Team:Heidelberg/Project/Mouse Infection
From 2010.igem.org
| Line 30: | Line 30: | ||
All but one virus were packaged by the AAV rep and cap gene with Adenovirus 5 (Ad5) as a helper plasmid. Accordingly, one virus construct was packaged into a shuffled cap gene from our [https://2010.igem.org/Team:Heidelberg/Project/Capsid_Shuffling/Homology_Based homology based capsid shuffling] attempt. | All but one virus were packaged by the AAV rep and cap gene with Adenovirus 5 (Ad5) as a helper plasmid. Accordingly, one virus construct was packaged into a shuffled cap gene from our [https://2010.igem.org/Team:Heidelberg/Project/Capsid_Shuffling/Homology_Based homology based capsid shuffling] attempt. | ||
<br/> | <br/> | ||
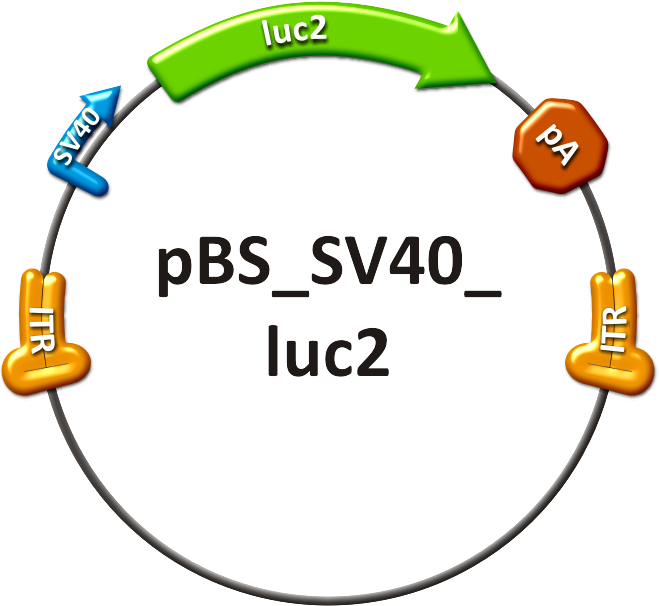
| - | # The positive control (see sidebar, fig. 1) consisted of the SV40 promoter driving a firefly luciferase (luc2) gene, thereby leading to an unspecific expression of the luciferase protein in all mice tissues. | + | # The positive control (see sidebar, fig. 1) consisted of the SV40 promoter driving a firefly luciferase (luc2) gene, thereby leading to an unspecific expression of the luciferase protein in all mice tissues. In addition to packaging this construct into a wild type AAV virus, the positive control was also packaged as a transgene into our [https://2010.igem.org/Team:Heidelberg/Project/Capsid_Shuffling/Homology_Based shuffled capsid] which after random selection <!--selection pressure--> was already able to [https://2010.igem.org/Team:Heidelberg/Notebook/Homology_Based/October#16/10/2010 positively transduce Huh7 and HepG2 cells] <i>in vitro</i>. |
| - | In addition to packaging this construct into a wild type AAV virus, the positive control was also packaged as a transgene into our [https://2010.igem.org/Team:Heidelberg/Project/Capsid_Shuffling/Homology_Based shuffled capsid] which after random selection <!--selection pressure--> was already able to [https://2010.igem.org/Team:Heidelberg/Notebook/Homology_Based/October#16/10/2010 positively transduce Huh7 and HepG2 cells] <i>in vitro</i>. | + | |
# The off-targeting construct (see sidebar, fig. 2) was composed of an SV40 promoter driving a firefly luciferase (luc2) gene with binding sites against miR-122 behind it. In order to achieve the highest expression in all mice cells but the liver cells - a single perfect binding site of miR-122 was used for <i>in vivo</i> study. | # The off-targeting construct (see sidebar, fig. 2) was composed of an SV40 promoter driving a firefly luciferase (luc2) gene with binding sites against miR-122 behind it. In order to achieve the highest expression in all mice cells but the liver cells - a single perfect binding site of miR-122 was used for <i>in vivo</i> study. | ||
# The synthetic tuning construct (see sidebar, fig. 3) consisted of two viruses injected at the same time in the mice. The one virus packaged the expression construct of shRNA haat driven by the H1 promoter ("tuning" construct, see sidebar, fig.3). The second virus packaged the following transgene: SV40 promoter driving luc2 with shRNA haat binding site behind it ("tuned" construct, see sidebar, fig. 4). In order to ensure a synthetic tuning effect, a perfect binding site and one with a bulge that was introduced at position 9-12 were used for <i>in vivo</i> experiments, respectively. Those two binding sites should lead to a significant knockdown in the first case and a slight repression of luciferase expression in the latter as compared to the positive control. | # The synthetic tuning construct (see sidebar, fig. 3) consisted of two viruses injected at the same time in the mice. The one virus packaged the expression construct of shRNA haat driven by the H1 promoter ("tuning" construct, see sidebar, fig.3). The second virus packaged the following transgene: SV40 promoter driving luc2 with shRNA haat binding site behind it ("tuned" construct, see sidebar, fig. 4). In order to ensure a synthetic tuning effect, a perfect binding site and one with a bulge that was introduced at position 9-12 were used for <i>in vivo</i> experiments, respectively. Those two binding sites should lead to a significant knockdown in the first case and a slight repression of luciferase expression in the latter as compared to the positive control. | ||
| Line 63: | Line 62: | ||
__TOC__ | __TOC__ | ||
<br/> | <br/> | ||
| - | |||
| - | |||
<br/> | <br/> | ||
| - | + | === construct schemes === | |
<br/> | <br/> | ||
| - | [[Image: | + | [[Image:PBS SV40 luc2.png|frameless|center|250px|'''Figure 1: Positive control construct.''']] |
<br/> | <br/> | ||
| - | [[Image:PBS SV40 luc2 | + | [[Image:PBS SV40 luc2 miR122.png|frameless|center|250px|'''Figure 2: Off-targeting construct.''']] |
<br/> | <br/> | ||
| - | [[Image: | + | [[Image:PBS_H1_shR_hAAT.png|frameless|center|250px|'''Figure 3: Tuning construct.''']] |
<br/> | <br/> | ||
| - | [[Image:Image:PBS SV40 TetO2 luc2.png| | + | [[Image:PBS SV40 luc2 hAAT.png|frameless|center|250px|'''Figure 4: Tuned construct.''']] |
| + | <br/> | ||
| + | [[Image:Image:PBS TetR miR122 4x.png|frameless|center|250px|'''Figure 5: Repressor construct.''']] | ||
| + | <br/> | ||
| + | [[Image:Image:PBS SV40 TetO2 luc2.png|frameless|center|250px|'''Figure 6: Operator construct.''']] | ||
{{:Team:Heidelberg/Bottom}} | {{:Team:Heidelberg/Bottom}} | ||
Revision as of 13:51, 27 October 2010

|
|
|||
 "
"