Switch
From 2010.igem.org
|
||||||||
|
|
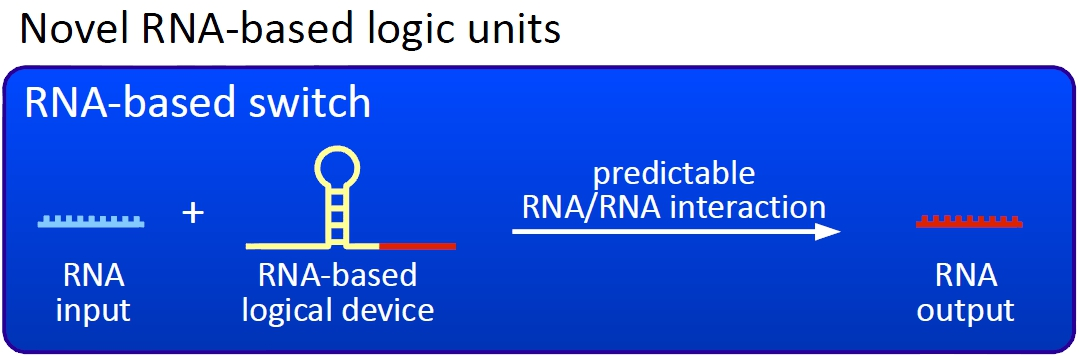
conceptbioLOGICSOur switches are based on the principle of transcriptional antitermination. Transcription can be cancelled by the formation of a RNA stem loop in the nascent RNA-chain, which then causes the RNA polymerase to stop transcription and fall off the DNA. We use sequences as transcriptional switches, that are calculated to be capable of stem loop formation. We plan to control the termination at these switches with a small RNA molecule (our RNA "signal") that is complementary to a part of the stem loop forming sequence. This small functional RNA inhibits the stem loop formation by complementary base-pairing and hence avoids transcription termination. The signal is composed of two parts: While the first part provides specifity, the second part causing stem loop disintegration can in principle be the same for all bioLOGICS. Therefore variation of the first part allows the construction of an endless number of switches. The second part causing stem loop disintegration is based on a working system (see attenuation) established by nature. Different stem loops were tested in this effort: Regulatory parts from the E. coli trp-operon, his-operon and one based on previous iGEM-work.
|
|||||||
 "
"