Team:Groningen/Expression
From 2010.igem.org
Joelkuiper (Talk | contribs) m (→References) |
(→Timed expression of chaplins in a biofilm) |
||
| Line 22: | Line 22: | ||
'''''srfA''''' | '''''srfA''''' | ||
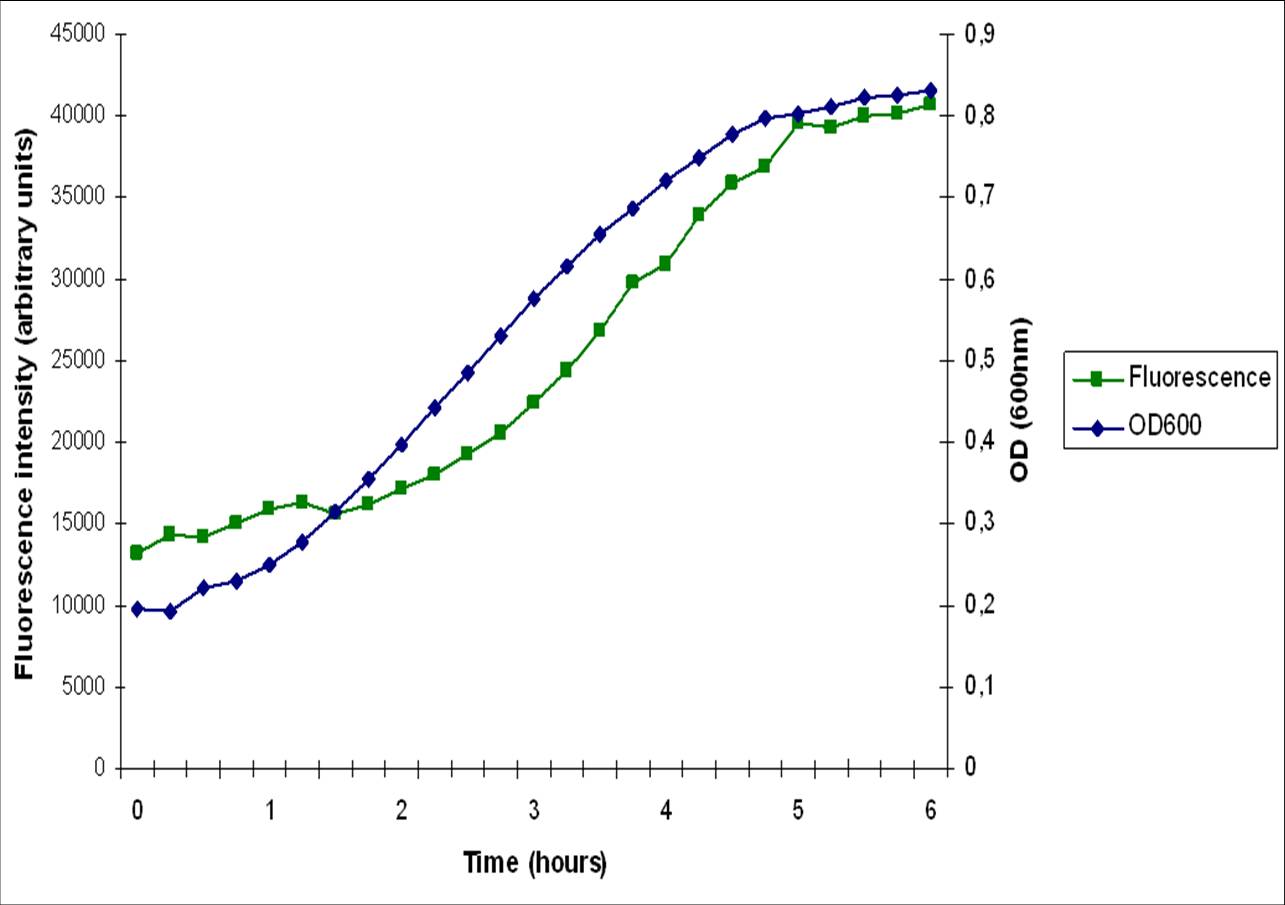
| - | The [http://dbtbs.hgc.jp/COG/prom/srfAA-srfAB-comS-srfAC-srfAD.html ''srfA'' operon] has been reported to be important for natural competence and sporulation in ''Bacillus subtilis''. All these activities occur in biofilms, the promoter is not active until the end of exponential growth. It is controlled by the [https://2010.igem.org/Team:Groningen/Expression_model#ComXPA_quorum_sensing_system ComXPA quorum sensing system] and hence active in states of high cell densities. Therefore the ''srfA'' promoter would be suitable for chaplin expression. Two different lengths of the ''srfA'' promoter where chosen due to uncertainties concerning the region between the response element and the transcription start side of the SrfAA protein. In the original promoter this region is unusually long, by shortening it 190bp’s we hope to achieve a higher transcription efficiency. So we came up with two different promoters, the [http://partsregistry.org/wiki/index.php?title=Part:BBa_K305008 original] one and the [http://partsregistry.org/wiki/index.php?title=Part:BBa_K305007 shortened] one. | + | The [http://dbtbs.hgc.jp/COG/prom/srfAA-srfAB-comS-srfAC-srfAD.html ''srfA'' operon] has been reported to be important for natural competence and sporulation in ''Bacillus subtilis''. All these activities occur in biofilms, the promoter is not active until the end of exponential growth. It is controlled by the [https://2010.igem.org/Team:Groningen/Expression_model#ComXPA_quorum_sensing_system ComXPA quorum sensing system] and hence active in states of high cell densities. Therefore the ''srfA'' promoter would be suitable for chaplin expression. Two different lengths of the ''srfA'' promoter where chosen due to uncertainties concerning the region between the response element and the transcription start side of the SrfAA protein. In the original promoter this region is unusually long, by shortening it 190bp’s we hope to achieve a higher transcription efficiency. So we came up with two different promoters, the [http://partsregistry.org/wiki/index.php?title=Part:BBa_K305008 original] one and the [http://partsregistry.org/wiki/index.php?title=Part:BBa_K305007 shortened] one. Fusions with GFP confirmed our assumption that the short srfA variant leads to a higher transcription. While the fluorescence off the short variant was clearly above background levels, the long variant did not give convincing results. |
| + | [[Image:florescence_srfA.jpg]] | ||
| Line 29: | Line 30: | ||
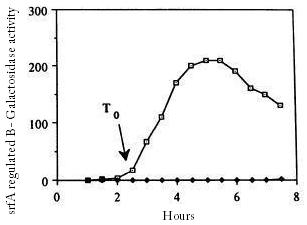
[[Image:igemgroningen_yqxm_prmoteractivity.jpg|right|200px|yqxm|thumb|yqxM promotor activity during cell growth of different mutants (Axel G. 1999)]] | [[Image:igemgroningen_yqxm_prmoteractivity.jpg|right|200px|yqxm|thumb|yqxM promotor activity during cell growth of different mutants (Axel G. 1999)]] | ||
| - | The [http://dbtbs.hgc.jp/COG/prom/yqxM-sipW-tasA.html ''yqxM-sipW-tasA''] operon is controlled by the ''yqxM'' promoter. It is needed for biofilm formation because TasA is a key protein of the extracellular matrix. The promotor gets activated via a cascade of other regulatory elements, including SrfA, in response to quorum sensing. Since the chaplins should work in a similar way to TasA we think the [http://partsregistry.org/wiki/index.php?title=Part:BBa_K305006 ''yqxM''] promoter would be very suitable for chaplin expression during the stationary phase. | + | The [http://dbtbs.hgc.jp/COG/prom/yqxM-sipW-tasA.html ''yqxM-sipW-tasA''] operon is controlled by the ''yqxM'' promoter. It is needed for biofilm formation because TasA is a key protein of the extracellular matrix. The promotor gets activated via a cascade of other regulatory elements, including SrfA, in response to quorum sensing. Since the chaplins should work in a similar way to TasA we think the [http://partsregistry.org/wiki/index.php?title=Part:BBa_K305006 ''yqxM''] promoter would be very suitable for chaplin expression during the stationary phase. We fused the yqxM promoter with GFP but could not observe any expression, since the GFP worked with the srfA promoter we conclude that the yqxM promoter does not work. |
Revision as of 17:18, 27 October 2010
Subtilin induced expression of chaplins
The Biofilm forming capacity of Bacillus subtilis makes it a good host for our application. In addition, B. subtilis is known for its ability to produce and export large amounts of protein at high cell densities. However, despite its track record as an efficient production organism and the fact that both B. subtilis and Streptomyces coelicolor are gram-positive bacteria, it is not certain wether chaplins can be heterologously expressed in B. subtilis. Improper folding, unsuccessful export, or even the very nature of the chaplins, could still lead to hampered expression. We took several steps to ensure optimal expression. The Coding sequences of the chaplins were codon optimized for B. subtilis and synthesized. We placed a ribosome binding site in front of the coding sequences that is known to work well in B. subtilis, and flanked these constructs with the biobrick prefix and suffix.
SURE expression system
Because it is uncertain how chaplin expression will affect B. subtilis, the initial expression attempts were performed with the stringently controlled, subtilin-regulated gene expression (SURE) system (Bongers et al, 2005). This system uses the subtilin sensing machinery present in a strain of B. subtilis that autoinduces the production of more of the lantibiotic subtilin. The subtilin sensor histidine kinase SpaK phosphorylates the response regulator SpaR, which can then bind to so-called spa boxes in the promoter regions of genes involved in subtilin biosynthesis (Kleerebezem et al, 2004). In the SURE system, a B. subtilis strain naturally lacking the subtilin biosynthesis genes has the spaRK genes introduced into its genome. A plasmid carrying a spa box promoter that is transformed to this strain can then drive the expression of proteins upon subtilin induction of SpaRK signalling.
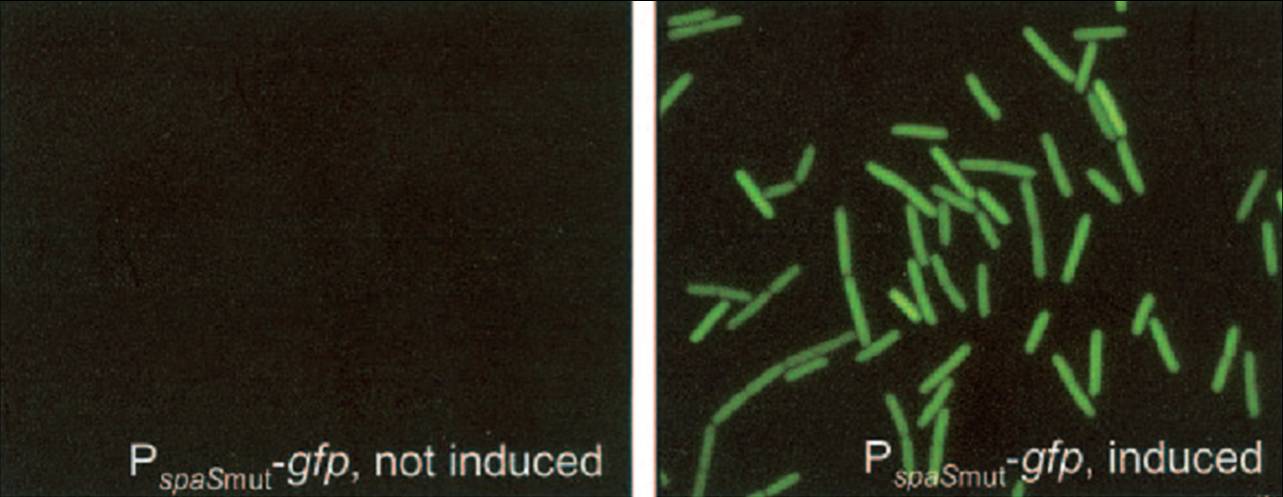
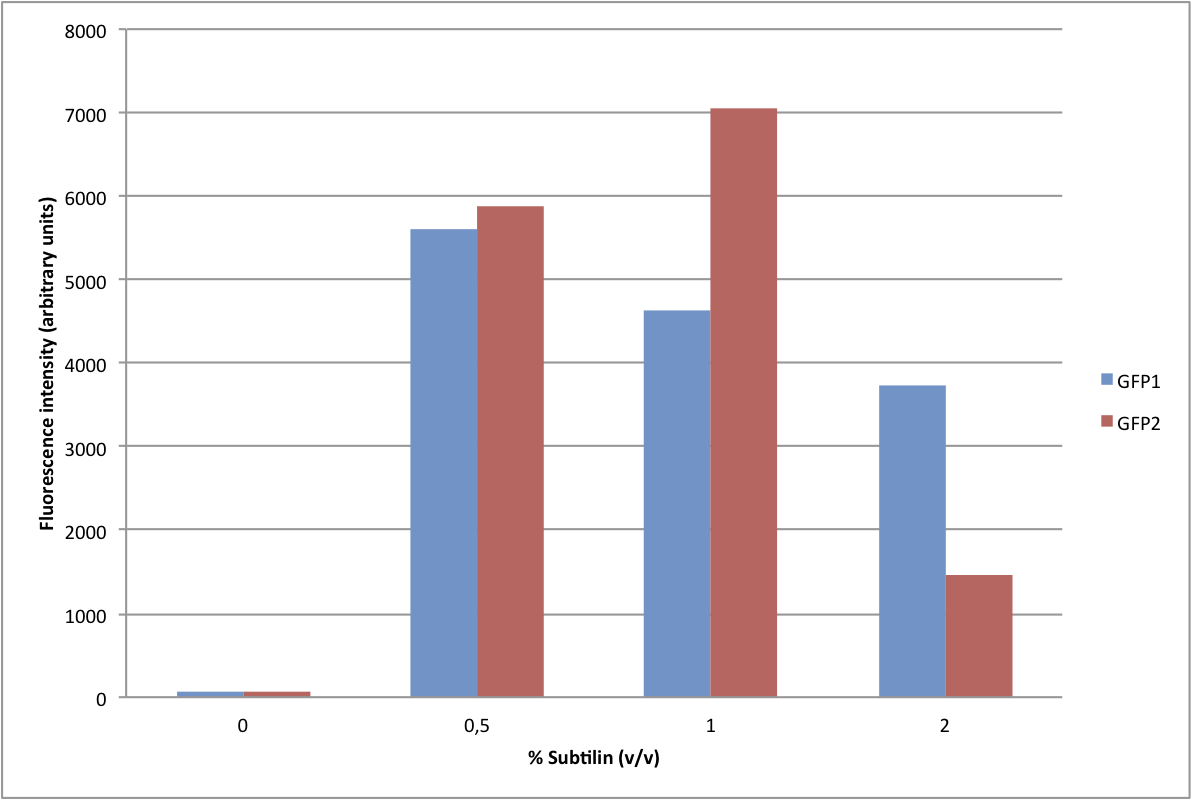
We have adapted this system to make it BioBrick compatible for easy expression of our chaplins, combinations of chaplins, or any other biobrick part that is composed of an RBS followed by a protein coding sequence. We introduced the BioBrick prefix and suffix into the expression plasmid, downstream of the mutated spaS promoter, producing our subtilin inducible expression backbone part, BBa_K305011. To test the expression and find a suitable subtilin concentration for induction of the chaplins we made use of GFP fluorescence measurements. We inserted the part BBa_E0240 into the BioBrick site and induced liquid cultures of B. subtilis carrying this plasmid (and the spaRK genes) with different volumes of subtilin-containing culture supernatant of a subtilin producing strain of B. subtilis. These results demonstrate that addition of 0.5 to 1%(vol/vol) of subtilin to the culture is sufficient to reach optimal induction. > Chaplins
Timed expression of chaplins in a biofilm
An important question is which promoter we should use to control the chaplin expression. We assume that an ideal promoter would not be active until the biofilm has formed because the expression of hydrophobic proteins might influence the formation of it. Two promoters where found that are active in biofilms but not during normal growth.
srfA
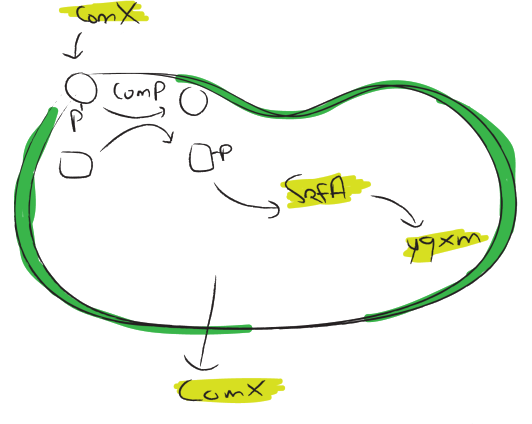
The srfA operon has been reported to be important for natural competence and sporulation in Bacillus subtilis. All these activities occur in biofilms, the promoter is not active until the end of exponential growth. It is controlled by the ComXPA quorum sensing system and hence active in states of high cell densities. Therefore the srfA promoter would be suitable for chaplin expression. Two different lengths of the srfA promoter where chosen due to uncertainties concerning the region between the response element and the transcription start side of the SrfAA protein. In the original promoter this region is unusually long, by shortening it 190bp’s we hope to achieve a higher transcription efficiency. So we came up with two different promoters, the original one and the shortened one. Fusions with GFP confirmed our assumption that the short srfA variant leads to a higher transcription. While the fluorescence off the short variant was clearly above background levels, the long variant did not give convincing results.
yqxM
The yqxM-sipW-tasA operon is controlled by the yqxM promoter. It is needed for biofilm formation because TasA is a key protein of the extracellular matrix. The promotor gets activated via a cascade of other regulatory elements, including SrfA, in response to quorum sensing. Since the chaplins should work in a similar way to TasA we think the yqxM promoter would be very suitable for chaplin expression during the stationary phase. We fused the yqxM promoter with GFP but could not observe any expression, since the GFP worked with the srfA promoter we conclude that the yqxM promoter does not work.
References
Bongers RS, Veening JW, Van Wieringen M, Kuipers OP, and Kleerebezem M. Development and characterization of a subtilin-regulated expression system in Bacillus subtilis: strict control of gene expression by addition of subtilin. Environ Microbiol 2005 Dec; 71(12) 8818-24. pmid:16332878
Kleerebezem, M., R. Bongers, G. Rutten, W. M. de Vos, and O. P. Kuipers. 2004. Autoregulation of subtilin biosynthesis in Bacillus subtilis: the role of the spa-box in subtilin-responsive promoters. Peptides 25:1415–1424
Stöver AG, Driks A. Regulation of synthesis of the Bacillus subtilis transition-phase, spore-associated antibacterial protein TasA. J. Bacteriol. Sept. 1999, p. 5476-5481, Vol. 181, No. 17
Nakano MM, Xia LA, Zuber P. Transcription initiation region of the srfA operon, which is controlled by the comP-comA signal transduction system in Bacillus subtilis. PMC208261
Frances Chu, Daniel B. Kearns, Anna McLoon, Yunrong Chai, Roberto Kolter and Richard Losicka, A Novel Regulatory Protein Governing Biofilm Formation in Bacillus subtilis PMC2430766
Hayashi K, Ohsawa T, Kobayashi K, Ogasawara N, Ogura M. The H2O2 stress-responsive regulator PerR positively regulates srfA expression in Bacillus subtilis. PMC1251593
 "
"