Team:UCL London/Genetic Circuit
From 2010.igem.org
Genetic Circuit
The SyntheticBiology.org website defines Synthetic Biology as:
A) the design and construction of new biological parts, devices, and systems, and B) the re-design of existing, natural biological systems for useful purposes.
In our case, we did both! When it comes to the modelling, we have successfully designed and constructed a whole new system, based on new and existing biological parts.
The challenges around the biological systems are the following, and summarise the scope of the formation of the iGEM competition:
• Defining the device physics of molecular components
• Building and simulating models
• Using defined components from a standard library (biobricks)
• Building new biological entities based on quantitatively predictive designs
Mathematical Equations
The design of a biological system should include all the information needed to describe what is happening within the cell (Carlson, 2010). That is why a set of mathematical equations were developed to describe the system's performance and the simulations, based on these equations, could be visualised.
The cartoon description used to describe the protein expression as a result of the transcription of DNA into RNA by polymerase which is in turn translated into protein. The ribosomes, which are not included in the animation, play an important role in the production of the proteins, according to the “central dogma”, defined by Francis Crick in 1957 that information flows from DNA to RNA to protein.
The genetic switch, that we aimed to build in the bacterium E.coli, was to be independent of the application of a chemical signal, such as IPTG, that would induce the cells to start producing the protein. The bacterium, when the switch is on, is predicted to make red fluorescent protein; when the switch is off, the protein is no longer made and any remaining protein decays.
Video Simulations
The BCCS-Bristol have created the video simulations of the genetic circuit which is a visual representation of the protein expression. The BSim simulation was on Gene Regulatory Network model and the video shows first the Linear system, which is what it is currently in the biopharmaceutical industry. The system is normalized, with zero initial conditions. The 2nd part of the video shows that bacterial RFP expression based on the hypoxia promoter and the positive feedback loop , which is much quicker than the normal linear one. Click on the BSim_BCCS-Bristol collaboration for UCL iGEM 2010 to watch the video simulation!
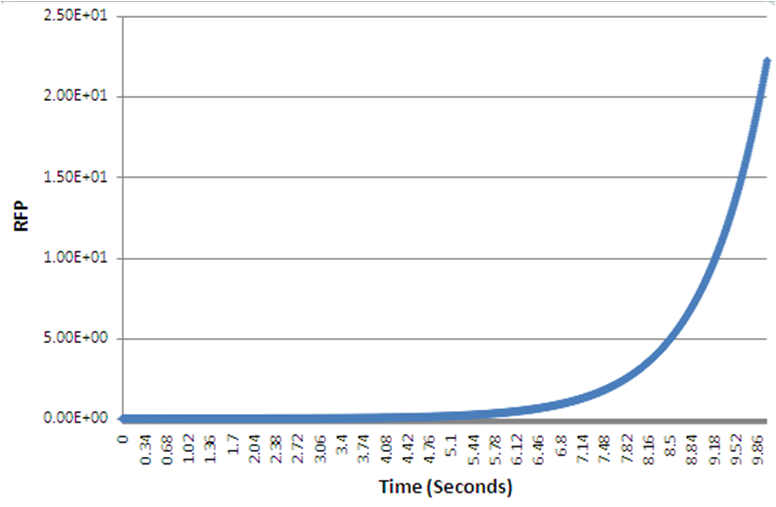
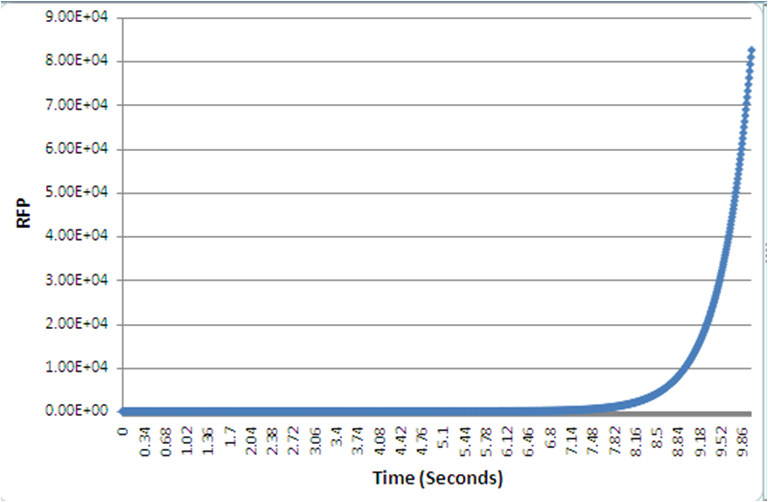
As in with our case, the numerical simulation of all the biochemical reactions identified, was to describe the system, and specific values such as the rate constant, are unknown, leading to the simulation exploring a wide variety of possible values from literature. A 4-fold increase was shown, over 10s, given normalized parameters. Despite the timescale, and in a different set of parameters, still the positive feedback loop yields quicker protein expression than the conventional, standard IPTG method.
Even if we did have particular values for the variables appearing in the model, it would still be impossible to make accurate predictions about its behaviour in real life. According to Carslon (2010), the line between model and simulation becomes blurred when the only way to assess a particular model is depending upon the simulation of the model. However, again the design stage with the simulation differs significantly from the actual model. This is where the difficulty lays, as scientists and engineers, work on projects together by repeatedly comparing simulation and measurement through experiments. The BCCS-Bristol team for this year iGEM competition, has assisted us, in the production of a standard simulation, with the potential to run multiple simulations using their BSim programme, to explore the behaviour of the model, over a range of conditions.
Modelling Animation
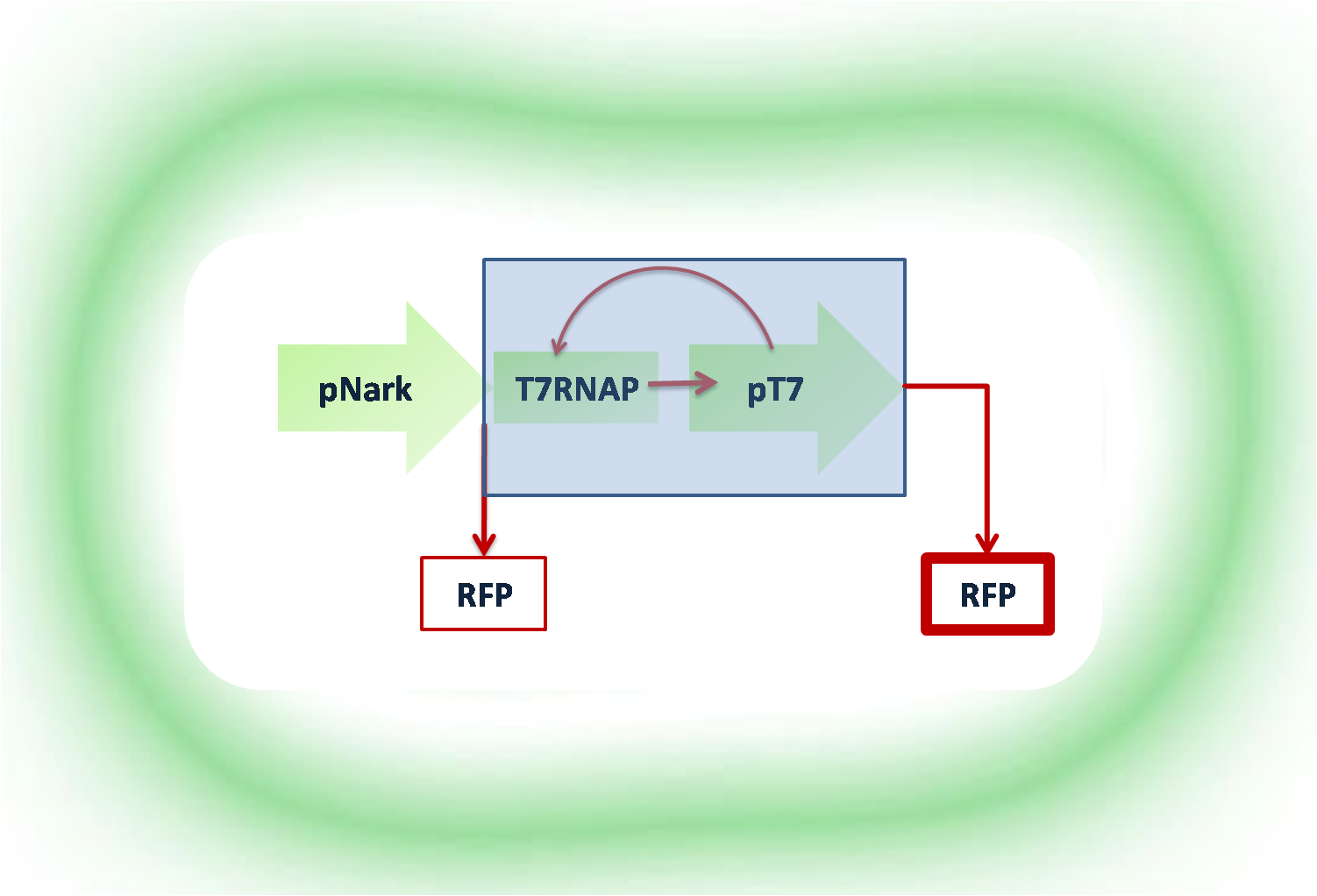
The fundamental that the genetic circuit works on, is that a signal from outside the cell, that is when the oxygen spike appears, declaring the low oxygen concentration present, induces the cell to begin producing the proteins. Traditionally, this was done with the manual addition by the operator of IPTG to the cells. The employment of inducible promoters, i.e. pNark, that works by interfering with a repressor protein, LacI, a protein that inhibits transcription by binding to the promoter. The polymerase, T7RNA P, is located downstream of the pNark, where it binds to DNA and begins transcription. The repressor protein, LacI, blocks the access to the pLacI promoter, because it is prevented by the pNark promoter. Therefore the role of the pNark inducible promoter is double: to accept the signal for cell induction and production of mRNA and proteins through the loop and also to inhibit the repressor binding to DNA. The addition of IPTG into the system, will cause the induction of the pLacI promoter which was repressed by the LacI proteins during the duration of the positive feedback loop and will be the regulator. The anti-PA binds to the promoter activator and reversibly stops the positive loop and hence, transcription terminates.
According to Purnick and Weiss (2009), the first wave of synthetic biology includes the combination of promoters, RBS (ribosome binding sites) and transcriptional repressors for the formation of functional modules e.g. controlled protein expression. Such a novel circuit design was based on principles like the creation and analysis of a genetic circuit, the experimental evaluation of the circuit and the combatibility of the results with literature. Moreover, they refer to directed evolution, which is about the optimisation of biobricks and the system i.e. the biological circuit.
The graphs, as produced with the BSim programme, show the protein expression. The positive feedback loop gives a much quicker red fluorescent protein expression, or within a specific timeframe it yields a higher protein expression compared to the normal IPTG method.
Therefore, as more “mature” engineering fields exist like mechanical and electrical, the project was “expressed” in those terms. For example the standard computational programmes SolidWorks and Verilog are powerful tools for the expression of equipment and circuits respectively in the engineering fields mentioned above. Comparatively engineering toolboxes that will enable the visual construction of models, using those programmes for the needs of a synthetic biology, will enable a quantitative understanding of how parts of a biological system when come together behave in different manners. Through this combination it is believed that the whole perspective of Synthetic Biology will be transformed (Carlson, 2009).
The construction of a database with well characterised genetic parts, with assigned mathematically abstracted functions that can be modelled in a software and turning electronic circuits into molecule representations via DNA synthesis could lead to a more economically and social value of what Synthetic Biology projects can offer. The currently available computational tools for the representation of genetic networks, include Antimony, Athena, BioJade, GenoCAD, OptCircuit, SynBioSS and many more software tools which were constructed to integrate all the relevant aspects of a biological system (Purnick and Weiss, 2009).


 "
"


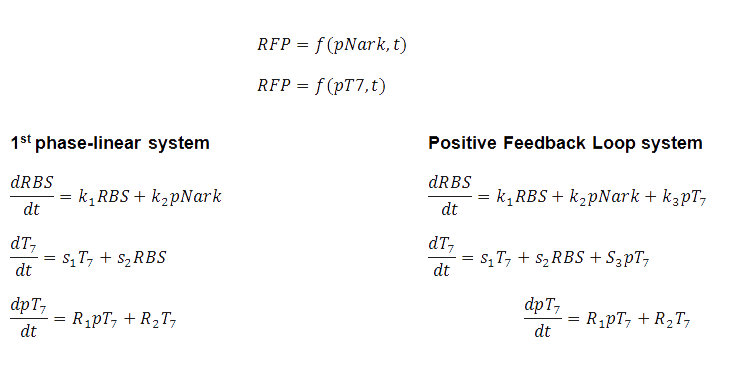

 Twitter
Twitter Facebook
Facebook UCL
UCL Flickr
Flickr YouTube
YouTube