Team:HokkaidoU Japan/Notebook/September14
From 2010.igem.org
(Difference between revisions)
| (11 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{Template:HokkaidoU_Japan}}<div class="linkbar"><div class="prev">[[Team:HokkaidoU_Japan/Notebook/September13|September 13]]</div>[[Team:HokkaidoU_Japan/Notebook|Notebook]]<div class="next">[[Team:HokkaidoU_Japan/Notebook/September15|September 15]]</div></div> | {{Template:HokkaidoU_Japan}}<div class="linkbar"><div class="prev">[[Team:HokkaidoU_Japan/Notebook/September13|September 13]]</div>[[Team:HokkaidoU_Japan/Notebook|Notebook]]<div class="next">[[Team:HokkaidoU_Japan/Notebook/September15|September 15]]</div></div> | ||
| - | =pSB1C3, araC Promoter | + | =Digestion and ligation of pSB1C3, araC Promoter and GFP= |
| + | |||
==Digestion== | ==Digestion== | ||
| - | [[Image:HokkaidoU Japan 20100914a.jpg|200px|right|thumb|]] | + | |
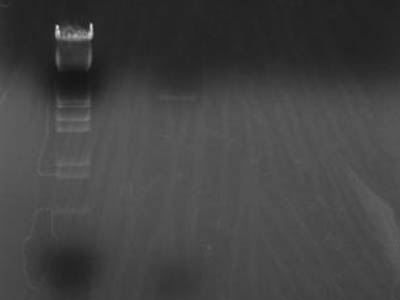
| - | + | [[Image:HokkaidoU Japan 20100914a.jpg|200px|right|thumb|Electrophoresed digestion product]] | |
| - | {|style="text-align:center; float:left | + | |
| + | Used pSB1C3 which was PCRed september 6th. DNA concentration was 2256 ng/uL. So diluted 20 times. | ||
| + | |||
| + | {|style="text-align:center; float:left;" class="protocol" | ||
| + | !Reagent | ||
| + | !Amount | ||
|- | |- | ||
|10x M buffer | |10x M buffer | ||
| Line 25: | Line 31: | ||
|3 | |3 | ||
|- | |- | ||
| - | |style="border-top:1px solid # | + | |style="border-top:1px solid #996;"|'''Total''' |
| - | |style="border-top:1px solid # | + | |style="border-top:1px solid #996;"|'''50 uL''' |
|} | |} | ||
| - | {|style="text-align:center; float:left | + | {|style="text-align:center; float:left;" class="protocol" |
| + | !Reagent | ||
| + | !Amount | ||
|- | |- | ||
|10x M buffer | |10x M buffer | ||
| Line 48: | Line 56: | ||
|2 | |2 | ||
|- | |- | ||
| - | |style="border-top:1px solid # | + | |style="border-top:1px solid #996;"|'''Total''' |
| - | |style="border-top:1px solid # | + | |style="border-top:1px solid #996;"|'''50 uL''' |
|} | |} | ||
| - | {|style="text-align:center; float:left | + | {|style="text-align:center; float:left;" class="protocol" |
| + | !Reagent | ||
| + | !Amount | ||
|- | |- | ||
|10x M buffer | |10x M buffer | ||
| Line 71: | Line 81: | ||
|1 | |1 | ||
|- | |- | ||
| - | |style="border-top:1px solid # | + | |style="border-top:1px solid #996;"|'''Total''' |
| - | |style="border-top:1px solid # | + | |style="border-top:1px solid #996;"|'''50 uL''' |
|} | |} | ||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
| - | + | →Incubated at 37C for 120 min<br> | |
| - | + | →Added Sample Buffer 10 uL and electrophoresed | |
| - | * | + | * pSB1C3 band wasn't visible |
| - | + | → Other 2 were gel extracted | |
| - | * 40 | + | * Diluted that final volume would be 40 uL |
| + | ==[[Team:HokkaidoU_Japan/Protocols|Ethanol precipitation]]== | ||
| + | [[Image:HokkaidoU Japan 20100914b.jpg|200px|right|thumb|Electrophoresis after ethanol precipitation]] | ||
| - | + | * Added ethanol acetate[3 M] 4 uL to 40 uL of previously purified DNA solution | |
| - | [[ | + | * Added EtOH[100%] 44 uL froze in dry ice |
| - | * | + | * Melted and centrifuged at 15000 rpm, 4C 5 min |
| - | * | + | * Discarded supernatant and added EtOH[70%] 100 uL rinsed |
| - | * | + | * Centrifuged at 15,996rpm, 4C 5 min |
| - | * | + | * Discarded supernatant and dried |
| - | * | + | * Diluted in 2 uL of TE each |
| - | promoter: 125 ng/ | + | *Anticipated concentrations<br> |
| - | + | promoter: 125 ng/2uL<br> | |
| - | + | GFP: 106 ng/2uL | |
| - | + | ||
| + | * pSB1C3 was cut(EcoR1, Pst1) on September 6th | ||
| + | * Did ethanol precipitation and electrophoresed to check concentration | ||
| + | * It was about 20 ng/uL | ||
| + | →Used 2 uL of it | ||
==Ligation== | ==Ligation== | ||
| - | {|style="text-align:center;" | + | |
| + | {|style="text-align:center;" class="protocol" | ||
| + | !Reagent | ||
| + | !Amount | ||
|- | |- | ||
|pSB1C3 | |pSB1C3 | ||
| Line 115: | Line 133: | ||
|0.4 uL | |0.4 uL | ||
|- | |- | ||
| - | |style="border-top:1px solid # | + | |style="border-top:1px solid #996;"|'''Total''' |
| - | |style="border-top:1px solid # | + | |style="border-top:1px solid #996;"|'''12.4 uL''' |
|} | |} | ||
| - | + | *50 uL were transformed by electroporation and remaining 50 uL by heat shock (chemical) | |
Latest revision as of 08:20, 27 October 2010
Digestion and ligation of pSB1C3, araC Promoter and GFP
Digestion
Used pSB1C3 which was PCRed september 6th. DNA concentration was 2256 ng/uL. So diluted 20 times.
| Reagent | Amount |
|---|---|
| 10x M buffer | 5 uL |
| DW | 34 |
| pSB1C3 | 1 |
| BSA | 5 |
| EcoR I | 2 |
| Pst I | 3 |
| Total | 50 uL |
| Reagent | Amount |
|---|---|
| 10x M buffer | 5 uL |
| DW | 27 |
| Promoter | 8 |
| BSA | 5 |
| EcoR I | 3 |
| Spe I | 2 |
| Total | 50 uL |
| Reagent | Amount |
|---|---|
| 10x M buffer | 5 uL |
| DW | 32 |
| GFP | 3 |
| BSA | 5 |
| Xba I | 4 |
| Pst I | 1 |
| Total | 50 uL |
→Incubated at 37C for 120 min
→Added Sample Buffer 10 uL and electrophoresed
- pSB1C3 band wasn't visible
→ Other 2 were gel extracted
- Diluted that final volume would be 40 uL
Ethanol precipitation
- Added ethanol acetate[3 M] 4 uL to 40 uL of previously purified DNA solution
- Added EtOH[100%] 44 uL froze in dry ice
- Melted and centrifuged at 15000 rpm, 4C 5 min
- Discarded supernatant and added EtOH[70%] 100 uL rinsed
- Centrifuged at 15,996rpm, 4C 5 min
- Discarded supernatant and dried
- Diluted in 2 uL of TE each
- Anticipated concentrations
promoter: 125 ng/2uL
GFP: 106 ng/2uL
- pSB1C3 was cut(EcoR1, Pst1) on September 6th
- Did ethanol precipitation and electrophoresed to check concentration
- It was about 20 ng/uL
→Used 2 uL of it
Ligation
| Reagent | Amount |
|---|---|
| pSB1C3 | 2 uL |
| promoter | 2 uL |
| GFP | 2 uL |
| Mighty Mix | 6 uL |
| T4 ligase | 0.4 uL |
| Total | 12.4 uL |
- 50 uL were transformed by electroporation and remaining 50 uL by heat shock (chemical)
 "
"