Team:Imperial College London/Modelling/Protein Display
From 2010.igem.org
m |
|||
| Line 61: | Line 61: | ||
<div class="accordionButton">Objectives</div> | <div class="accordionButton">Objectives</div> | ||
<div class="accordionContent"> | <div class="accordionContent"> | ||
| - | The motivation to | + | The motivation to develop this model came from our design process. The idea of having surface proteins that could be cleaved and then activate the receptor was very innovatory. However, with this new approach questions arose that could not be answered without using computer models. Hence, the following aims for this model were specified: |
<ol> | <ol> | ||
<li>Determine the concentration of Schistosoma elastase or TEV protease that should be added to the bacteria in order to trigger a response. This should allow us to correlate the required concentration for the activation with the concentration of Schistosoma elastase in the water.</li> | <li>Determine the concentration of Schistosoma elastase or TEV protease that should be added to the bacteria in order to trigger a response. This should allow us to correlate the required concentration for the activation with the concentration of Schistosoma elastase in the water.</li> | ||
| Line 84: | Line 84: | ||
<h2>1. Elements of the system</h2> | <h2>1. Elements of the system</h2> | ||
<ol> | <ol> | ||
| - | <li>The surface protein | + | <li>The surface protein consists of a cell wall binding domain, linker, AIP (Auto Inducing Peptide). It is expressed by constitutive gene expression. It was assumed that the bacteria would be fully grown before carrying out the detection so the cell wall would be covered with as many surface proteins as the cell can maintain.</li> |
| - | <li>Schistosoma elastase (enzyme released by the parasite) cleaves AIP from cell wall binding domain at the linker site. In laboratory we used TEV protease as we could not get handle of Schistosoma elastase, so the model was adjusted appropriately (TEV enzyme kinetic parameters were used).</li> | + | <li>Schistosoma elastase (enzyme released by the parasite) cleaves the AIP from the cell wall binding domain at the linker site. In the laboratory we used TEV protease as we could not get handle of Schistosoma elastase, so the model was adjusted appropriately (TEV enzyme kinetic parameters were used).</li> |
<li>ComD receptor is being activated by high enough AIP concentration. Once activated, ComD signals into the cell to activate the colour response.</li> | <li>ComD receptor is being activated by high enough AIP concentration. Once activated, ComD signals into the cell to activate the colour response.</li> | ||
</ol> | </ol> | ||
Revision as of 10:09, 15 October 2010
| Surface Protein Model | ||||||||||||||||||||||||||
|
Objectives
The motivation to develop this model came from our design process. The idea of having surface proteins that could be cleaved and then activate the receptor was very innovatory. However, with this new approach questions arose that could not be answered without using computer models. Hence, the following aims for this model were specified:
Detailed Description
This model consists of 5 parts that had to be developed:
1. Elements of the system
2. Interactions between elementsApart from proteins being expressed from genes, there was only one chemical reaction identified in this part of the system. It is cleavage of protein which is an enzymatic reaction:
This results in 4 partial differential equations (PDEs), which is of similar form as the 1-step amplification model. However, most of the constants and initial concentrations are different. For detailed description and derivation of PDEs, please refer to "Detailed Description" part of Modelling Output. 3. Threshold concentration of AIP (20/08/2010)The optimal peptide concentration required to activate ComD is 10 ng/ml [1]. This is the threshold value for ComD activation. However, the minimum concentration of peptide to give a detectable activation is 0.5ng/ml.The threshold for minimal activation of receptor is cth=4.4658x10-9 mol/L. Click on the button below to uncover the calculations. Converting 10 ng/ml to 4.4658x10-9 mol/L
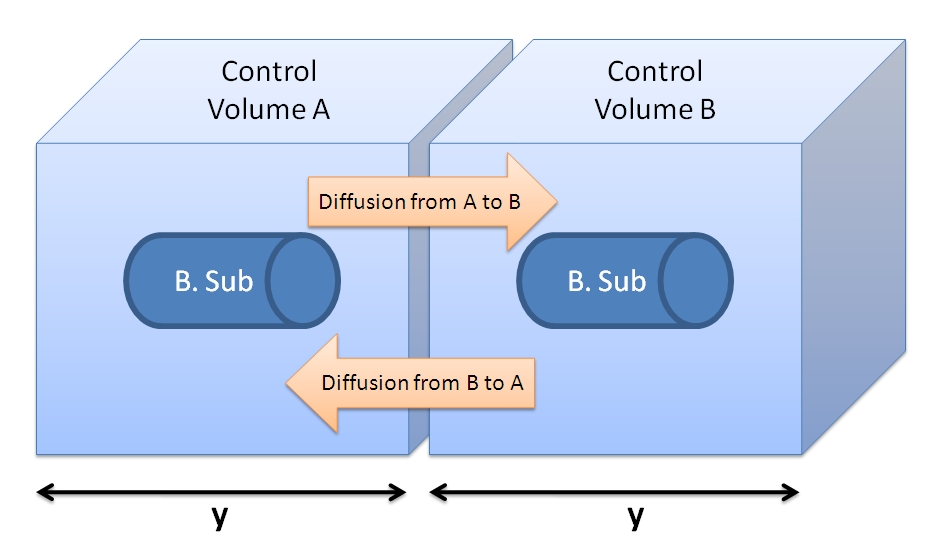
4. Control volume selectionNote that this enzymatic reaction is to be modelled outside the cell. Hence, it is important to consider the cell separations. It is worth considering whether the diffusion or fluid movements will play significant role. Initially we defined control volume as if bacteria were growing closely in colonies on the plate. The details can be revealed by clicking on the button below.Initial Choice of Control Volume
Control volume initial choice (23/08/2010)
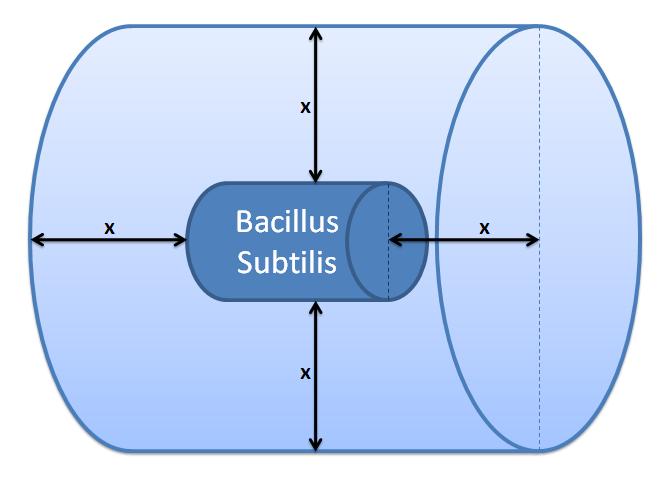
This control volume is considered to be wrong by us, but the details were kept for the reference. The control volume: The inner boundary is determined by the bacterial cell (proteins after being displayed and cleaved cannot diffuse back into bacterium). The outer boundary is more time scale dependent. We have assumed that after mass cleavage of the display-proteins by TEV, many of these AIPs will bind to the receptors quite quickly (eg. 8 seconds). Our volume is determined by the distance that AIPs could travel outwards by diffusion within that short time. In this way, we are sure that the concentration of AIPs outside our control volume after a given time is approximately 0. This approach is not very accurate and can lead us to false negative conclusions (as in reality there will be a concentration gradient, with highest conentration on the cell wall).
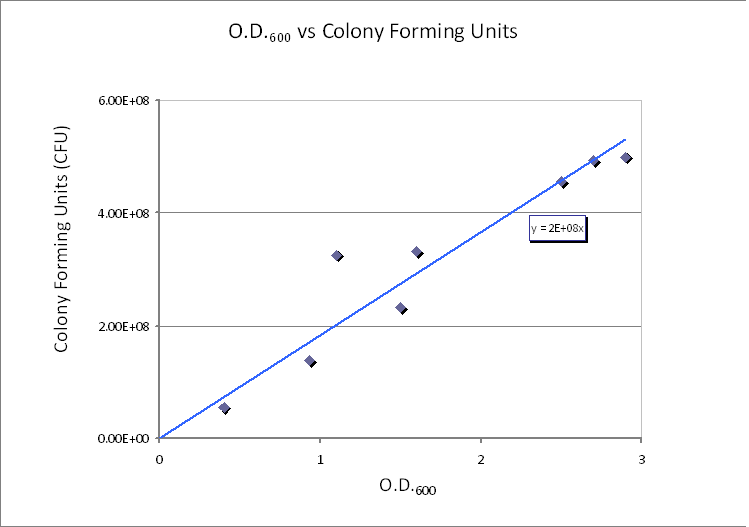
We realized that our initial choice of control volume was not accurate because this assumption was treating bacteria as the medium. However, in reality bacteria live in colonies very close to each other. As a product our bacteria was meant to be used in suspension, so we had to reconsider the issue. Using CFU to estimate the spacing between cells (24/08/2010) CFU stands for Colony-forming unit. It is a measure of bacterial numbers. For liquids, CFU is measured per ml. Since we already have data of CFU/ml from the Imperial iGEM 2008, this is an easy way to estimate the number of cells in a given volume using a spectrometer at 600nm wavelength. The graph below is taken from the Imperial iGEM 2008 Wiki page [4].
Side length of CV = y = 1.26x10-4 dm = 1.26x10-5 m. Choice of Control Volume allows simplifications (24/08/2010)
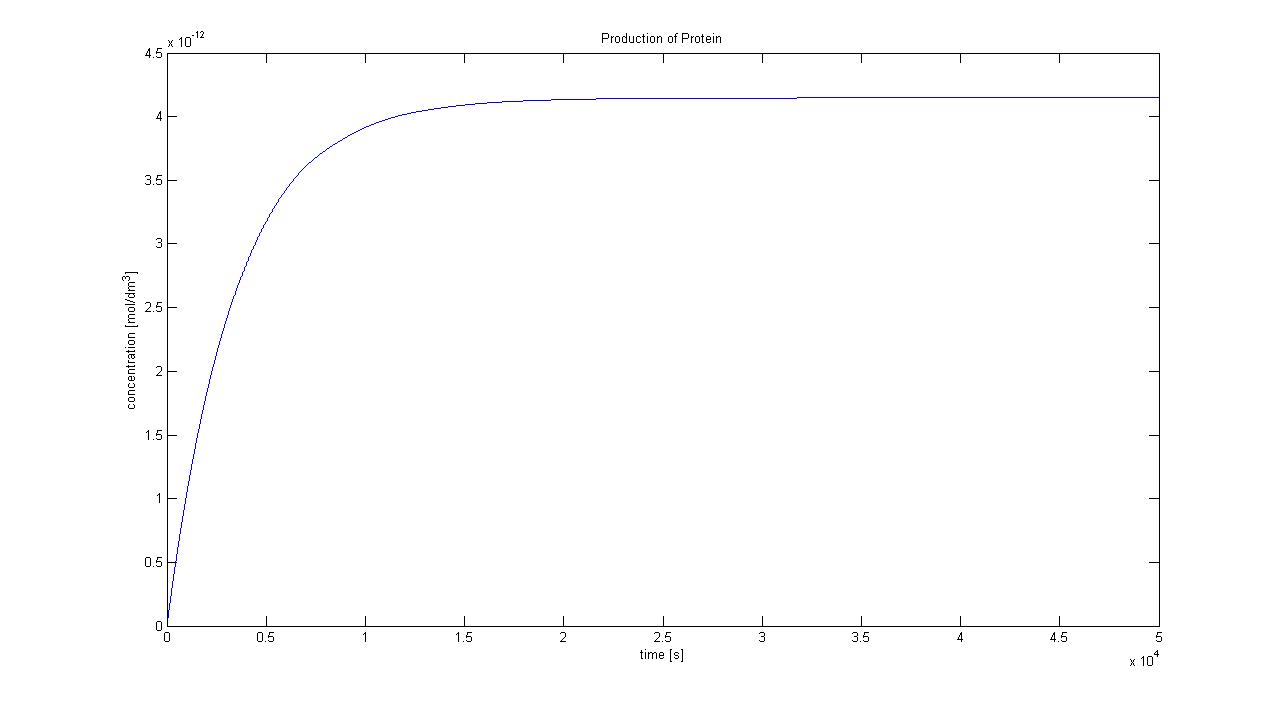
5. Protein production (23/08/2010)
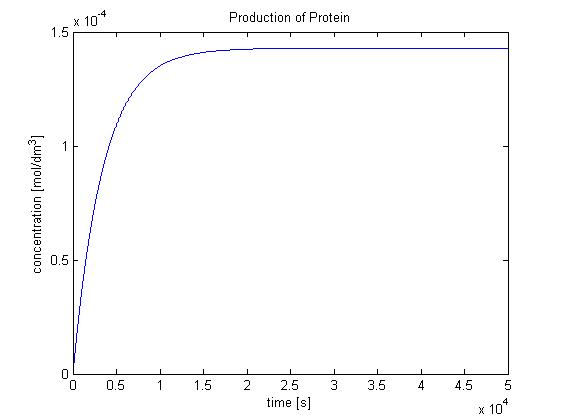
Hence, we can deduce that the concentration that the protein expression will tend to: c = 1.4289x10-4 mol/dm3 = cfinal. Therefore, we can model the protein production by transcription and translation and adjust the production constant so the concentration value will tend towards cfinal. Using a similar model to the simple production of Dioxygenase for the Output Amplification Model (Model preA), we obtain the following graph:
The degradation rate was kept constant, and the production rate was changed according to the final concentration. Protein production in Control Volume (23/08/2010)The previously determined constants of protein production in B.sub to obtain the concentration of proteins are not valid in the Control Volume. It has to be adjusted (multiplied) by the following factor: factor=Vbacillus/VCV = 5.7974x10-6 (for the particular numbers presented above)
References
Parameters & Constants
Constants for the Protein Display Model
References
Results & Conclusions
Risk of False positives (31/08/2010) It was pointed out that we should assess the risk of false positive activation of the receptor. We are particularly concerned about the display protein not binding to the cell wall, but instead diffusing into the extra-cellular environment. In order to be able to assess the risk of false positives, we need to do further research into the affinity of AIP with attached linker and transmembrane proteins for the receptor as compared to the affinity of the AIP itself for the receptor. This paper [1] might have some information on affinity comparison. We need to know how proteins are being transported from intracellular to transmembrane space. Understanding this concept could give us an idea of what could go wrong. References
Download: MatLab Files
|
 "
"




![Graphs showing the simulation using [TEV]<sub>0</sub> = 4x10<sup>-4</sup> mol/dm<sup>3</sup>. The graph on the right hand-side below shows that the AIP threshold (red line) is reached after 22 s.](http://www.openwetware.org/images/7/77/Protein_display.png)
