Team:Imperial College London/Lab Diaries/XylE team
From 2010.igem.org
| Line 244: | Line 244: | ||
*The new GFP + TEV primer arrived, was diluted and used to set up the appropriate PCR. | *The new GFP + TEV primer arrived, was diluted and used to set up the appropriate PCR. | ||
[[Image:Assay 3 sept.jpg|thumb|center|400px|Catechol assay on XylE-trasformed cells in a 96-well plate (A to H decreasing cell concentration, 1-10 decreasing catechol concentration, column 11 and 12 negative and control)]] | [[Image:Assay 3 sept.jpg|thumb|center|400px|Catechol assay on XylE-trasformed cells in a 96-well plate (A to H decreasing cell concentration, 1-10 decreasing catechol concentration, column 11 and 12 negative and control)]] | ||
| + | |} | ||
| + | |||
| + | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
| + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"| Lab notes and schedule | ||
| + | |- | ||
| + | | | ||
| + | |||
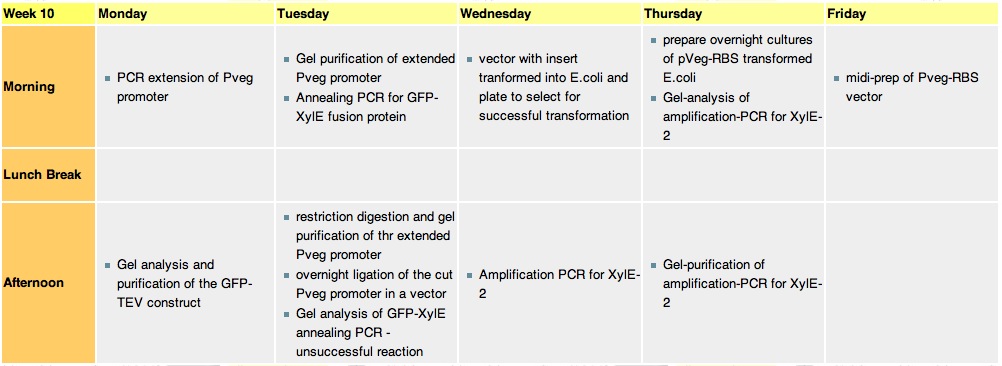
| + | =='''Week 10''' == | ||
| + | |||
| + | [[Image:wk10.jpg|thumb|800px|6.9.10-11.9.10]] | ||
| + | |||
| + | ===Monday, 6th-Sept=== | ||
| + | *PCR extension of Pveg promoter: EcoRI---Pveg---RBS-SpeI | ||
| + | |||
| + | *I performed a catechol assay on the picked transformed colonies to deduce which ones were successfully transformed with the insert plus vector. 1-5 7 and 10 failed to turn yellow (1-5 were background controls)leaving 8 yellow colonies. | ||
| + | *I had to perform a colony PCR on two selected colonies 8 and 14 to check the correct insert+terminator was present. | ||
| + | *Colony 8 when run on a gel analysis showed the correct size 924 XylE + 97 J23101 + 35 B0014. | ||
| + | *This was set up as an overnight culture. | ||
| + | *Gel analysis of the GFP-TEV construct showed satisfactory bands. | ||
| + | |||
| + | ===Tuesday 7th Sept=== | ||
| + | *Mini prep of the overnight Colony 8 culture (by Wolf) | ||
| + | *Midi prep of overnight culture | ||
| + | *gel purification of PCR product EcoRI--Pveg--RBS--SpeI | ||
| + | *overnight restriction digestion of EcoRI--Pveg--RBS--SpeI with SpeI | ||
| + | *started data analysis of plate assay | ||
| + | *Annealing PCR reaction included 2 samples without and one with additional primers (XbaI-His-GFP Fwd, SpeI-XylE Rev). While the primer-including reaction did not show any clearly identifiable bands, the others showed clear, if very weak bands at 800 and 1000bp, which represent the GFP and XylE constructs respectively. No band was identifiable at the 1.7 kb range, which would have indicated a successful annealing reaction. However, problems with the lense of the gel-analyser were only discovered later to have severely reduced the band brightness. Potentially a PCR-reaction with appropriate primers to amplify an annealed product(XbaI-His-GFP Fwd, SpeI-XylE Rev), could have been successful. However the reaction mixture was disposed of before this became clear. | ||
| + | |||
| + | ===Wednesday, 8th Sep=== | ||
| + | *A second PCR extension of Pveg promoter to introduce the RBS and cut sites. It was also gel purified and stored in gel lumbs in the freezer. (maybe needed later so keep in mind). | ||
| + | *Overnight restriction digestion completed. Then, it was run on the gel to check if it worked and then gel purified again. | ||
| + | *Vector PSBI-C3 was digested to remove the terminator and make it sticky for the insert (Pveg-RBS). Then it was run on the gel to check if restriction has worked, but the gel didn't run far enough to determine easily between undigested and digested vector. | ||
| + | * For further annealing reactions for GFP-XylE constructs additional XylE(2) template was required -> amplification PCR for XylE(2). | ||
| + | |||
| + | ==Thursday, 9th Sep== | ||
| + | *Gel with digested and undigested vector PSBI-C3 was run and then the digested vector was purified. | ||
| + | *Gel was run to determine the DNA concentration ratio for the ligation of PSBI-C3 and Pveg-RBS. | ||
| + | *Vector PSBI-C3 was dephosphorylated. | ||
| + | *Ligation of PSBI-C3 and Pveg-RBS has been set up overnight. | ||
| + | *Gel-analysis and gel-purification of the XylE(2) amplification PCR product. | ||
| + | ==Thursday, 10th Sep== | ||
| + | *The transformation of E.Coli with Pveg-RBS in PSBI-C3 and PSBI-C3 by itself (to check see how successful dephosphoryaltion of PSBI-C3 was and estimate the percentage of bacteria that contain the insert) was completed | ||
| + | |||
| + | *concentration of the midi prep of J23101-XylE-B0014 was determined to be ~600ng/ul (using new protocol) | ||
| + | *Kyasha kindly digested my midi with EcoRI and SpeI and performed a gel analysis. The results show a potential additional plasmid contaminating my midi however the concentration of DNA was extremely high. NB Chris said that it could be sheared DNA from a midi prep step. | ||
| + | |||
| + | *the midi prepped plasmid was transformed into testing E.coli strain TOP10. | ||
| + | |||
| + | ===Friday, 10 sep=== | ||
| + | |||
| + | The transformation was a SUCCESS. 2x replica plates were made plate 1# 1-6 plate #2 6-11; colony 6 and colony 9 of plates 1 and 2 respectively were transfered into a 5ml liquid culture + 5ul CmR. These will later be turned into glycerol stocks. After the replica plates have grown up mini preps on a number of colonies shall be performed - this hopefully will eliminate the contaminating plasmid DNA. This will be followed by a midi prep. | ||
| + | |||
| + | ===Saturday, 11 Sep=== | ||
| + | |||
| + | The transofrmation of E.Coli with PSB1-C3 with insert did not work :( | ||
|} | |} | ||
Revision as of 09:41, 21 October 2010
| Lab Diaries | Overview | Surface Protein Team | XylE Team | Vectors Team | Modelling Team |
| Here are the technical diaries for our project. We've split them up into three lab teams and the modelling team. We think it's really important that absolutely anyone can find out what we've been doing. For a really detailed look at what we did, and when, you've come to the right place! | |
| Team XylE |
Follow our progress: Click me!
| XylE team Lab Objectives |
|
| Lab notes and schedule |
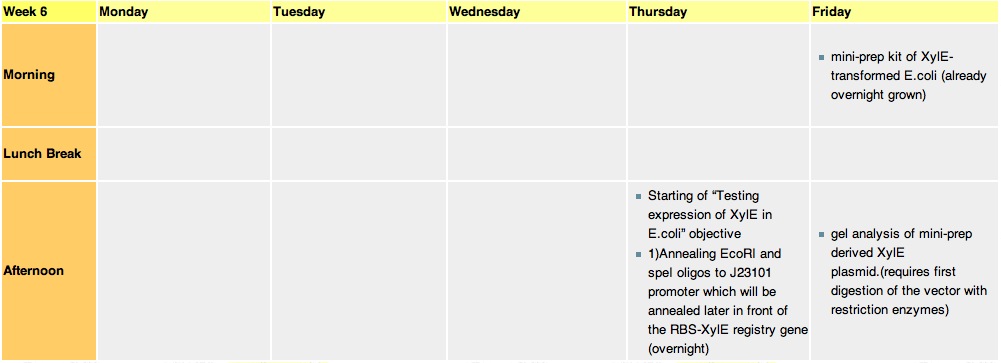
Week 6Thursday, 12-Aug-2010
we constructed the standard E.coli promoter J23101 with sticky ends. These ends are complementary to restriction sites made by EcoRI and SpeI enzyme. This promoter will be later used in 3A assemply to construct a promoter-RBS-XylE design in a psB1C3 vector. E.coli will be transformed with this final construct plasmid to assess XylE activity and characterization. It will also be one of the submitted biobricks.
these cultures are going to be used tomorrow for mini-prepping. Miniprep will allow us to isolate E.coli's plasmid DNA(which contains the XylE gene). Friday, 13-Aug-2010
Mini-prep is usually used to confirm that our gene of interest has not been changed in any way, as the isolated plasnid id sent for sequencing. However, since XylE was taken from the registry, we assume that it is fine and no sequencing is required. The mini-prep will later be used for the midi-prep (that gives out higher yeilds of DNA needed for cloning).
|
|
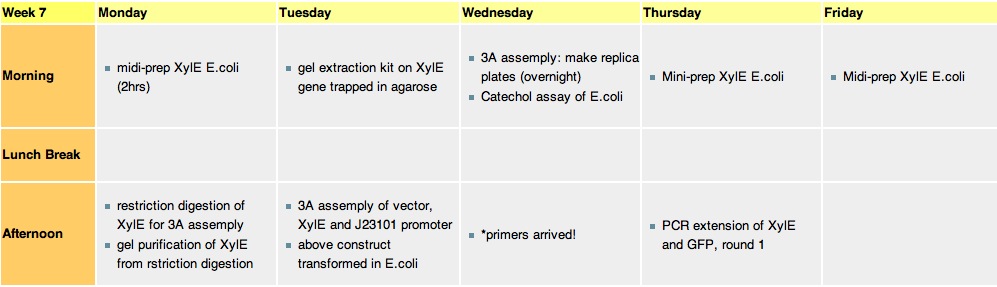
Week 7Monday, 16-Aug-2010
Tuesday, 17-Aug-2010
Thursday, 19-Aug-2010
Friday, 20-Aug-2010The J23101 gene in a biobrick vector containg RFP gene
|
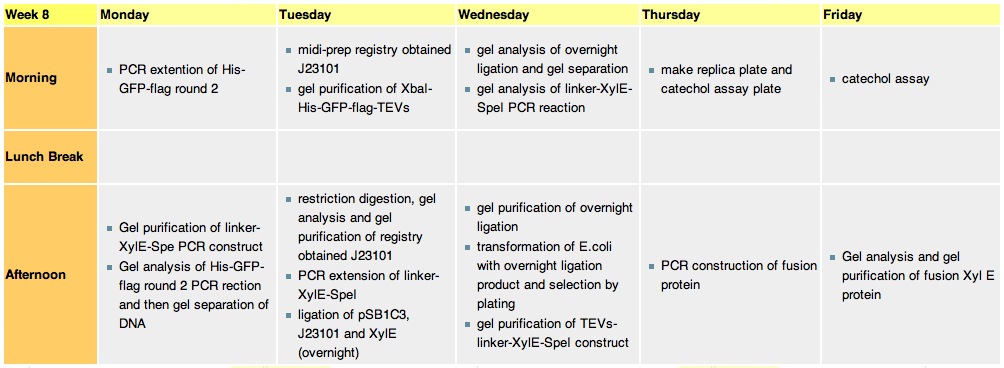
Week 8Monday, 23-Aug
Tuesday, 24th-Aug
Wednesday, 25th-AugPerformed gel analysis on the purified XylE and J23101 to obtain ratios for ligation. First gel was scrapped as it produced appauling(explanation for Nick:really bad) results, 2nd gel run was successful.
Thursday, 26th-Aug
Friday, 27th-Aug
File:Catechol Assay before.jpg Plate before adding catechol assay File:Catechol assay after (27-8).jpg After addition of catechol colonies turn yellow-orange in seconds!!
Saturday, 28th-Aug
Sunday, 29th-Aug
|
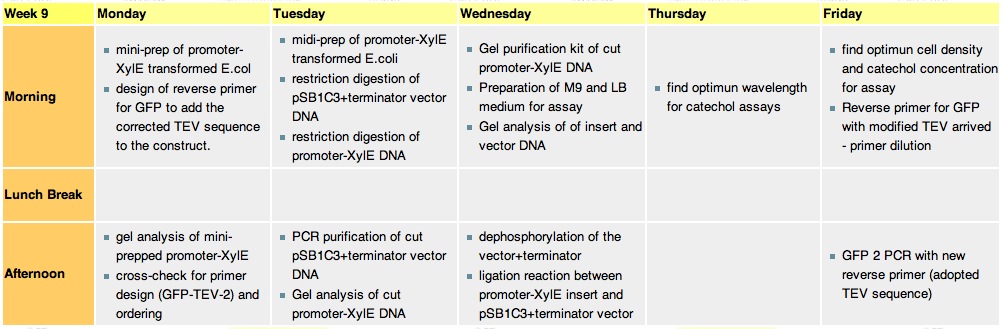
Week 9
Monday, 30th-Aug
Tuesday, 31st-Aug
Thursday, 2nd-SeptFile:200-600nm spectr.jpg Spectra of XylE transformed E.coli after addition of catechol assay. The broad peak around 380nm wavelength arises is due to the presence of the product of the enzymatic reaction involving pyrocatechol and XylE enzyme. This peak if absent if a culture of XylE transformed cells are measured without the addition of catechol
Friday, 3rd-Sept
File:Assay 3 sept.jpg Catechol assay on XylE-trasformed cells in a 96-well plate (A to H decreasing cell concentration, 1-10 decreasing catechol concentration, column 11 and 12 negative and control) |
| Lab notes and schedule |
Week 10Monday, 6th-Sept
Tuesday 7th Sept
Wednesday, 8th Sep
Thursday, 9th Sep
Thursday, 10th Sep
Friday, 10 sepThe transformation was a SUCCESS. 2x replica plates were made plate 1# 1-6 plate #2 6-11; colony 6 and colony 9 of plates 1 and 2 respectively were transfered into a 5ml liquid culture + 5ul CmR. These will later be turned into glycerol stocks. After the replica plates have grown up mini preps on a number of colonies shall be performed - this hopefully will eliminate the contaminating plasmid DNA. This will be followed by a midi prep. Saturday, 11 SepThe transofrmation of E.Coli with PSB1-C3 with insert did not work :( |
 "
"