Team:Cambridge/References/ProjectBioluminescence/Luciferase
From 2010.igem.org
(Difference between revisions)
(→luxCDABE (Vibrio Fischeri/Harvyi)) |
|||
| (61 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | == | + | {{:Team:Cambridge/Templates/headerMinimalprototype}} |
| + | {{:Team:Cambridge/Templates/RefBar}} | ||
| + | {{:Team:Cambridge/Templates/headerbar|colour=#96d446|title=Bioluminescence: Luciferases}} | ||
| - | == | + | ==Mutant luciferase (P. pyralis)== |
| + | *[http://www.ncbi.nlm.nih.gov/nuccore/AB261988.1 Mutant Enhanced Photon Initiating Complex (EPIC) Luciferase (P.pyralis) Sequence] | ||
| + | *[http://www.ncbi.nlm.nih.gov/pubmed/10529195 Site-directed mutagenesis of firefly luciferase active site amino acids: a proposed model for bioluminescence color] | ||
| + | *[http://www.ncbi.nlm.nih.gov/pubmed/12044905 Improved practical usefulness of firefly luciferase by gene chimerization and random mutagenesis] | ||
| - | == | + | ==Other Insect luciferases== |
| + | *For colours we could use the click beetle luciferase [http://partsregistry.org/wiki/index.php/Part:BBa_J70005 red] or [http://partsregistry.org/wiki/index.php/Part:BBa_J70006 green]. They're only at the 'planning' stage though. | ||
| - | *[Team:Cambridge/ | + | ==Bacterial Bioluminescence== |
| - | + | *[http://arjournals.annualreviews.org/doi/pdf/10.1146/annurev.mi.42.100188.001055?cookieSet=1 paper with hard numbers, pathways] | |
| - | * | + | *[http://www.ncbi.nlm.nih.gov/pmc/articles/PMC372803/pdf/microrev00032-0137.pdf The definitive summary paper] |
| - | * | + | *[[Team:Cambridge/VibrioFischeri| Vibrio Fischeri]] |
| - | * | + | *[[Team:Cambridge/VibrioHarvei| Vibrio Harveyi]] |
| - | * | + | *[[Team:Cambridge/Phosphoreum| Vibrio Phosphoreum]] |
| - | * | + | *[[Team:Cambridge/Photorhabdus| Photorhabdus]] |
| - | * | + | *[[Team:Cambridge/ProjectBioluminescence/Luciferase/WikiGeneticsLuxCDABE| Genetics of luxCDABE (adapted from wikipedia)]] |
| - | * | + | *[https://2006.igem.org/Lux_operon| 2006 attempts at BioBricking the lux operon] |
| + | *[[Team:Cambridge/ProjectBioluminescence/Luciferase/Notes| Notes]] | ||
| + | *[http://partsregistry.org/Lux The lux wiki from parts registry] | ||
| + | *[http://www.ncbi.nlm.nih.gov/nuccore/AF170104.1?report=graph&log$=seqview NCBI Lux operon sequence] | ||
| + | *[http://partsregistry.org/Part:BBa_G10001 parts registry lux operon] -currently waiting on an email to see how useful/available this part is | ||
| + | *[[Team:Cambridge/ProjectBioluminescence/Luciferase/IMPORTANT INFO | IMPORTANT INFO]] | ||
| + | * The Photorhabdus luminescens [http://partsregistry.org/Part:BBa_J70008 luxCDABE operon] in the registry. It's not well characterised | ||
| + | |||
| + | [[Image:Luxoperondiagram.jpg]] | ||
| + | |||
| + | ==Plasmid experiment== | ||
| + | |||
| + | *[http://departments.kings.edu/biology/lux/bacterial.html Vibrio plasmid experiment] -possible source of lux operon | ||
| + | |||
| + | [[Image:4449975.pdf | article detailing proceedure and plasmids]] | ||
| + | |||
| + | *[http://departments.kings.edu/biology/lux/index.html] - Transformation protocol for students from basics - includes several plasmids | ||
| + | |||
| + | *Plan | ||
| + | **set plasmid 555 into a BioBrick | ||
| + | **replace the promoter? sensitivity tuners? | ||
| + | |||
| + | *references | ||
| + | ** [http://www.pnas.org/content/84/19/6639.short Overproduction and purification of the luxR gene product: Transcriptional activator of the Vibrio fischeri luminescence system (Kaplan & Greenburg 1987) ] | ||
| + | ** [http://jb.asm.org/cgi/content/abstract/172/7/3974 Critical regions of the Vibrio fischeri luxR protein defined by mutational analysis (Slock et al 1990)] | ||
| + | |||
| + | ==Transformation Experiment to test Firefly luciferase in registry== | ||
| + | |||
| + | * Firefly luciferase is in the registry as [http://partsregistry.org/Part:BBa_I712019 BBa_I712019], and have been sent to us as part of the spring 2010 distribution. | ||
| + | |||
| + | *We need to find a new vector with a bacterial ribosyme binding site, initiator, and a stop codon. Having searched via the catalog in the plasmids section, then in expression plasmids, we have selected a constitutive plasmid: | ||
| + | ** The plasmid selected is [http://www.partsregistry.org/Part:BBa_J13002 BBa_J13002], which is a TetR repressed POPS/RIPS generator, again part of the spring 2010 distribution - kit plate 1, well 13B, plasmid pSB1A2. It was proven to work by UMN iGEM 2009. | ||
| + | ==Wavelengths== | ||
| + | 548 nm http://pubs.acs.org/doi/pdf/10.1021/bi7015052 <html><img src="http://www4a.wolframalpha.com/Calculate/MSP/MSP545519beai43119h6bi900003450dhf2577a40b8?MSPStoreType=image/gif&s=62&w=99&h=20"></html> | ||
| + | |||
| + | 613 nm http://www.promega.com/pnotes/85/10904_11/10904_11.pdf <html><img src="http://www4a.wolframalpha.com/Calculate/MSP/MSP52619bedhd64d9hi9240000123ei80gh721227b?MSPStoreType=image/gif&s=40&w=99&h=20"></html> | ||
| + | <html> | ||
| + | </div> | ||
| + | </html> | ||
| + | |||
| + | {| class="wikitable" | ||
| + | !Luciferase!!Colour(at pH7.8)!!λ<sub>max</sub>(nm)!!(pH 7.8)!!(pH 6.0)!!Base change!!Amino acid change!! | ||
| + | |- | ||
| + | |Genjii||yellow-green||||562||609 | ||
| + | |- | ||
| + | |C-M-1||orange||||607||614||G857-->A||Ser286-->Asn | ||
| + | |- | ||
| + | |C-M-2||red||||609||611||G976-->A||Gly326-->Ser | ||
| + | |- | ||
| + | |C-M-3||red||||612||612||C1297-->T||His433-->Tyr | ||
| + | |- | ||
| + | |C-M-4||yellow-orange||||595||609||C1354-->T||Pro452-->Ser | ||
| + | |- | ||
| + | |C-M-6||green||||558||558||G715-->A||Val239-->Ile | ||
| + | |- | ||
| + | |C-M-11||yellow||||565||612 | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | {{:Team:Cambridge/Templates/footerMinimal}} | ||
Latest revision as of 12:18, 7 October 2010

Link dumps: Bioluminescence |
Firefly luciferases|
Lucferin recovery|
Light output|
Experiments|
Modelling
Bioluminescence: Luciferases
Mutant luciferase (P. pyralis)
- Mutant Enhanced Photon Initiating Complex (EPIC) Luciferase (P.pyralis) Sequence
- Site-directed mutagenesis of firefly luciferase active site amino acids: a proposed model for bioluminescence color
- Improved practical usefulness of firefly luciferase by gene chimerization and random mutagenesis
Other Insect luciferases
- For colours we could use the click beetle luciferase red or green. They're only at the 'planning' stage though.
Bacterial Bioluminescence
- paper with hard numbers, pathways
- The definitive summary paper
- Vibrio Fischeri
- Vibrio Harveyi
- Vibrio Phosphoreum
- Photorhabdus
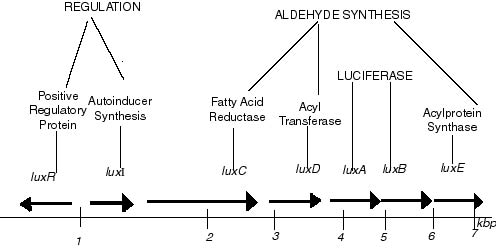
- Genetics of luxCDABE (adapted from wikipedia)
- 2006 attempts at BioBricking the lux operon
- Notes
- The lux wiki from parts registry
- NCBI Lux operon sequence
- parts registry lux operon -currently waiting on an email to see how useful/available this part is
- IMPORTANT INFO
- The Photorhabdus luminescens luxCDABE operon in the registry. It's not well characterised
Plasmid experiment
- Vibrio plasmid experiment -possible source of lux operon
- [1] - Transformation protocol for students from basics - includes several plasmids
- Plan
- set plasmid 555 into a BioBrick
- replace the promoter? sensitivity tuners?
- references
Transformation Experiment to test Firefly luciferase in registry
- Firefly luciferase is in the registry as BBa_I712019, and have been sent to us as part of the spring 2010 distribution.
- We need to find a new vector with a bacterial ribosyme binding site, initiator, and a stop codon. Having searched via the catalog in the plasmids section, then in expression plasmids, we have selected a constitutive plasmid:
- The plasmid selected is BBa_J13002, which is a TetR repressed POPS/RIPS generator, again part of the spring 2010 distribution - kit plate 1, well 13B, plasmid pSB1A2. It was proven to work by UMN iGEM 2009.
Wavelengths
548 nm http://pubs.acs.org/doi/pdf/10.1021/bi7015052
613 nm http://www.promega.com/pnotes/85/10904_11/10904_11.pdf
| Luciferase | Colour(at pH7.8) | λmax(nm) | (pH 7.8) | (pH 6.0) | Base change | Amino acid change | |
|---|---|---|---|---|---|---|---|
| Genjii | yellow-green | 562 | 609 | ||||
| C-M-1 | orange | 607 | 614 | G857-->A | Ser286-->Asn | ||
| C-M-2 | red | 609 | 611 | G976-->A | Gly326-->Ser | ||
| C-M-3 | red | 612 | 612 | C1297-->T | His433-->Tyr | ||
| C-M-4 | yellow-orange | 595 | 609 | C1354-->T | Pro452-->Ser | ||
| C-M-6 | green | 558 | 558 | G715-->A | Val239-->Ile | ||
| C-M-11 | yellow | 565 | 612 |
 "
"