Home
People
Project Details
Protocols
Completed Systems
Notebook
Biosafety
Human Impact
References
Support
|
Monday 7/26
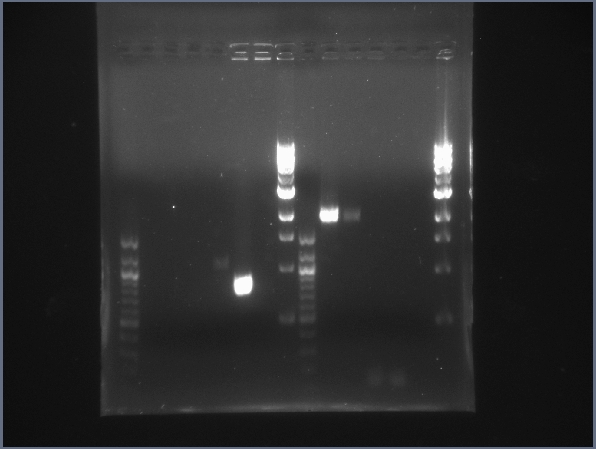 CPCR gel featuring L16, L19, L23, L24, L31 & L34. (7/26) 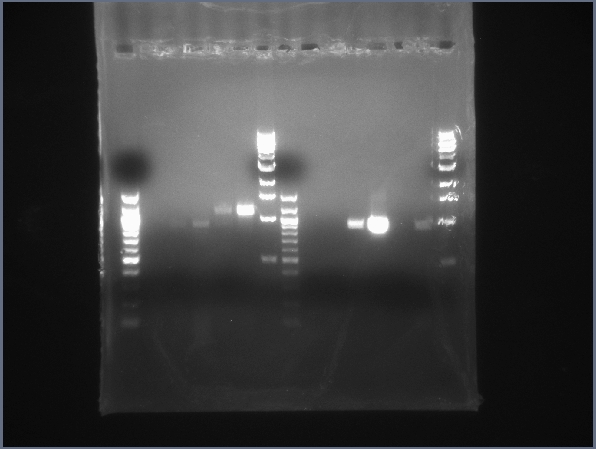 CPCR gel featuring L34, L35, L40, L42, L43 & L44. (7/26) - Started PCR reaction to extract the HSP (K112400) from its Berkeley Standard backbone (BBb). (Used same PCR parameters as CPCR from 7/22.
- Group meeting!
- By next week: Need to design test-construct experiments, as they will be ready sometime early next week.
- Can use PCR machine to achieve easy temperature gradients
- Use spectrophotometer and eventually microscope to assay fluorescence.
- Consider a plate assay?
- AIBN issues:
- Consider using low-concentration DMSO to help dissolve AIBN in LB/cell mixture.
- Need to figure out how much unreacted (or otherwise) AIBN remains after mixing with cells.
- Need to test crosslinking using hydrolyzed and epoxidized oil mixed with lysate and with cells to tweak concentration of AIBN needed, and to see what it makes.
- TGase:
- purchase urea?
- To prevent a solid plug forming upon addition of TGase powder to sample, prepare liquid stock of concentrated TGase.
- Wiki:
- Should visit biofab.org for research material for human impact segment.
- In general, we should set goals for both the end of SURF and for the end of the summer and determine what we need to do to get on track for them.
- Printing in 2D
- Working AND gate, at least in bulk fluid
- Characterization of 3D hydrogel/plastic
- Crosslinking data
- PCR purified K112400 PCR product & digested as plasmid insert.
- Transformed RFP bricks of verious resistances to use as backbone for future ligations
- Ran 2 gels of today's CPCR products.
Tuesday 7/27
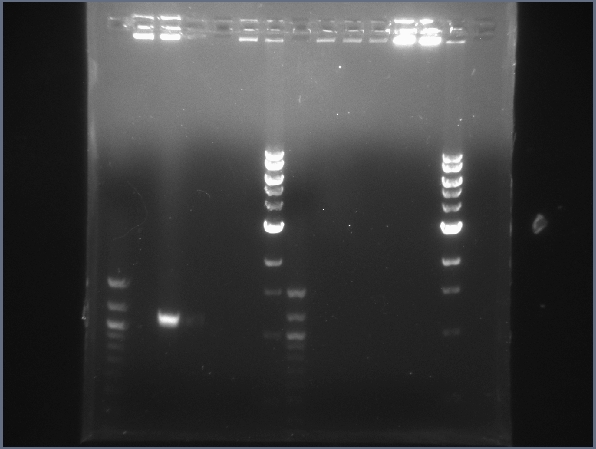 CPCR gel featuring L16, L19, L31, L34, & L35. (7/22) - Began more CPCR on failed ones from yesterday: L16, L19, L31, L34, L35
- Began more digests of miniprepped DNA from K112400
- Made more electrocompetent cells. Began two test transformations.
- Transformed ligation products: L42, L46, L47 and the HSP.
- Began 11 new ligations:
- L48: R0082 + I13504 + Cm
- L31: L17 + K215000 + Kan
- L34: L17 + L21 + Amp
- L35: L18 + L21 + Amp
- L22: K156012 + B0034 + Cm
- L24: K156014 + B0015 + Cm
- L20: C0077 + B0015 + Cm
- L49: K112400 + L40 + Cm
- L25: B0015 + M30109 + Tet
- L26: R0077 + K124017 + Cm
- L50: L18 + L27 + Amp
- Note: Need to upstream-digest more of: J23119, L17, K156014, B0015 for future ligations.
Wednesday 7/28
- Began CPCR reactions on the HSP, L42, L46, L48.
- Transformed all ligations from yesterday
- Re-ran gel from yesterday
- Same results: looks like 19-4 is a success
- Ran gel of today's CPCR to determine if the HSP ligations worked.
- All bands appear to have a 100-400bp smear and no other bands, making the results indeterminate.
Thursday 7/29
- Re-did the following ligations: L16, L31, L34, L35, HSP (in pSB1C3, for submission), L42, L46, L47
- Started epic amounts of colony PCR:
- 3 colonies of each: L42, L46, L31, HSP-Amp, L35, L50, L34, L48, L24, L49, L22, L20
- Gels for each are included to the right ->
- Successful: L42-4/-5, L46-4/-5, L47-4, HSP-Amp-5/-6
Friday 7/30
- Prepped 47-4 (HSP + B0034), L42-4/-5, L46-4/-5 (HSP + R0077), HSP-Amp-5/-6
- Began a ton of digests:
- 2x Tet backbone
- U19: K156012 (us)
- U20: K156014 (us)
- U21: C0077 (us)
- U22: L19 (us)
- U23: L46 (us)
- U24: L47 (us)
- U25: L17 (us)
- D17: B0034 (ds)
- D18: B0015 (ds)
- D19: L18 (ds)
- D20: L19 (ds)
- D21: HSP-Amp (ds)
Weekend 7/31-8/1
Saturday 7/31
- Transformed L22, L24, L33, L43
- Failed: L51, L52, L53, L25 (cuvette arced)
- Ran gel of M30109 CPCR product: no bands!
Sunday 8/1
- Removed transformation plates: L22 & L24 were successful; L33 & L53 failed.

|
 "
"



