Home
People
Project Details
Protocols
Completed Systems
Notebook
Biosafety
Human Impact
References
Support
|
Monday 7/12
- Group meeting today!
- Sequencing results were really crappy (quality < 30% confidence), so began 2 5mL LB cultures of the light-lysis construct (picked from the original transformation plate).
Tuesday 7/13
- Sent off two new samples of the light-lysis construct for sequencing using the VF2 and VR primers. Results should be available tonight or tomorrow.
- Used remaining cells to inoculate new cultures of each and to extract DNA for a test digest:
- Digested each (ideally identical) sample with EcoRI-HF and PstI, both for 1hr and for 10min.
- The digest appears inconclusive, as a smear is visible from the top to about 3Kb, while little or no DNA is visible in the insert range (~1.5Kb, see below).
- Will run another gel tonight with other bricks to check efficacy of digests and will look forward to sequencing results soon.
- Second digest can be found below. Lane 1: 4μL 100bp ladder (NEB); Lane 2: 5μL pSB1T3 digest; Lane 3: 10μL lysis gene downstream digest; Lane 4: 10μL GFP upstream digest; Lane 5: 4μL 1kb ladder (NEB).
- Lanes appear to have correct bands; digests and ligations appear to be working better now. (We now run digests for 60 min at 37°C and ligations for 30 min at RT, contrary to the times suggested by NEB.)
- Ordered more SpeI from NEB, since we're basically out.
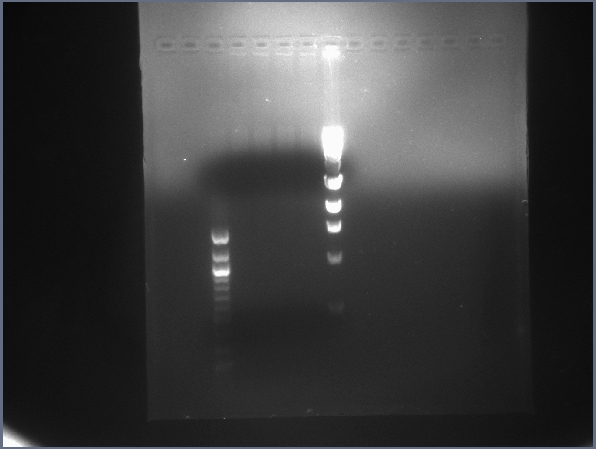 Gel image for the light-lysis test digest. (7/13) 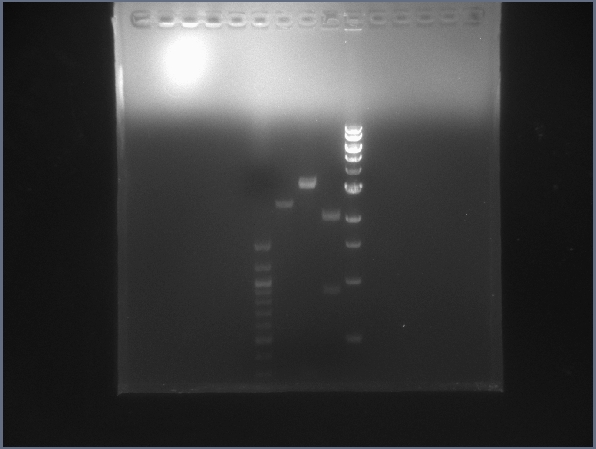 Digest test gel featuring digested backbone, US and DS digested bricks. (7/13) - Performed initial growth assay with cells in different solutions mixed with varying levels of AIBN and other conditions.
Wednesday 7/14
- Received bricks from the Registry of Standard Parts today.
- The light-lysis sequencing results could not be returned due to insufficient DNA concentration. Picked two new colonies from the original plate. Also made cultures of the previous ligations (PR aka Ligation1 and lyT aka Ligation2). Will prep tomorrow for sequencing.
- Met with the Caltech Office of Technology Transfer (Drs. Hannah Dvorak-Carbone & Jennifer Hodas)
- Discussed patent law and IP issues relevant to our project
- Began designing primers for making glutamine & lysine peptides to experiment with transglutaminase (TGase).
- Transformed pBAD18 into both DH5α and V1012 to test competence & troubleshoot transformation failures. (Used 2μL DNA and 60μL cells.)
- Prepared SOC from SOB and made 10mL aliquots to freeze.
- Ran gel to test ligation products from today (and troubleshoot the light-lysis construct some more...)
- Lanes 2 & 3 contain Ligation1 & 2 digests. They have bands, but they do not appear to be the right size - we'll hasve to look at these a bit more. The other lanes are raw ligation product from today's ligations.
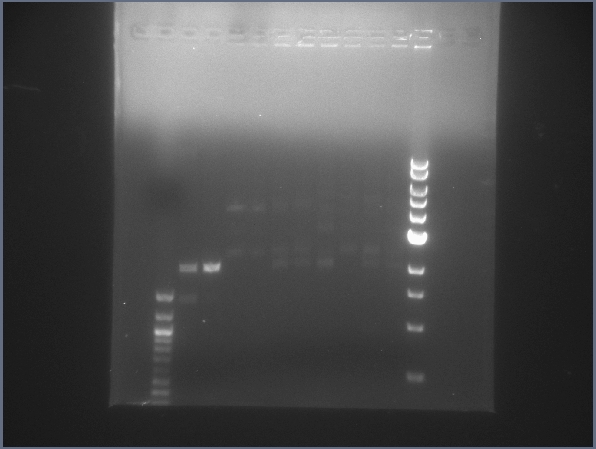 Gel of today's ligation products & light-lysis ligation digests. (7/14) Thursday 7/15
- The V1012 transformation was successful, but the V1012 was not. Will troubleshoot V1012 further, but it appears that the DH5α cells are competent.
- May want to make more V1012 cells competent this weekend.
- It appears that using at least 50μL of comp cells and 2μL of DNA works most effectively. We will also use SOC to rescue the transformations from now on.
- The PR (Ligation1) and PRlyT (Ligation3) cultures grew, but the lyT (Ligation2) culture did not. This could indicate an issue with the lyT product (perhaps the colonies on the plate are contamination).
- Will reperform the Ligation3 reaction and see if that helps. (The gel of the Ligation2 digest (above) looks correct, so there shouldn't be a problem with the ligation product.)
- Prepped the cultures (to send for sequencing)
- PRlyT1 = 50.1 ng/μL; PRlyT2 = 34.5 ng/μL (will send 20μL of each for seq.)
- PR1 = 197.0 ng/μL; PR2 = 201.0 ng/μL (will send 10μL of each for seq.)
- Team signed the OTT Invention Disclosure form and submitted a project description to the OTT. Provisional patent application will be filed tomorrow.
- Designed primers for BBb_K112400 to change from Berkley standard to the correct BBa standard.
- Will order these tomorrow so they are ready when the heat-shock promoter arrives.
- Prepared 500mL 2YT media, 1L diH2O, 2 250mL Erlenmeyer flasks and 50mL 10% glycerol. Inoculated 1 5mL culture of 2YT with DH5α cells.
- Will make more competent cells tomorrow.
- Made minipreps of the bricks received on Wednesday.
Friday 7/16
- Lab clean-up scheduled for today.
- All 11 ligation transformations failed (except the positive control). This seems like a DNA issue - we will try transforming more DNA today after our competent cells are ready.
- Performed solubility and reactivity test on AIBN on soybean and linseed oil.
- Performed live-dead assay on cells under different conditions with solutions of varying AIBN concentrations.
- Ordered primers to fix a BioBrick part.
- Ordered a synthesized BioBrick part encoding a poly-lysine and poly-glutamine polypeptide sequence.
Weekend 7/17-18
- DH5α competence test: positive control grew cells, negative did not -> Cells are competent and not contaminated.
- Transformation plates: L15 (light-lysis construct): failed; L4: >1000 colonies (will reculture to verify antibiotic resistance); L8: 2 colonies (will also reculture to verify that colonies aren't contamination)
- Will also start cultures of K124017, as all digests of this brick appear incorrect. We want to try again with fresh DNA.
- Live-dead assay results: All plates from the start point in time contained films of cells. The plates created an hour later from glycerol solutions kept at 4°C showed little change in the amount of cells grown. Plates created from the room temperature solutions, showed either large colonies and/or less developed films.
- Transformed 5μL ligation product for ligations L4-L14 (again).
- Hydrolyzed soybean oil

|
 "
"



