Team:Imperial College London/Lab Diaries/XylE team
From 2010.igem.org
| Line 123: | Line 123: | ||
{| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
|style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"| | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"| | ||
| + | |- | ||
| | | | ||
| Line 158: | Line 159: | ||
*the His-GFP-Flag DNA was gel purified | *the His-GFP-Flag DNA was gel purified | ||
*transformation of XL-1Blue cells with J23101 in J62001 vector from the registry. One more attemp to construct a successful promoter-xylE ligation, since we believe that the strand annealed promoter was of bad quality | *transformation of XL-1Blue cells with J23101 in J62001 vector from the registry. One more attemp to construct a successful promoter-xylE ligation, since we believe that the strand annealed promoter was of bad quality | ||
| + | |} | ||
| + | |||
| + | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
| + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"| | ||
| + | |- | ||
| + | | | ||
=='''Week 8'''== | =='''Week 8'''== | ||
| Line 203: | Line 210: | ||
* Gel analysis of the attempted annealing reaction of GFP-2 XylE-2 showed unsufficiently clear bands for gel-purification. A new reaction is being prepared: 10 rounds of annealing PCR, followed by addition of primers (5' primer for GFP-2 and 3' primer for XylE-2) in order to introduce an amplification step in the reaction. --- Following Kirstins advice, we are discarding this reaction and wait for the arrival of new primers for XylE-2 (5' + TEV) and GFP-2 (3' +TEV). | * Gel analysis of the attempted annealing reaction of GFP-2 XylE-2 showed unsufficiently clear bands for gel-purification. A new reaction is being prepared: 10 rounds of annealing PCR, followed by addition of primers (5' primer for GFP-2 and 3' primer for XylE-2) in order to introduce an amplification step in the reaction. --- Following Kirstins advice, we are discarding this reaction and wait for the arrival of new primers for XylE-2 (5' + TEV) and GFP-2 (3' +TEV). | ||
| + | |||
| + | |} | ||
{| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
|style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"| | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"| | ||
| + | |- | ||
| | | | ||
| - | |||
=='''Week 9'''== | =='''Week 9'''== | ||
| Line 235: | Line 244: | ||
*The new GFP + TEV primer arrived, was diluted and used to set up the appropriate PCR. | *The new GFP + TEV primer arrived, was diluted and used to set up the appropriate PCR. | ||
[[Image:Assay 3 sept.jpg|thumb|center|400px|Catechol assay on XylE-trasformed cells in a 96-well plate (A to H decreasing cell concentration, 1-10 decreasing catechol concentration, column 11 and 12 negative and control)]] | [[Image:Assay 3 sept.jpg|thumb|center|400px|Catechol assay on XylE-trasformed cells in a 96-well plate (A to H decreasing cell concentration, 1-10 decreasing catechol concentration, column 11 and 12 negative and control)]] | ||
| + | |} | ||
Revision as of 09:38, 21 October 2010
| Lab Diaries | Overview | Surface Protein Team | XylE Team | Vectors Team | Modelling Team |
| Here are the technical diaries for our project. We've split them up into three lab teams and the modelling team. We think it's really important that absolutely anyone can find out what we've been doing. For a really detailed look at what we did, and when, you've come to the right place! | |
| Team XylE |
Follow our progress: Click me!
| XylE team Lab Objectives |
|
| Lab notes and schedule |
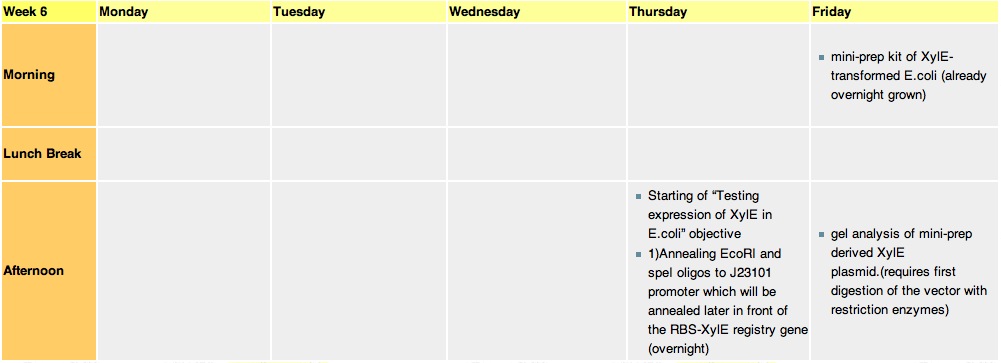
Week 6Thursday, 12-Aug-2010
we constructed the standard E.coli promoter J23101 with sticky ends. These ends are complementary to restriction sites made by EcoRI and SpeI enzyme. This promoter will be later used in 3A assemply to construct a promoter-RBS-XylE design in a psB1C3 vector. E.coli will be transformed with this final construct plasmid to assess XylE activity and characterization. It will also be one of the submitted biobricks.
these cultures are going to be used tomorrow for mini-prepping. Miniprep will allow us to isolate E.coli's plasmid DNA(which contains the XylE gene). Friday, 13-Aug-2010
Mini-prep is usually used to confirm that our gene of interest has not been changed in any way, as the isolated plasnid id sent for sequencing. However, since XylE was taken from the registry, we assume that it is fine and no sequencing is required. The mini-prep will later be used for the midi-prep (that gives out higher yeilds of DNA needed for cloning).
|
|
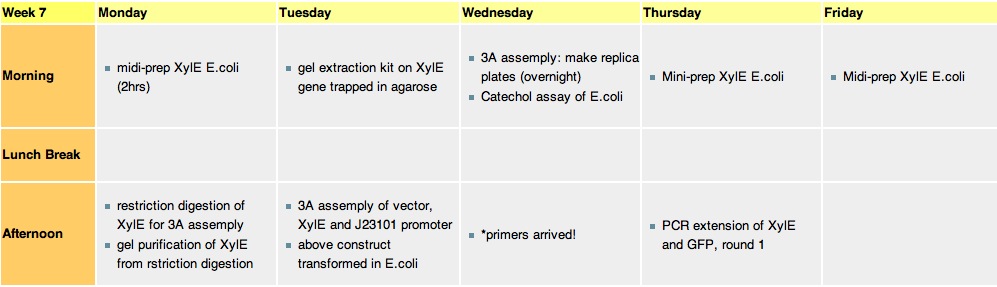
Week 7Monday, 16-Aug-2010
Tuesday, 17-Aug-2010
Thursday, 19-Aug-2010
Friday, 20-Aug-2010The J23101 gene in a biobrick vector containg RFP gene
|
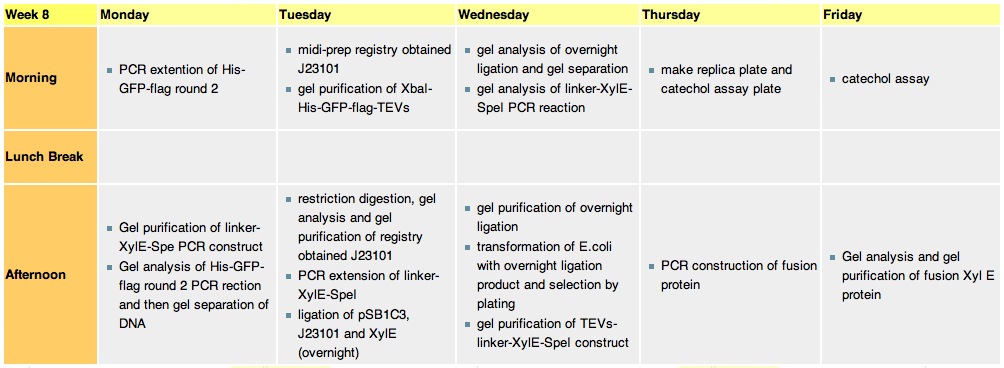
Week 8Monday, 23-Aug
Tuesday, 24th-Aug
Wednesday, 25th-AugPerformed gel analysis on the purified XylE and J23101 to obtain ratios for ligation. First gel was scrapped as it produced appauling(explanation for Nick:really bad) results, 2nd gel run was successful.
Thursday, 26th-Aug
Friday, 27th-Aug
File:Catechol Assay before.jpg Plate before adding catechol assay File:Catechol assay after (27-8).jpg After addition of catechol colonies turn yellow-orange in seconds!!
Saturday, 28th-Aug
Sunday, 29th-Aug
|
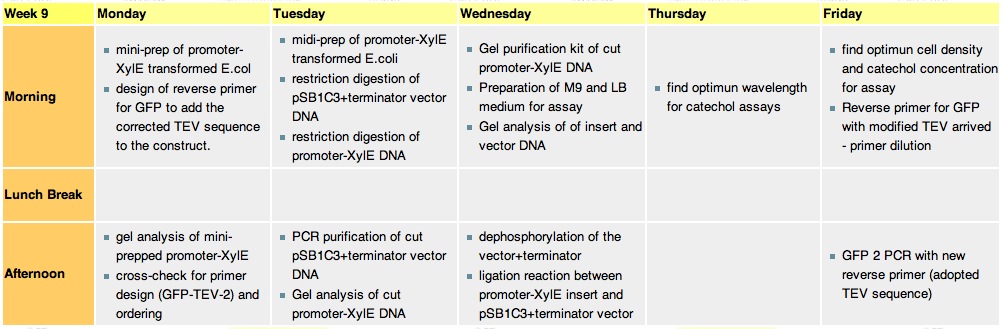
Week 9
Monday, 30th-Aug
Tuesday, 31st-Aug
Thursday, 2nd-SeptFile:200-600nm spectr.jpg Spectra of XylE transformed E.coli after addition of catechol assay. The broad peak around 380nm wavelength arises is due to the presence of the product of the enzymatic reaction involving pyrocatechol and XylE enzyme. This peak if absent if a culture of XylE transformed cells are measured without the addition of catechol
Friday, 3rd-Sept
File:Assay 3 sept.jpg Catechol assay on XylE-trasformed cells in a 96-well plate (A to H decreasing cell concentration, 1-10 decreasing catechol concentration, column 11 and 12 negative and control) |
 "
"