 July 2010
From 2010.igem.org
PhilipHall (Talk | contribs) (→Results) |
Shethharsh08 (Talk | contribs) |
||
| Line 29: | Line 29: | ||
*'''Lane 9''': 1 Kbp Molecular weight marker | *'''Lane 9''': 1 Kbp Molecular weight marker | ||
| - | + | ==Discussion== | |
This procedure was followed in order to determine whether our ''lacI'' had been ligated into the pSB1AT3 in the way we desired (i.e. in front of RFP). | This procedure was followed in order to determine whether our ''lacI'' had been ligated into the pSB1AT3 in the way we desired (i.e. in front of RFP). | ||
| - | The gel shows bands for colonies 1 and 2 that | + | The gel shows bands for colonies 1 and 2, that indicates that they contain vector only (size 3-3.5 Kbp) and for 3-7 that indicate that they contain vector plus RFP (size 4.5 Kbp), as with the original plasmid stock. |
==Conclusion== | ==Conclusion== | ||
The conclusion that can be drawn from this gel is that the lacI has not been inserted into the plasmid as we desired. However due to the feint second band we are unsure as to whether plasmid has fully digested, to that end we will leave the digest running overnight and run the samples on a gel tomorrow. | The conclusion that can be drawn from this gel is that the lacI has not been inserted into the plasmid as we desired. However due to the feint second band we are unsure as to whether plasmid has fully digested, to that end we will leave the digest running overnight and run the samples on a gel tomorrow. | ||
{{Team:Newcastle/footer}} | {{Team:Newcastle/footer}} | ||
Revision as of 08:17, 11 August 2010

| |||||||||||||
| |||||||||||||
Contents |
LacI BioBrick Construction
Aims
- To use PCR to extract lacI (promoter, ribosome-binding site (RBS) & coding sequence (CDS)) from plasmid pMutin4 and ligate into vector pSB1AT3 in front of red fluorescent protein (RFP).
Materials
- The seven overnight cultures
- Restriction enzyme EcoR1
Protocol
- E. coli DH5α was transformed again with our ligation mix.
- The seven overnight cultures underwent plasmid extraction.
- The seven plasmid extracts were then restriction digested with EcoR1 to linearise.
- The seven digests were then run on a gel.
Results
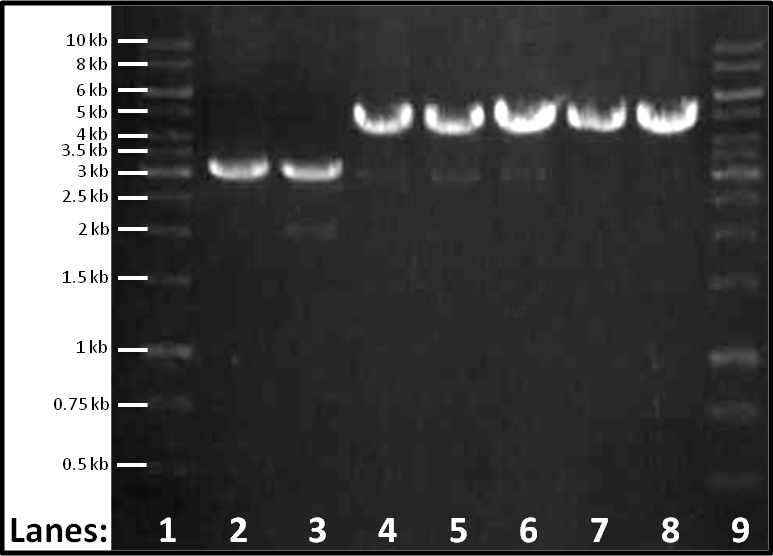
Figure 1: Gel electrophoresis of the seven plasmid extracts which have been digested with EcoR1.
- Lane 1: 1 Kbp Molecular weight marker
- Lane 2-8: Colonies 1-7 in order
- Lane 9: 1 Kbp Molecular weight marker
Discussion
This procedure was followed in order to determine whether our lacI had been ligated into the pSB1AT3 in the way we desired (i.e. in front of RFP). The gel shows bands for colonies 1 and 2, that indicates that they contain vector only (size 3-3.5 Kbp) and for 3-7 that indicate that they contain vector plus RFP (size 4.5 Kbp), as with the original plasmid stock.
Conclusion
The conclusion that can be drawn from this gel is that the lacI has not been inserted into the plasmid as we desired. However due to the feint second band we are unsure as to whether plasmid has fully digested, to that end we will leave the digest running overnight and run the samples on a gel tomorrow.
 
|
 "
"