Team:Lethbridge/Notebook/Lab Work/June
From 2010.igem.org
JustinVigar (Talk | contribs) (→June 30/2010) |
Adam.smith4 (Talk | contribs) (→June 28/2010) |
||
| (20 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | + | <div style="background-color:#000000; color:white"> | |
| + | <html> | ||
| - | + | <br> | |
| - | + | <table border="0" width="100%" style="background-color:#000000"> | |
| - | + | <tr> | |
| - | + | <th> | |
| - | + | <image src="https://static.igem.org/mediawiki/2010/2/29/UofLteamlogo.jpg" width="200px"/> | |
| - | + | ||
| - | + | </th> | |
| - | + | ||
| - | + | ||
| - | + | <th> | |
| - | + | ||
| - | + | ||
| - | + | <image src="https://static.igem.org/mediawiki/2010/9/91/UofLLabWork.JPG" height="300px"/> | |
| - | + | ||
| - | + | ||
| - | + | </th> | |
| - | + | ||
| - | + | ||
| - | + | <th> | |
| - | + | ||
| - | + | ||
| - | + | <image src="https://static.igem.org/mediawiki/2010/2/29/UofLteamlogo.jpg" width="200px"/> | |
| - | + | ||
| - | + | ||
| - | + | </th> | |
| - | + | ||
| - | + | ||
| - | + | </tr> | |
| - | + | ||
| - | + | ||
| - | + | </table> | |
| + | |||
| + | |||
| + | |||
| + | <br> | ||
| + | |||
| + | <align="centre"> | ||
| + | <table border="0" width="100%" style="background-color:#000000"> | ||
| + | |||
| + | <tr> | ||
| + | |||
| + | <th> | ||
| + | |||
| + | <a href="https://2010.igem.org/Team:Lethbridge"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/2/22/UofLHome.jpg" width="80"/> | ||
| + | </a> | ||
| + | |||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Team"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/0/0d/UofLTeam.jpg" width="80"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Project"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/8/8d/UofLProjectbutton.jpg" width="80"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Lab_Work"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/7/73/UofLNotebookbutton.jpg" width="80"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Parts"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/8/84/UofLPartsSubmittedToTheRegistrybutton.jpg" width="80"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Modeling"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/e/e1/UofLModelingbutton.jpg" width="80"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Ethics"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/2/26/UofLEthicsbutton.jpg" width="80"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Safety"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/0/00/UofLSafetybutton.jpg" width="80"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Art"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/0/0a/UofLArt.jpg" width="80"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/News"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/c/c3/UofLNewsButton.jpg" width="80"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | </table> | ||
| + | </body> | ||
| + | </html> | ||
| + | <hr> | ||
| + | |||
| + | <html> | ||
| + | <center> | ||
| + | <font color="white">Feel free to look around our notebook! | ||
| + | </center> | ||
| + | </html> | ||
| + | |||
| + | <html> | ||
| + | <body> | ||
| + | <center> | ||
| + | <table border="0" width="28%" style="background-color:#000000"> | ||
| + | |||
| + | <tr> | ||
| + | <th> | ||
| + | <div class="miniBar"> | ||
| + | <div class="countdown"><object type="application/x-shockwave-flash" data="http://www.oneplusyou.com/bb/files/countdown/countdown.swf?co=FFFFFF&bgcolor=000000&date_month=10&date_day=27&date_year=0&un=THE WIKI FREEZE&size=normal&mo=10&da=27&yr=2010" width="300" height="100"><param name="movie" value="http://www.oneplusyou.com/bb/files/countdown/countdown.swf?co=FFFFFF&bgcolor=000000&date_month=10&date_day=27&date_year=0&un=THE WIKI FREEZE&size=normal&mo=10&da=27&yr=2010" /><param name="bgcolor" value="#000000" /></object><img src="http://www.oneplusyou.com/q/img/bb_badges/countdown.jpg" alt="" style="display: none;" height="1" width="1" /></div> | ||
| + | <div class="miniContainer"> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Lab_Work"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/7/73/UofLNotebookbutton.jpg" width="80"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Protocols"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/9/91/UofLprotocolsbutton.jpg" width="60"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Calendar"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/7/73/UofLcalendar.jpg" width="60"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th> | ||
| + | <div class="miniBar"> | ||
| + | <div class="countdown"><object type="application/x-shockwave-flash" data="http://www.oneplusyou.com/bb/files/countdown/countdown.swf?co=FFFFFF&bgcolor=000000&date_month=11&date_day=05&date_year=0&un=THE IGEM JAMBOREE&size=normal&mo=11&da=05&yr=2010" width="300" height="100"><param name="movie" value="http://www.oneplusyou.com/bb/files/countdown/countdown.swf?co=FFFFFF&bgcolor=000000&date_month=11&date_day=05&date_year=0&un=THE IGEM JAMBOREE&size=normal&mo=11&da=05&yr=2010" /><param name="bgcolor" value="#000000" /></object><img src="http://www.oneplusyou.com/q/img/bb_badges/countdown.jpg" alt="" style="display: none;" height="1" width="1" /></div> | ||
| + | <div class="miniContainer"> | ||
| + | </th> | ||
| + | <tr> | ||
| + | </table> | ||
| + | </center> | ||
| + | </body> | ||
| + | </html> | ||
| + | |||
| + | <hr> | ||
| + | |||
| + | <html> | ||
| + | <center> | ||
| + | <font color="white">Here you can check out the work we have done in the lab, click on a month to take a look! | ||
| + | </center> | ||
| + | </html> | ||
| + | |||
| + | <html> | ||
| + | <body> | ||
| + | <center> | ||
| + | <table border="0" width="50%" style="background-color:#000000"> | ||
| + | |||
| + | <tr> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Lab_Work/April"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/8/8a/UofLapril.jpg" width="60"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Lab_Work/May"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/7/7b/UofLmaybutton.jpg" width="60"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Lab_Work/June"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/8/80/UofLjunebutton.jpg" width="80"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Lab_Work/July"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/5/53/UofLjulybutton.jpg" width="60"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Lab_Work/August"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/1/15/UofLaugustbutton.jpg" width="60"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Lab_Work/September"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/4/4d/UofLseptemberbutton.jpg" width="60"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | <th><a href="https://2010.igem.org/Team:Lethbridge/Notebook/Lab_Work/October"> | ||
| + | <img src="https://static.igem.org/mediawiki/2010/4/4e/UofLoctoberbutton.jpg" width="60"/> | ||
| + | </a> | ||
| + | </th> | ||
| + | |||
| + | |||
| + | |||
| + | <tr> | ||
| + | </table> | ||
| + | </center> | ||
| + | </body> | ||
| + | </html> | ||
| + | <hr> | ||
| - | + | <BLOCKQUOTE> | |
| - | + | ||
| - | + | ||
| - | =June 2010= | + | =<font color="white">June 2010= |
| - | ==June 1/2010== | + | ==<font color="white">June 1/2010== |
JV quantified the amount of DNA in gels run to date using ImageJ software. Results to be posted in [[Team:Lethbridge/Notebook/Working_Plasmids|working plasmids box]].<br> | JV quantified the amount of DNA in gels run to date using ImageJ software. Results to be posted in [[Team:Lethbridge/Notebook/Working_Plasmids|working plasmids box]].<br> | ||
| Line 68: | Line 218: | ||
*BglII Endonuclease (Bba_K112106)<br> | *BglII Endonuclease (Bba_K112106)<br> | ||
| - | ==June 2/2010== | + | ==<font color="white">June 2/2010== |
(In Lab: JV)<br> | (In Lab: JV)<br> | ||
| Line 141: | Line 291: | ||
Killed enzymes by incubating reactions for 10 minutes at 80<sup>o</sup>C</ul> | Killed enzymes by incubating reactions for 10 minutes at 80<sup>o</sup>C</ul> | ||
| - | ==June 2/2010 - Evening== | + | ==<font color="white">June 2/2010 - Evening== |
<b>Objective:</b> Set up new ligations of pLacI and sRBS-Lum-dT according to Tom Knight's protocol. Previous ligation had very little DNA.<br> | <b>Objective:</b> Set up new ligations of pLacI and sRBS-Lum-dT according to Tom Knight's protocol. Previous ligation had very little DNA.<br> | ||
<b>Relevant Information:</b><br> | <b>Relevant Information:</b><br> | ||
| Line 190: | Line 340: | ||
Incubate for 20 minutes at 80<sup>o</sup>C to heat kill<br> | Incubate for 20 minutes at 80<sup>o</sup>C to heat kill<br> | ||
| - | ==June 3/2010== | + | ==<font color="white">June 3/2010== |
Carried out protocol described in June 2/2010 - Evening<br> | Carried out protocol described in June 2/2010 - Evening<br> | ||
Analyzed results on 1% agarose gel.Load order as follows:<br> | Analyzed results on 1% agarose gel.Load order as follows:<br> | ||
| Line 220: | Line 370: | ||
| - | ==June 3/2010 - Evening== | + | ==<font color="white">June 3/2010 - Evening== |
<b>Objective:</b> Repeat restriction of pSB1T3 and ligate with pLacI and sRBS-Lum-dT. Previous ligations all used up on gel.<br> | <b>Objective:</b> Repeat restriction of pSB1T3 and ligate with pLacI and sRBS-Lum-dT. Previous ligations all used up on gel.<br> | ||
<b>Method:</b><br> | <b>Method:</b><br> | ||
| Line 265: | Line 415: | ||
<b>Continue Ligation on Saturday (See below).</b><br> | <b>Continue Ligation on Saturday (See below).</b><br> | ||
| - | ==June 5/2010 == | + | ==<font color="white">June 5/2010 == |
(In the lab:AS)<br> | (In the lab:AS)<br> | ||
<b>Objective:</b> Ligate restriction products from June 3/2010.<br> | <b>Objective:</b> Ligate restriction products from June 3/2010.<br> | ||
| Line 289: | Line 439: | ||
| - | ==June 6/2010 == | + | ==<font color="white">June 6/2010 == |
(In Lab: JV, HS)<br> | (In Lab: JV, HS)<br> | ||
| Line 321: | Line 471: | ||
None of the plates showed any growth. | None of the plates showed any growth. | ||
| - | ==June 7/2010 == | + | ==<font color="white">June 7/2010 == |
(In Lab: JV, HB, TF, AV)<br> | (In Lab: JV, HB, TF, AV)<br> | ||
| Line 390: | Line 540: | ||
<font color ="Red">IMAGE TO COME!!!!!!</font><br> | <font color ="Red">IMAGE TO COME!!!!!!</font><br> | ||
| - | ==June 7/2010 - Evening == | + | ==<font color="white">June 7/2010 - Evening == |
(In Lab: AS, KG)<br> | (In Lab: AS, KG)<br> | ||
| Line 428: | Line 578: | ||
<font color ="Red">FILL ME IN!!!!!!</font><br> | <font color ="Red">FILL ME IN!!!!!!</font><br> | ||
| - | ==June 8/2010== | + | ==<font color="white">June 8/2010== |
(In the lab: JV, AV)<br> | (In the lab: JV, AV)<br> | ||
<b>Objective:</b> Follow the overexpression of our pLacI-sRBS-Lum-dT construct.<br> | <b>Objective:</b> Follow the overexpression of our pLacI-sRBS-Lum-dT construct.<br> | ||
| Line 542: | Line 692: | ||
Looks like the mms6 DNA was not cut at all. Therefore is doubtful that the ligations will work.<br><br> | Looks like the mms6 DNA was not cut at all. Therefore is doubtful that the ligations will work.<br><br> | ||
| - | ==June 8/2010 - Evening== | + | ==<font color="white">June 8/2010 - Evening== |
<b>Objective:</b> Transform pLacI-sRBS-Lum-dT constructs assembled via three antibiotic assembly on June 3/2010. Also transform mms6-dT constructs assembled today using old insertion method.<br> | <b>Objective:</b> Transform pLacI-sRBS-Lum-dT constructs assembled via three antibiotic assembly on June 3/2010. Also transform mms6-dT constructs assembled today using old insertion method.<br> | ||
<b>Method:</b> Follow [[Team:Lethbridge/Notebook/Protocols|transformation protocol]]. | <b>Method:</b> Follow [[Team:Lethbridge/Notebook/Protocols|transformation protocol]]. | ||
<b>Results:</b> No colonies anywhere<br> | <b>Results:</b> No colonies anywhere<br> | ||
| - | ==June 9/2010== | + | ==<font color="white">June 9/2010== |
(in the lab: TF, JV)<br> | (in the lab: TF, JV)<br> | ||
<b>Objective:</b> Transform mms6-dT ligation reactions from June 8/2010.<br> | <b>Objective:</b> Transform mms6-dT ligation reactions from June 8/2010.<br> | ||
| Line 557: | Line 707: | ||
<b>Results:</b> No transformants on plates.<br><br> | <b>Results:</b> No transformants on plates.<br><br> | ||
| - | ==June 10/2010== | + | ==<font color="white">June 10/2010== |
(In the lab: AV, HB, JV)<br> | (In the lab: AV, HB, JV)<br> | ||
<b>Objective:</b> Repeat ligation of mms6 (B9,D9,D10) and dT (C1).<br> | <b>Objective:</b> Repeat ligation of mms6 (B9,D9,D10) and dT (C1).<br> | ||
| Line 612: | Line 762: | ||
*B0010-B0012: 120 colonies | *B0010-B0012: 120 colonies | ||
| - | ==June 14/2010 Evening== | + | ==<font color="white">June 14/2010 Evening== |
(In the lab: TF, DM, HS)<br> | (In the lab: TF, DM, HS)<br> | ||
<b>Objective:</b> Restriction Digest of RBS-xylE with EcoRI and SpeI<br> | <b>Objective:</b> Restriction Digest of RBS-xylE with EcoRI and SpeI<br> | ||
| Line 636: | Line 786: | ||
*Incubate overnight | *Incubate overnight | ||
| - | ==June 15/2010== | + | ==<font color="white">June 15/2010== |
(In the lab: JV)<br> | (In the lab: JV)<br> | ||
| Line 696: | Line 846: | ||
DNA was then purified using the protocol for purification of PCR products (BioBasic). DNA was eluted with 35µL of elution buffer.<br> | DNA was then purified using the protocol for purification of PCR products (BioBasic). DNA was eluted with 35µL of elution buffer.<br> | ||
| - | ==June 15/2010 Evening== | + | ==<font color="white">June 15/2010 Evening== |
(In the lab: AS)<br> | (In the lab: AS)<br> | ||
| Line 737: | Line 887: | ||
<b>Results: </b>Everything cuts exactly as it should. Continue with ligation and analysis. <font color ="Red">IMAGE TO COME!!!!</font><br> | <b>Results: </b>Everything cuts exactly as it should. Continue with ligation and analysis. <font color ="Red">IMAGE TO COME!!!!</font><br> | ||
| - | ==June 16/2010== | + | ==<font color="white">June 16/2010== |
(in lab: JV) | (in lab: JV) | ||
<b>Objective: </b>To miniprep Adam's overnight cultures of xylE and rbs. Completing this will give us a working stock of this plasmid.<br> | <b>Objective: </b>To miniprep Adam's overnight cultures of xylE and rbs. Completing this will give us a working stock of this plasmid.<br> | ||
| Line 754: | Line 904: | ||
<b>Results:</b> There wasn't any growth after overnight incubation. Possibility of an antibiotic mix-up. | <b>Results:</b> There wasn't any growth after overnight incubation. Possibility of an antibiotic mix-up. | ||
| - | ==June 16/2010 Evening== | + | ==<font color="white">June 16/2010 Evening== |
(in lab: ADS) | (in lab: ADS) | ||
| Line 821: | Line 971: | ||
60µL of each VF2 & VR primers (10µL) -> send 65µL | 60µL of each VF2 & VR primers (10µL) -> send 65µL | ||
| - | ==June 17/2010== | + | ==<font color="white">June 17/2010== |
(in lab: JV, HB) | (in lab: JV, HB) | ||
<b>Objective:</b>Run Agarose gel of overnight samples from June 16/10 to test T4 DNA ligase<br> | <b>Objective:</b>Run Agarose gel of overnight samples from June 16/10 to test T4 DNA ligase<br> | ||
| Line 843: | Line 993: | ||
<font color ="Red">IMAGE TO COME!!!!</font><br> | <font color ="Red">IMAGE TO COME!!!!</font><br> | ||
| - | ==June 17/2010 Evening== | + | ==<font color="white">June 17/2010 Evening== |
(in lab: JS, AS, KG)<br> | (in lab: JS, AS, KG)<br> | ||
| Line 868: | Line 1,018: | ||
*controls (no forward or reverse primer and no DNA)<br> | *controls (no forward or reverse primer and no DNA)<br> | ||
| - | ==June 18/2010== | + | ==<font color="white">June 18/2010== |
(in lab: JV)<br> | (in lab: JV)<br> | ||
| Line 889: | Line 1,039: | ||
<br> | <br> | ||
| - | ==June 21/2010== | + | ==<font color="white">June 21/2010== |
(in lab: JV)<br> | (in lab: JV)<br> | ||
| Line 915: | Line 1,065: | ||
<b>Results:</b>PCR samples were run on a gel (1% 1XTAE) with the PCR samples from June 22/10. This gel was run on June 22/10<br> | <b>Results:</b>PCR samples were run on a gel (1% 1XTAE) with the PCR samples from June 22/10. This gel was run on June 22/10<br> | ||
| - | ==June 22/2010== | + | ==<font color="white">June 22/2010== |
(in lab: JV, HS)<br> | (in lab: JV, HS)<br> | ||
| Line 983: | Line 1,133: | ||
Ran the gel for 40 minutes at 100V.<br> | Ran the gel for 40 minutes at 100V.<br> | ||
| - | ==June 22/2010 Evening== | + | ==<font color="white">June 22/2010 Evening== |
(in lab: KG, AS)<br> | (in lab: KG, AS)<br> | ||
| Line 1,040: | Line 1,190: | ||
<tr><td>Imitrages polymerase</td><td>0.2</td></tr></table><br> | <tr><td>Imitrages polymerase</td><td>0.2</td></tr></table><br> | ||
| - | ==June 23/2010== | + | ==<font color="white">June 23/2010== |
(in lab: JV, HS)<br> | (in lab: JV, HS)<br> | ||
| Line 1,098: | Line 1,248: | ||
*all Phusion tubes are <font color ="Blue">blue</font><br> | *all Phusion tubes are <font color ="Blue">blue</font><br> | ||
| - | <b>Results:</b> | + | <b>Results:</b> |
| + | <font color="red">IMAGE!!!</font> | ||
| Line 1,135: | Line 1,286: | ||
<font color ="Red">Picture to come!!!!!!!!!</font><br> | <font color ="Red">Picture to come!!!!!!!!!</font><br> | ||
| - | ==June 24/2010== | + | ==<font color="white">June 24/2010== |
(in lab: JV, HS, AV)<br> | (in lab: JV, HS, AV)<br> | ||
| Line 1,237: | Line 1,388: | ||
<tr><td>33</td><td>1kb ladder</td><td>0.5 ladder + 2 Dye (6X) + 9.5 Milli-Q H<sub>2</sub>O</td></tr> | <tr><td>33</td><td>1kb ladder</td><td>0.5 ladder + 2 Dye (6X) + 9.5 Milli-Q H<sub>2</sub>O</td></tr> | ||
</table><br> | </table><br> | ||
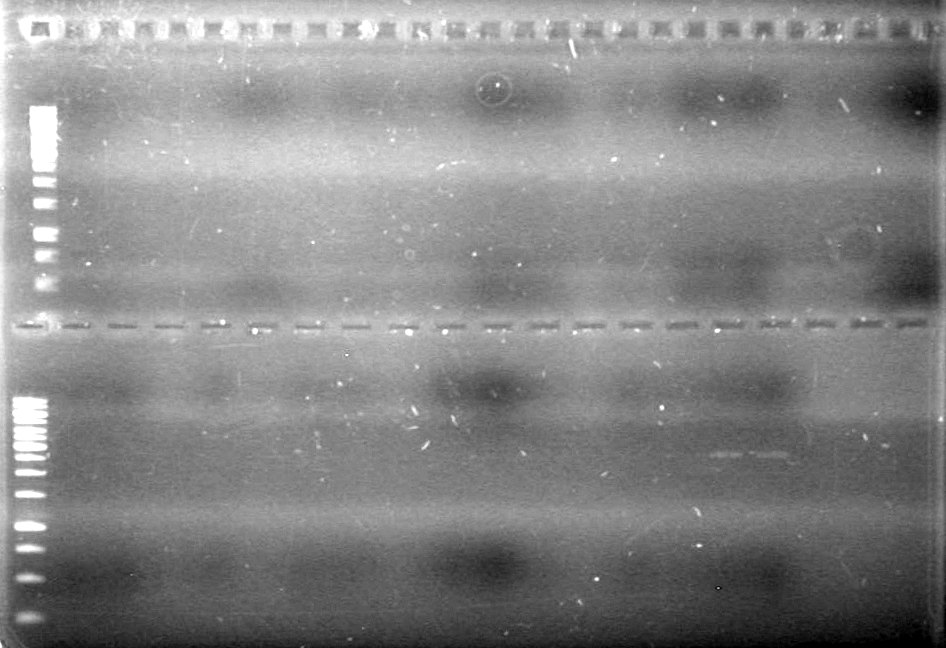
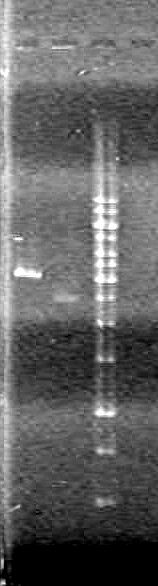
| + | <b>Results:</b> [[image:Lethbridge_100624PCR.JPG|200px]] | ||
| - | <font color =" | + | ==<font color="white">June 28/2010== |
| - | + | ||
| - | + | ||
(in lab: JV, HS, AV, HB)<br> | (in lab: JV, HS, AV, HB)<br> | ||
| Line 1,298: | Line 1,448: | ||
<tr><td>18</td><td>pET28(a) RD</td><td>5 DNA + 2 Dye (6X) + 5 Milli-Q H<sub>2</sub>O</td></tr> | <tr><td>18</td><td>pET28(a) RD</td><td>5 DNA + 2 Dye (6X) + 5 Milli-Q H<sub>2</sub>O</td></tr> | ||
<tr><td>19</td><td>pET28(a) RD</td><td>5 DNA + 2 Dye (6X) + 5 Milli-Q H<sub>2</sub>O</td></tr> | <tr><td>19</td><td>pET28(a) RD</td><td>5 DNA + 2 Dye (6X) + 5 Milli-Q H<sub>2</sub>O</td></tr> | ||
| - | <tr><td>20</td><td>pET28(a)</td><td>5 DNA + 2 Dye (6X) + 5 Milli-Q H<sub>2</sub>O</td></tr> | + | <tr><td>20</td><td>pET28(a)</td><td>5 DNA + 2 Dye (6X) + 5 Milli-Q H<sub>2</sub>O</td></tr></table> |
Ran gel at 100V for 90 minutes<br> | Ran gel at 100V for 90 minutes<br> | ||
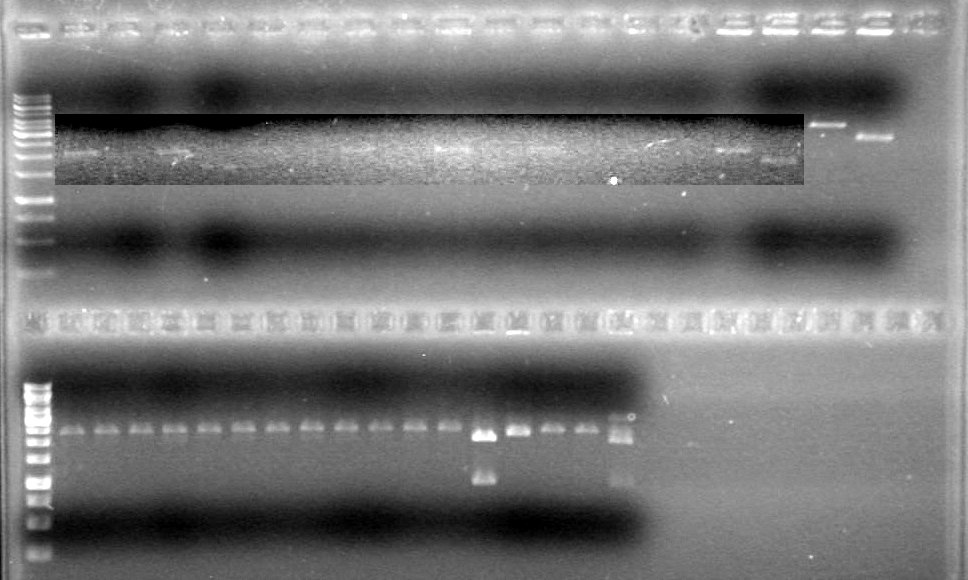
<b>Results:</b>pET28(a) bands are all smeared, and are not useful from inserting parts.<br> | <b>Results:</b>pET28(a) bands are all smeared, and are not useful from inserting parts.<br> | ||
| + | [[image:Lethbridge_100628AV (2).JPG|200px]]<br> | ||
| - | <font color =" | + | ==<font color="white">June 30/2010== |
| + | (in lab: JV)<br> | ||
| + | |||
| + | <b>Objective:</b>: Isolate plasmid DNA from DH5α<br> | ||
| + | |||
| + | <b>Method:</b>Used Kothe Maxiprep protocol.<br> | ||
| + | Cell Pellet Weights:<br> | ||
| + | *xylE = 1.15g | ||
| + | *Mms6 = 1.13g | ||
| + | *pET28(a) = 0.67g | ||
| + | *lumazine synthase = 1.1g<br> | ||
| + | |||
| + | <b>Results:</b><br> | ||
 "
"