Team:SDU-Denmark/labnotes2
From 2010.igem.org
(Difference between revisions)
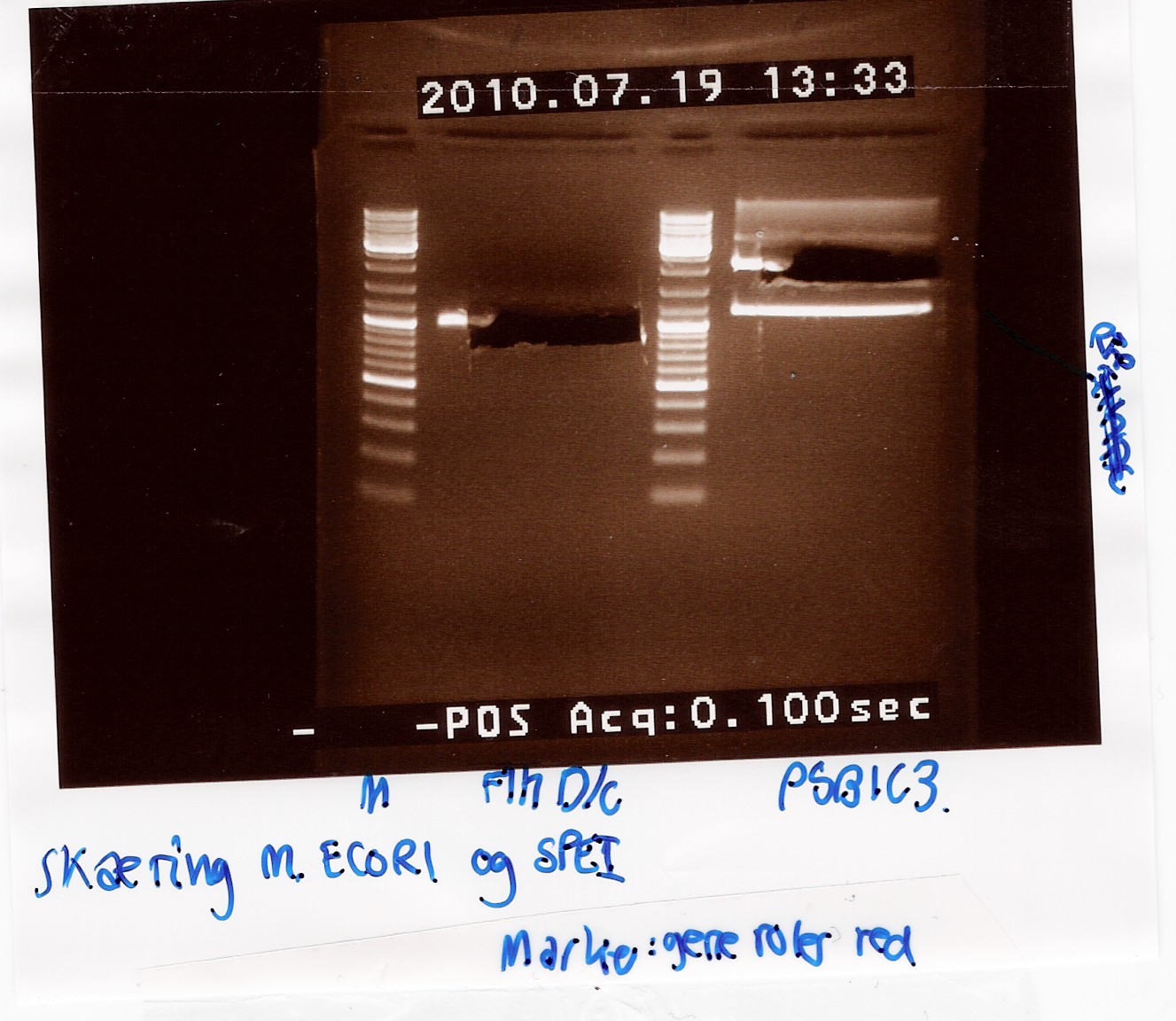
(→Gel extraction of flhD/C) |
(→Group: Retinal) |
||
| Line 392: | Line 392: | ||
3 parallel transformations were carried out for L1, L2 and L3 respectively (see [https://2010.igem.org/Team:SDU-Denmark/labnotes2#Ligation_of_flhD.2FC_and_pSB1C3 Ligation]). L1.1, L1.2, L2.1, L2.2 and L3.1 were transformed using compotent cells from 19/7 ([https://2010.igem.org/Team:SDU-Denmark/labnotes2#Transformation_of_K081005_in_pSB1A2_.28constitutive_promoter_and_RBS_combined.29.2CR0011_in_pSB1A2.2C_pSB3C5_w._J04450_and_pSB3T5_w._J04450_in_Top_10_E.Coli Compotent cells]). <br> | 3 parallel transformations were carried out for L1, L2 and L3 respectively (see [https://2010.igem.org/Team:SDU-Denmark/labnotes2#Ligation_of_flhD.2FC_and_pSB1C3 Ligation]). L1.1, L1.2, L2.1, L2.2 and L3.1 were transformed using compotent cells from 19/7 ([https://2010.igem.org/Team:SDU-Denmark/labnotes2#Transformation_of_K081005_in_pSB1A2_.28constitutive_promoter_and_RBS_combined.29.2CR0011_in_pSB1A2.2C_pSB3C5_w._J04450_and_pSB3T5_w._J04450_in_Top_10_E.Coli Compotent cells]). <br> | ||
Prior to transformation test plasmid was washed with 10xTE pH 8.0. <br> | Prior to transformation test plasmid was washed with 10xTE pH 8.0. <br> | ||
| - | --[[User:Tipi|Tipi]] 15:19, 20 July 2010 (UTC) | + | --[[User:Tipi|Tipi]] 15:19, 20 July 2010 (UTC)<br><br> |
| + | |||
| + | === Coloni PCR of R0011 in pSB1A2 and K081005 in pSB1A2 === | ||
| + | Start date: 20/07 End date: 20/07<br> | ||
| + | ''Methods:'' Coloni PCR and gel electrophoresis <br><br> | ||
| + | ''Protocol:''[https://2010.igem.org/Team:SDU-Denmark/protocols#CP1.2 CP1.2] <br><br> | ||
| + | ''Experiment done by:'' Maria and LC<br><br> | ||
| + | ''Notes:''<br> | ||
| + | 6 colonies transformed with pSB1A2 w. R0011 and 4 colonies transformed with pSB1A2 w. K081005 (see [https://2010.igem.org/Team:SDU-Denmark/labnotes2#Transformation_of_K081005_in_pSB1A2_.28constitutive_promoter_and_RBS_combined.29.2CR0011_in_pSB1A2.2C_pSB3C5_w._J04450_and_pSB3T5_w._J04450_in_Top_10_E.Coli Transformation]) was used for coloni PCR. <br> | ||
| + | Colonies with K081005 was denoted A1, A2 A3 and A4 respectively.<br> | ||
| + | Colonies with R0011 was denotes B1, B2, B3, B4, B5 and B6 respectively<br> | ||
| + | PCR was made with only half the amount of premix.<br> | ||
| + | Premix for 12 PCR reactions: <br> | ||
| + | <table style="text-align: left; width: 100px;" border="1" | ||
| + | cellpadding="2" cellspacing="2"> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top;">10xtaq buffer<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top;">30uL<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top;">MgCl2<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top;">12uL<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top;">VF2<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top;">12uL<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top;">VR<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top;">12uL<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top;">dNTP<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top;">6uL<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top;">H2O<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top;">42uL<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top;">Total vol.<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top;">114uL<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | </table> <br><br> | ||
| + | 0.5uL Taq polymerase from ampliqon was used.<br> | ||
| + | Taq PCR program:<br> | ||
| + | <table style="text-align: left; height: 260px; width: 225px;" border="1" | ||
| + | cellpadding="2" cellspacing="2"> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top; width: 102px;">1:Start<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 52px;">94C<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 45px;">2 min<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top; width: 102px;">2: Denaturing<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 52px;">94C<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 45px;">1 min<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top; width: 102px;">3: Annealing<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 52px;">55C<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 45px;">1 min<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top; width: 102px;">4: Elongation<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 52px;">72C<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 45px;">30 s<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top; width: 102px;">5:<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 52px;">GO TO<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 45px;">rep. 29x<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top; width: 102px;">6: End <br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 52px;">72C<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 45px;">3 min<br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="vertical-align: top; width: 102px;">7: Hold<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 52px;">4C<br> | ||
| + | </td> | ||
| + | <td style="vertical-align: top; width: 45px;"><br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | </table> <br><br> | ||
| + | PCR product was loaded onto a 2% agarose gel. Gene ruler 100bp DNA ladder was used as marker.<br><br> | ||
| + | ''Results:''<br><br> | ||
| + | ''Analysis:''<br> | ||
| + | --[[User:Tipi|Tipi]] 16:21, 20 July 2010 (UTC) | ||
</div> | </div> | ||
<div id="rightcolumn"> | <div id="rightcolumn"> | ||
Revision as of 16:21, 20 July 2010
 "
"